

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.5
(852 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
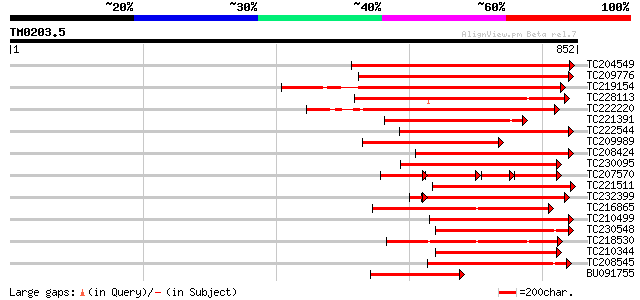
Score E
Sequences producing significant alignments: (bits) Value
TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-l... 422 e-118
TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-l... 410 e-114
TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.1... 405 e-113
TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 364 e-101
TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like prot... 331 9e-91
TC221391 similar to UP|O81906 (O81906) Serine/threonine kinase-l... 317 2e-86
TC222544 weakly similar to UP|O23743 (O23743) SFR1 protein precu... 314 1e-85
TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 283 2e-76
TC208424 weakly similar to UP|O65466 (O65466) Serine/threonine k... 283 2e-76
TC230095 weakly similar to UP|Q9XED4 (Q9XED4) Receptor-like prot... 281 1e-75
TC207570 weakly similar to UP|O65470 (O65470) Serine/threonine k... 107 7e-74
TC221511 similar to UP|Q40096 (Q40096) Receptor protein kinase, ... 264 1e-70
TC232399 similar to UP|Q39086 (Q39086) Receptor kinase , partial... 258 8e-69
TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial... 257 2e-68
TC210499 weakly similar to UP|Q9LDS6 (Q9LDS6) Serine/threonine k... 253 3e-67
TC230548 weakly similar to UP|Q8LLI4 (Q8LLI4) S-locus receptor-l... 244 9e-65
TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partia... 244 9e-65
TC210344 similar to UP|Q9AVE0 (Q9AVE0) SRKb, partial (21%) 243 3e-64
TC208545 weakly similar to UP|Q9S972 (Q9S972) ARK2 product/recep... 239 3e-63
BU091755 239 5e-63
>TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-like protein,
partial (54%)
Length = 2032
Score = 422 bits (1085), Expect = e-118
Identities = 203/335 (60%), Positives = 250/335 (74%)
Frame = +3
Query: 514 KEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLS 573
KEG D ++ FDF +I AT+ FS NKLG GG+G VYKG L G+ +AVKRLS
Sbjct: 744 KEGKTAYDIPTVDSLQFDFSTIEAATNKFSADNKLGEGGFGEVYKGTLSSGQVVAVKRLS 923
Query: 574 SVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKS 633
S QG +EFKNEVV++AKLQHRNLVRL G+C++G+EKIL+YEY+PNKSLD +FDP K
Sbjct: 924 KSSGQGGEEFKNEVVVVAKLQHRNLVRLLGFCLQGEEKILVYEYVPNKSLDYILFDPEKQ 1103
Query: 634 ALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFG 693
LDW R+ I+ GIARG+ YLH+DSRLR+IHRDLK SNILLDG+M PKISDFG+ARIFG
Sbjct: 1104 RELDWGRRYKIIGGIARGIQYLHEDSRLRIIHRDLKASNILLDGDMNPKISDFGMARIFG 1283
Query: 694 GKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTL 753
+T+ NT R+VGTYGYM+PEYA+ G+FS KSD++SFGV+L+EI+SGKKN+ FYQ G
Sbjct: 1284 VDQTQGNTSRIVGTYGYMAPEYAMHGEFSVKSDVYSFGVLLMEILSGKKNSSFYQTDGAE 1463
Query: 754 SLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIML 813
LL YAW+LW + L+LMD L E+YN N+ IR H+GLLCVQ++P DRP M+ +V+ML
Sbjct: 1464 DLLSYAWQLWKDGTPLELMDPILRESYNQNEVIRSIHIGLLCVQEDPADRPTMATIVLML 1643
Query: 814 DSETATLPTPKQPTFFTRKDLSSTASSSLQFDSSI 848
DS T TLPTP QP FF L FD SI
Sbjct: 1644 DSNTVTLPTPTQPAFFVHSGTDPNMPKELPFDQSI 1748
>TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-like protein
, partial (47%)
Length = 1305
Score = 410 bits (1053), Expect = e-114
Identities = 193/323 (59%), Positives = 246/323 (75%)
Frame = +2
Query: 525 IEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFK 584
+E FDF +I AT FS+ANKLG GG+G VYKG L G+E+AVKRLS +S QG +EFK
Sbjct: 29 VESLRFDFSTIEAATQKFSEANKLGEGGFGEVYKGLLPSGQEVAVKRLSKISGQGGEEFK 208
Query: 585 NEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDI 644
NEV ++AKLQHRNLVRL G+C++G+EKIL+YE++ NKSLD +FDP K LDW R+ I
Sbjct: 209 NEVEIVAKLQHRNLVRLLGFCLEGEEKILVYEFVANKSLDYILFDPEKQKSLDWTRRYKI 388
Query: 645 LLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRV 704
+ GIARG+ YLH+DSRL++IHRDLK SN+LLDG+M PKISDFG+ARIFG +T+ANT R+
Sbjct: 389 VEGIARGIQYLHEDSRLKIIHRDLKASNVLLDGDMNPKISDFGMARIFGVDQTQANTNRI 568
Query: 705 VGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWT 764
VGTYGYMSPEYA+ G++S KSD++SFGV++LEIISGK+N+ FY+ LL YAWKLW
Sbjct: 569 VGTYGYMSPEYAMHGEYSAKSDVYSFGVLILEIISGKRNSSFYETDVAEDLLSYAWKLWK 748
Query: 765 ENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPK 824
+ L+LMD SL E+Y N+ IRC H+GLLCVQ++P DRP M++VV+MLDS + TL P
Sbjct: 749 DEAPLELMDQSLRESYTRNEVIRCIHIGLLCVQEDPIDRPTMASVVLMLDSYSVTLQVPN 928
Query: 825 QPTFFTRKDLSSTASSSLQFDSS 847
QP F+ L+ D S
Sbjct: 929 QPAFYINSRTEPNMPKGLKIDQS 997
>TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.110 precursor
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (31%)
Length = 1363
Score = 405 bits (1040), Expect = e-113
Identities = 220/426 (51%), Positives = 280/426 (65%)
Frame = +1
Query: 409 SPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVI 468
S C +W +L +K + R+L +R+ S++E K + K + + L GVV+
Sbjct: 43 SGCVMWFGDLLDIKLYSVAESGRRLHIRLPPSELESIKSKKSSKIIIGTSVAAPL-GVVL 219
Query: 469 LACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVP 528
C ++ RR IA K K + +SI RQ + ++VP
Sbjct: 220 AIC-----FIYRRNIADKSKTK-KSIDRQL------------------------QDVDVP 309
Query: 529 YFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVV 588
FD +I ATD F NK+G GG+GPVYKGKL GG+EIAVKRLSS+S QGI EF EV
Sbjct: 310 LFDMLTITAATDNFLLNNKIGEGGFGPVYKGKLVGGQEIAVKRLSSLSGQGITEFITEVK 489
Query: 589 LIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGI 648
LIAKLQHRNLV+L G CIKG EK+L+YEY+ N SL++F+FD KS LLDW RF+I+LGI
Sbjct: 490 LIAKLQHRNLVKLLGCCIKGQEKLLVYEYVVNGSLNSFIFDQIKSKLLDWPRRFNIILGI 669
Query: 649 ARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTY 708
ARGLLYLHQDSRLR+IHRDLK SN+LLD ++ PKISDFG+AR FGG +TE NT RVVGTY
Sbjct: 670 ARGLLYLHQDSRLRIIHRDLKASNVLLDEKLNPKISDFGMARAFGGDQTEGNTNRVVGTY 849
Query: 709 GYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKL 768
GYM+PEYA DG FS KSD+FSFG++LLEI+ G KN TL+L+GYAW LW E
Sbjct: 850 GYMAPEYAFDGNFSIKSDVFSFGILLLEIVCGIKNKSLCHENQTLNLVGYAWALWKEQNA 1029
Query: 769 LDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTF 828
L L+D + ++ + +RC HV LLCVQ P+DRP M++V+ ML SE + PK+P F
Sbjct: 1030LQLIDSGIKDSCVIPEVLRCIHVSLLCVQQYPEDRPTMTSVIQMLGSE-MDMVEPKEPGF 1206
Query: 829 FTRKDL 834
F R+ L
Sbjct: 1207FPRRIL 1224
>TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (53%)
Length = 1492
Score = 364 bits (935), Expect = e-101
Identities = 183/362 (50%), Positives = 250/362 (68%), Gaps = 39/362 (10%)
Frame = +1
Query: 518 EEKDNEGIEVPY---FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSS 574
+ +D++ IE+ F+F++I VAT+ FSD+NKLG+GG+G VY+G+L G+ IAVKRLS
Sbjct: 73 QHEDDDEIEIAQSLQFNFDTIRVATEDFSDSNKLGQGGFGAVYRGRLSDGQMIAVKRLSR 252
Query: 575 VSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVF------ 628
SSQG EFKNEV+L+AKLQHRNLVRL G+C++G E++LIYEY+PNKSLD F+F
Sbjct: 253 ESSQGDTEFKNEVLLVAKLQHRNLVRLLGFCLEGKERLLIYEYVPNKSLDYFIFGRS**V 432
Query: 629 ------------------------------DPTKSALLDWQMRFDILLGIARGLLYLHQD 658
DPTK A L+W+MR+ I+ G+ARGLLYLH+D
Sbjct: 433 HN*TVALYFTSGSGQRLNIHIPLKLMAVYADPTKKAQLNWEMRYKIITGVARGLLYLHED 612
Query: 659 SRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALD 718
S LR+IHRDLK SNILL+ EM PKI+DFG+AR+ +T+ANT R+VGTYGYM+PEYA+
Sbjct: 613 SHLRIIHRDLKASNILLNEEMNPKIADFGMARLVLMDQTQANTNRIVGTYGYMAPEYAMH 792
Query: 719 GQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGE 778
GQFS KSD+FSFGV++LEIISG KN+G + LL +AW+ W E + ++D SL
Sbjct: 793 GQFSMKSDVFSFGVLVLEIISGHKNSGIRHGENVEDLLSFAWRNWREGTAVKIVDPSLNN 972
Query: 779 AYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTRKDLSSTA 838
+ N+ +RC H+GLLCVQ+ DRP M+ +++ML+S + +LP P +P F+ S +
Sbjct: 973 -NSRNEMLRCIHIGLLCVQENLADRPTMTTIMLMLNSYSLSLPIPSKPAFYVSSRTGSIS 1149
Query: 839 SS 840
++
Sbjct: 1150AT 1155
>TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like protein kinase,
RLK3 precursor, partial (35%)
Length = 1304
Score = 331 bits (848), Expect = 9e-91
Identities = 176/382 (46%), Positives = 245/382 (64%), Gaps = 1/382 (0%)
Frame = +3
Query: 446 TRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSE 505
T GN KS ++L + V I+ +C+ ++ +R +++ ++LR+
Sbjct: 9 TYAGNKKSVSRVVLIVVPVVVSIILLLCVCYFILKRS-----RKKYNTLLRENF------ 155
Query: 506 RHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGR 565
G E E ++ F +I AT+ FS ++G GG+G VYKG L GR
Sbjct: 156 ----------GEESATLESLQ---FGLVTIEAATNKFSYEKRIGEGGFGVVYKGVLPDGR 296
Query: 566 EIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDA 625
EIAVK+LS S QG EFKNE++LIAKLQHRNLV L G+C++ EK+LIYE++ NKSLD
Sbjct: 297 EIAVKKLSKSSGQGANEFKNEILLIAKLQHRNLVTLLGFCLEEHEKMLIYEFVSNKSLDY 476
Query: 626 FVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISD 685
F+FD +S L+W R+ I+ GIA+G+ YLH+ SRL+VIHRDLK SN+LLD M PKISD
Sbjct: 477 FLFDSHRSKQLNWSERYKIIEGIAQGISYLHEHSRLKVIHRDLKPSNVLLDSNMNPKISD 656
Query: 686 FGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTG 745
FG+ARI + + T R+VGTYGYMSPEYA+ GQFS KSD+FSFGV++LEIIS K+NT
Sbjct: 657 FGMARIVAIDQLQGKTNRIVGTYGYMSPEYAMHGQFSEKSDVFSFGVIVLEIISAKRNTR 836
Query: 746 FYQYKGTLSLLGYAWKLWTENKLLDLMDLSL-GEAYNANQFIRCTHVGLLCVQDEPDDRP 804
+ LL Y W+ W + L++ D S+ E + ++ ++C +GLLCVQ++PDDRP
Sbjct: 837 SV-FSDHDDLLSYTWEQWMDEAPLNIFDQSIEAEFCDHSEVVKCIQIGLLCVQEKPDDRP 1013
Query: 805 NMSNVVIMLDSETATLPTPKQP 826
++ V+ L+S LP PK+P
Sbjct: 1014KITQVISYLNSSITELPLPKKP 1079
>TC221391 similar to UP|O81906 (O81906) Serine/threonine kinase-like protein
, partial (25%)
Length = 642
Score = 317 bits (811), Expect = 2e-86
Identities = 153/214 (71%), Positives = 182/214 (84%)
Frame = +2
Query: 564 GREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSL 623
G E+AVKRLS SSQG++EFKNE+VLIAKLQHRNLVRL G CI+G+EKIL+YEY+PNKSL
Sbjct: 2 GEEVAVKRLSRKSSQGLEEFKNEMVLIAKLQHRNLVRLLGCCIQGEEKILVYEYLPNKSL 181
Query: 624 DAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKI 683
D F+FDP K LDW RF+I+ GIARGLLYLH+DSRLR+IHRDLK SNILLD M PKI
Sbjct: 182 DCFLFDPVKQTQLDWAKRFEIIEGIARGLLYLHRDSRLRIIHRDLKASNILLDESMNPKI 361
Query: 684 SDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKN 743
SDFGLARIFGG + EANT RVVGTYGYMSPEYA++G FS KSD++SFGV+LLEI+SG+KN
Sbjct: 362 SDFGLARIFGGNQNEANTNRVVGTYGYMSPEYAMEGLFSIKSDVYSFGVLLLEIMSGRKN 541
Query: 744 TGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLG 777
T F + SL+GYAW LW+E ++++L+D SLG
Sbjct: 542 TSFRDTDDS-SLIGYAWHLWSEQRVMELVDPSLG 640
>TC222544 weakly similar to UP|O23743 (O23743) SFR1 protein precursor ,
partial (27%)
Length = 861
Score = 314 bits (804), Expect = 1e-85
Identities = 154/264 (58%), Positives = 200/264 (75%), Gaps = 2/264 (0%)
Frame = +3
Query: 586 EVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDIL 645
EVV+I+KLQHRNLVRL G CI+ DE++L+YE+MPNKSLD+F+FDP + +LDW+ RF+I+
Sbjct: 3 EVVVISKLQHRNLVRLLGCCIERDEQMLVYEFMPNKSLDSFLFDPLQRKILDWKKRFNII 182
Query: 646 LGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIF-GGKETEANTQRV 704
GIARG+LYLH+DSRLR+IHRDLK SNILLD EM PKISDFGLARI G + EANT+RV
Sbjct: 183 EGIARGILYLHRDSRLRIIHRDLKASNILLDDEMHPKISDFGLARIVRSGDDDEANTKRV 362
Query: 705 VGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWT 764
VGTYGYM PEYA++G FS KSD++ V+LLEI+SG++NT FY + +LSL YAWKLW
Sbjct: 363 VGTYGYMPPEYAMEGIFSEKSDVYXXXVLLLEIVSGRRNTSFYNNEQSLSLXXYAWKLWN 542
Query: 765 ENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPK 824
E + ++DL + + +RC H+GLLCVQ+ +RP +S VV ML SE LP P+
Sbjct: 543 EGNIKSIIDLEIQDPMFEKSILRCIHIGLLCVQELTKERPTISTVVXMLISEITHLPPPR 722
Query: 825 QPTFFTRKDL-SSTASSSLQFDSS 847
Q F +++ SS +S QF+S+
Sbjct: 723 QXXFVQKQNCQSSESSQKSQFNSN 794
>TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (32%)
Length = 738
Score = 283 bits (725), Expect = 2e-76
Identities = 136/213 (63%), Positives = 174/213 (80%)
Frame = +3
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
F+F +I AT+ FSDANKLG+GG+G VYKG L G+EIA+KRLS S+QG EFKNE++L
Sbjct: 99 FEFATIKFATNNFSDANKLGQGGFGIVYKGTLSDGQEIAIKRLSINSNQGETEFKNEILL 278
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIA 649
+LQHRNLVRL G+C E++LIYE++PNKSLD F+FDP K L+ ++R+ I+ GIA
Sbjct: 279 TGRLQHRNLVRLLGFCFARRERLLIYEFVPNKSLDYFIFDPNKRVNLN*EIRYKIIRGIA 458
Query: 650 RGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYG 709
RGLLYLH+DSRL V+HRDLKTSNILLDGE+ PKISDFG+AR+F +TEA+T +VGT+G
Sbjct: 459 RGLLYLHEDSRLNVVHRDLKTSNILLDGELNPKISDFGMARLFEINQTEASTTTIVGTFG 638
Query: 710 YMSPEYALDGQFSTKSDIFSFGVVLLEIISGKK 742
YM+PEY GQFS KSD+FSFGV++LEI+ G++
Sbjct: 639 YMAPEYIKHGQFSIKSDVFSFGVMILEIVCGQR 737
>TC208424 weakly similar to UP|O65466 (O65466) Serine/threonine kinase-like
protein , partial (34%)
Length = 913
Score = 283 bits (725), Expect = 2e-76
Identities = 134/238 (56%), Positives = 174/238 (72%)
Frame = +2
Query: 610 EKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLK 669
EKILIYEY+ NKSLD F+FDP K LDW R+ I++GIARG+ YLH+DS+LR+IHRDLK
Sbjct: 2 EKILIYEYITNKSLDHFLFDPVKQKELDWSRRYKIIVGIARGIQYLHEDSQLRIIHRDLK 181
Query: 670 TSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFS 729
SN+LLD M PKISDFG+A+IF +T+ NT R+VGTYGYMSPEYA+ GQFS KSD+FS
Sbjct: 182 ASNVLLDENMNPKISDFGMAKIFQADQTQVNTGRIVGTYGYMSPEYAMRGQFSVKSDVFS 361
Query: 730 FGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCT 789
FGV++LEI+SGKKNT FYQ LL +AWK WT L+L+D +L +Y+ N+ RC
Sbjct: 362 FGVLVLEIVSGKKNTDFYQSNHADDLLSHAWKNWTLQTPLELLDPTLRGSYSRNEVNRCI 541
Query: 790 HVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTRKDLSSTASSSLQFDSS 847
H+GLLCVQ+ P DRP+M+ + +ML+S + T+ P+QP R + + + D S
Sbjct: 542 HIGLLCVQENPSDRPSMATIALMLNSYSVTMSMPQQPASLLRGRGPNRLNQGMDSDQS 715
>TC230095 weakly similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase
homolog RK20-1, partial (30%)
Length = 819
Score = 281 bits (718), Expect = 1e-75
Identities = 136/242 (56%), Positives = 175/242 (72%)
Frame = +3
Query: 588 VLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLG 647
+LIAKLQH+NLVRL G+C + EKILIYEY+PNKSLD F+FD K L W RF I+ G
Sbjct: 6 LLIAKLQHKNLVRLVGFCQEDREKILIYEYVPNKSLDHFLFDSQKHRQLTWSERFKIVKG 185
Query: 648 IARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGT 707
IARG+LYLH+DSRL++IHRD+K SN+LLD + PKISDFG+AR+ + + T RVVGT
Sbjct: 186 IARGILYLHEDSRLKIIHRDIKPSNVLLDYGINPKISDFGMARMVATDQIQGCTNRVVGT 365
Query: 708 YGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENK 767
YGYMSPEYA+ GQFS KSD+FSFGV++L+IISGKKN+ ++ LL YAW W +
Sbjct: 366 YGYMSPEYAMHGQFSEKSDVFSFGVMVLDIISGKKNSCSFESCRVDDLLSYAWNNWRDES 545
Query: 768 LLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPT 827
L+D +L E+Y N+ +C +GLLCVQ+ PDDRP M +V L + + +P P +P
Sbjct: 546 PYQLLDSTLLESYVPNEVEKCMQIGLLCVQENPDDRPTMGTIVSYLSNPSFEMPFPLEPA 725
Query: 828 FF 829
FF
Sbjct: 726 FF 731
>TC207570 weakly similar to UP|O65470 (O65470) Serine/threonine kinase-like
protein , partial (30%)
Length = 1285
Score = 107 bits (268), Expect(4) = 7e-74
Identities = 54/82 (65%), Positives = 64/82 (77%)
Frame = +1
Query: 626 FVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISD 685
F D + LLDW+ R +I+ GIA+GLLYLH+ SRL+VIHRDLK SNILLD EM KISD
Sbjct: 295 FKTDSARKDLLDWEKRLNIIGGIAQGLLYLHKYSRLKVIHRDLKASNILLDHEMNAKISD 474
Query: 686 FGLARIFGGKETEANTQRVVGT 707
FG+ARIFG + +E NT RVVGT
Sbjct: 475 FGMARIFGVRVSEENTNRVVGT 540
Score = 90.9 bits (224), Expect(4) = 7e-74
Identities = 42/72 (58%), Positives = 56/72 (77%)
Frame = +2
Query: 557 YKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYE 616
++G L +E+A+KRLS S QG+ EF NE L+AKLQH NLV+L G+CI+ DE+IL+YE
Sbjct: 2 HEGNLSDQQEVAIKRLSKSSGQGLIEFTNEAKLMAKLQHTNLVKLLGFCIQRDERILVYE 181
Query: 617 YMPNKSLDAFVF 628
YM NKSLD ++F
Sbjct: 182 YMSNKSLDFYLF 217
Score = 77.4 bits (189), Expect(4) = 7e-74
Identities = 32/71 (45%), Positives = 46/71 (64%)
Frame = +1
Query: 759 AWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETA 818
AW+LW + L+L+D +L + N+ RC H+GLLCVQD+ DRP M ++V L ++T
Sbjct: 865 AWQLWNAGRALELIDSTLNGLCSQNEVFRCIHIGLLCVQDQATDRPTMVDIVSFLSNDTI 1044
Query: 819 TLPTPKQPTFF 829
LP P QP +F
Sbjct: 1045QLPQPMQPAYF 1077
Score = 63.2 bits (152), Expect(4) = 7e-74
Identities = 29/50 (58%), Positives = 37/50 (74%)
Frame = +2
Query: 709 GYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGY 758
GYM+PEYA+ G S K+D+FSFGV+LLEI+S KKN Y L+L+GY
Sbjct: 629 GYMAPEYAMKGVVSIKTDVFSFGVLLLEILSSKKNNSRYHSDHPLNLIGY 778
>TC221511 similar to UP|Q40096 (Q40096) Receptor protein kinase, partial
(21%)
Length = 765
Score = 264 bits (674), Expect = 1e-70
Identities = 128/215 (59%), Positives = 165/215 (76%)
Frame = +3
Query: 636 LDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGK 695
LDW RF+I+ GIA+GLLYLHQDSRLR+IHRDLK SN+LLD E+ PKISDFG+ARIFG
Sbjct: 3 LDWSKRFNIICGIAKGLLYLHQDSRLRIIHRDLKASNVLLDSELNPKISDFGMARIFGVD 182
Query: 696 ETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSL 755
+ E NT+R+VGTYGYM+PEYA DG FS KSD+FSFGV+LLEIISGK++ G+Y + +L
Sbjct: 183 QQEGNTKRIVGTYGYMAPEYATDGLFSVKSDVFSFGVLLLEIISGKRSRGYYNQNHSQNL 362
Query: 756 LGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDS 815
+G+AWKLW E + L+L+D S+ ++ + +Q + C HV LLCVQ P+DRP MS+V++ML S
Sbjct: 363 IGHAWKLWKEGRPLELIDKSIEDSSSLSQMLHCIHVSLLCVQQNPEDRPGMSSVLLMLVS 542
Query: 816 ETATLPTPKQPTFFTRKDLSSTASSSLQFDSSIVE 850
E LP PKQP FF + + +S+S Q SS E
Sbjct: 543 E-LELPEPKQPGFFGKYSGEADSSTSKQQLSSTNE 644
>TC232399 similar to UP|Q39086 (Q39086) Receptor kinase , partial (23%)
Length = 1167
Score = 258 bits (659), Expect = 8e-69
Identities = 122/222 (54%), Positives = 161/222 (71%)
Frame = +2
Query: 620 NKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEM 679
N +L+ + DP+KS LLDW+ R I+ GI RGLLYLH+DSRL++IHRDLK SN+LL +
Sbjct: 299 NATLNVYFLDPSKSKLLDWRKRCGIIEGIGRGLLYLHRDSRLKIIHRDLKASNVLLYEAL 478
Query: 680 QPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIIS 739
PKISDFG+ARIFGG E +ANT RVVGTYGYMSPEYA+ G FS KSD+FSFGV+++EI+S
Sbjct: 479 NPKISDFGMARIFGGTEDQANTNRVVGTYGYMSPEYAMQGLFSEKSDVFSFGVLVIEIVS 658
Query: 740 GKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDE 799
G++N+ FY LSLLG+AW W E +L ++D + + + +RC H+GLLCVQ+
Sbjct: 659 GRRNSRFYDDDNALSLLGFAWIQWREGNILSVIDPEIYDVTHHKDILRCIHIGLLCVQER 838
Query: 800 PDDRPNMSNVVIMLDSETATLPTPKQPTFFTRKDLSSTASSS 841
DRP M+ V+ ML+SE A LP P QP F +++ + S S
Sbjct: 839 AVDRPTMAAVISMLNSEVAFLPPPDQPAFVQSQNMLNLVSVS 964
Score = 47.0 bits (110), Expect = 4e-05
Identities = 20/28 (71%), Positives = 24/28 (85%)
Frame = +3
Query: 601 LWGYCIKGDEKILIYEYMPNKSLDAFVF 628
L+G C +GDEK+LIYEYM NKSLD F+F
Sbjct: 3 LFGCCAEGDEKMLIYEYMLNKSLDVFIF 86
>TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial (28%)
Length = 1358
Score = 257 bits (656), Expect = 2e-68
Identities = 130/273 (47%), Positives = 188/273 (68%), Gaps = 1/273 (0%)
Frame = +1
Query: 545 ANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGY 604
ANK+G GG+GPVYKG G IAVK+LSS S QG +EF NE+ +I+ LQH +LV+L+G
Sbjct: 4 ANKIGEGGFGPVYKGCFSDGTLIAVKQLSSKSRQGNREFLNEIGMISALQHPHLVKLYGC 183
Query: 605 CIKGDEKILIYEYMPNKSLDAFVFDPTKSAL-LDWQMRFDILLGIARGLLYLHQDSRLRV 663
C++GD+ +L+YEYM N SL +F + + LDW R+ I +GIARGL YLH++SRL++
Sbjct: 184 CVEGDQLLLVYEYMENNSLARALFGAEEHQIKLDWTTRYKICVGIARGLAYLHEESRLKI 363
Query: 664 IHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFST 723
+HRD+K +N+LLD ++ PKISDFGLA++ T +T R+ GT+GYM+PEYA+ G +
Sbjct: 364 VHRDIKATNVLLDQDLNPKISDFGLAKLDEEDNTHIST-RIAGTFGYMAPEYAMHGYLTD 540
Query: 724 KSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNAN 783
K+D++SFG+V LEII+G+ NT Q + + S+L +A L + ++DL+D LG +N
Sbjct: 541 KADVYSFGIVALEIINGRSNTIHRQKEESFSVLEWAHLLREKGDIMDLVDRRLGLEFNKE 720
Query: 784 QFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSE 816
+ + V LLC RP MS+VV ML+ +
Sbjct: 721 EALVMIKVALLCTNVTAALRPTMSSVVSMLEGK 819
>TC210499 weakly similar to UP|Q9LDS6 (Q9LDS6) Serine/threonine kinase-like
protein , partial (27%)
Length = 717
Score = 253 bits (645), Expect = 3e-67
Identities = 121/218 (55%), Positives = 167/218 (76%), Gaps = 2/218 (0%)
Frame = +3
Query: 632 KSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARI 691
+S LLDW+ RF+I+ GI++G+LYLH+ SRL++IHRDLK SNILLD M PKISDFGLAR+
Sbjct: 3 RSMLLDWKKRFNIIEGISQGILYLHKYSRLKIIHRDLKASNILLDENMNPKISDFGLARM 182
Query: 692 FGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKG 751
F +E+ T R+VGTYGYMSPEYA++G FSTKSD++SFGV+LLEI+SG+KNT FY
Sbjct: 183 FMQQESTGTTSRIVGTYGYMSPEYAMEGTFSTKSDVYSFGVLLLEIVSGRKNTSFYDVDH 362
Query: 752 TLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVI 811
L+L+G+AW+LW + + L L+D SL ++++ ++ RC HVGLLCV+ +DRP MSNV+
Sbjct: 363 LLNLIGHAWELWNQGESLQLLDPSLNDSFDPDEVKRCIHVGLLCVEHYANDRPTMSNVIS 542
Query: 812 MLDSETATLPTPKQPTFFT-RKDL-SSTASSSLQFDSS 847
ML +E+A + P++P F+ RK+ T+S L DS+
Sbjct: 543 MLTNESAPVTLPRRPAFYVERKNFDGKTSSKELCVDST 656
>TC230548 weakly similar to UP|Q8LLI4 (Q8LLI4) S-locus receptor-like kinase
RLK14, partial (20%)
Length = 692
Score = 244 bits (624), Expect = 9e-65
Identities = 120/206 (58%), Positives = 152/206 (73%)
Frame = +3
Query: 641 RFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEAN 700
RF I+ GIARGLLYLHQDSRLR+IHRDLK SN+LLD EM PKISDFGLAR+ GG + E
Sbjct: 3 RFGIINGIARGLLYLHQDSRLRIIHRDLKASNVLLDNEMNPKISDFGLARMCGGDQIEGE 182
Query: 701 TQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAW 760
T RVVGTYGYM+PEYA DG FS KSD+FSFGV+LLEI+SGKKN+ + +L+G+AW
Sbjct: 183 TSRVVGTYGYMAPEYAFDGIFSIKSDVFSFGVLLLEIVSGKKNSRLFYPNDYNNLIGHAW 362
Query: 761 KLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATL 820
LW E + +D SL ++ + +RC H+GLLCVQ P+DRPNM++VV++L +E A L
Sbjct: 363 MLWKEGNPMQFIDSSLEDSCILYEALRCIHIGLLCVQHHPNDRPNMASVVVLLSNENA-L 539
Query: 821 PTPKQPTFFTRKDLSSTASSSLQFDS 846
P PK P++ ++ + SSS F S
Sbjct: 540 PLPKDPSYLSKDISTERESSSKNFTS 617
>TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partial (25%)
Length = 1190
Score = 244 bits (624), Expect = 9e-65
Identities = 126/265 (47%), Positives = 179/265 (67%)
Frame = +3
Query: 566 EIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDA 625
EIAVK+LS S QG +F E+ I+ +QHRNLV+L+G CI+G +++L+YEY+ NKSLD
Sbjct: 3 EIAVKQLSVGSHQGKSQFITEIATISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQ 182
Query: 626 FVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISD 685
+F K L+W R+DI LG+ARGL YLH++SRLR++HRD+K SNILLD E+ PKISD
Sbjct: 183 ALFG--KCLTLNWSTRYDICLGVARGLTYLHEESRLRIVHRDVKASNILLDYELIPKISD 356
Query: 686 FGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTG 745
FGLA+++ K+T +T V GT GY++PEYA+ G + K+D+FSFGVV LE++SG+ N+
Sbjct: 357 FGLAKLYDDKKTHIST-GVAGTIGYLAPEYAMRGHLTEKADVFSFGVVALELVSGRPNSD 533
Query: 746 FYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPN 805
+ LL +AW+L +N ++DL+D L E +N + R + LLC Q P RP+
Sbjct: 534 SSLEGEKVYLLEWAWQLHEKNCIIDLVDDRLSE-FNEEEVKRVVGIALLCTQTSPTLRPS 710
Query: 806 MSNVVIMLDSETATLPTPKQPTFFT 830
MS VV ML + +P + +
Sbjct: 711 MSRVVAMLSGDIEVSTVTSKPGYLS 785
>TC210344 similar to UP|Q9AVE0 (Q9AVE0) SRKb, partial (21%)
Length = 870
Score = 243 bits (620), Expect = 3e-64
Identities = 115/188 (61%), Positives = 148/188 (78%)
Frame = +2
Query: 641 RFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEAN 700
R DI+ GIARG+LYLH+DSRL++IHRDLK SN+LLD +M PKISDFG+ARIF G E EAN
Sbjct: 8 RLDIINGIARGILYLHEDSRLKIIHRDLKASNVLLDYDMNPKISDFGMARIFAGSEGEAN 187
Query: 701 TQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAW 760
T +VGTYGYM+PEYA++G +S KSD+F FGV+LLEII+GK+N GFY T SLL YAW
Sbjct: 188 TATIVGTYGYMAPEYAMEGLYSIKSDVFGFGVLLLEIITGKRNAGFYHSNKTPSLLSYAW 367
Query: 761 KLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATL 820
LW E K L+L+D ++ ++F+R H+GLLCVQ++ DRP MS+VV+ML +E+ATL
Sbjct: 368 HLWNEGKGLELIDPMSVDSCPGDEFLRYMHIGLLCVQEDAYDRPTMSSVVLMLKNESATL 547
Query: 821 PTPKQPTF 828
P++P F
Sbjct: 548 GQPERPPF 571
>TC208545 weakly similar to UP|Q9S972 (Q9S972) ARK2 product/receptor-like
serine/threonine protein kinase ARK2, partial (20%)
Length = 1253
Score = 239 bits (611), Expect = 3e-63
Identities = 116/216 (53%), Positives = 159/216 (72%)
Frame = +2
Query: 629 DPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGL 688
D T+S LLDW+ RF+I+ GI++GLLYLH+ SRL+VIHRDLK SNILLD M PKISDFGL
Sbjct: 326 DGTRSKLLDWKKRFNIIEGISQGLLYLHKYSRLKVIHRDLKASNILLDENMNPKISDFGL 505
Query: 689 ARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQ 748
AR+F +E+ NT R+VGTYGYMSPEYA++G FS KSD++SFGV+LLEI+SG++NT FY
Sbjct: 506 ARMFTRQESTTNTSRIVGTYGYMSPEYAMEGVFSVKSDVYSFGVLLLEIVSGRRNTSFYD 685
Query: 749 YKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSN 808
L+L+G+AW+LW E L L+D SL E+ + ++ RC H+GLLCV+ ++RP MS
Sbjct: 686 GDRFLNLIGHAWELWNEGACLKLIDPSLTESPDLDEVQRCIHIGLLCVEQNANNRPLMSQ 865
Query: 809 VVIMLDSETATLPTPKQPTFFTRKDLSSTASSSLQF 844
++ ML ++ + P++P F+ + SS +F
Sbjct: 866 IISMLSNKN-PITLPQRPAFYFGSETFDGIISSTEF 970
>BU091755
Length = 424
Score = 239 bits (609), Expect = 5e-63
Identities = 113/141 (80%), Positives = 132/141 (93%)
Frame = +1
Query: 543 SDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLW 602
+D+ KLGRGGYGPVYKG GG++IAVKRLSSVS+QG++EFKNEV+LIAKLQHRNLVRL
Sbjct: 1 TDSYKLGRGGYGPVYKGTFPGGQDIAVKRLSSVSTQGLEEFKNEVILIAKLQHRNLVRLR 180
Query: 603 GYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLR 662
GYCIKGDEKIL+YEYMPNKSLD+F+FD T+++LLDW +RF+I++GIARG+LYLHQDSRLR
Sbjct: 181 GYCIKGDEKILLYEYMPNKSLDSFIFDRTRTSLLDWPIRFEIIVGIARGMLYLHQDSRLR 360
Query: 663 VIHRDLKTSNILLDGEMQPKI 683
VIHRDLKTSNILLD EM PKI
Sbjct: 361 VIHRDLKTSNILLDEEMNPKI 423
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,353,540
Number of Sequences: 63676
Number of extensions: 619141
Number of successful extensions: 5259
Number of sequences better than 10.0: 992
Number of HSP's better than 10.0 without gapping: 4488
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4554
length of query: 852
length of database: 12,639,632
effective HSP length: 105
effective length of query: 747
effective length of database: 5,953,652
effective search space: 4447378044
effective search space used: 4447378044
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0203.5


