

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.3
(475 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
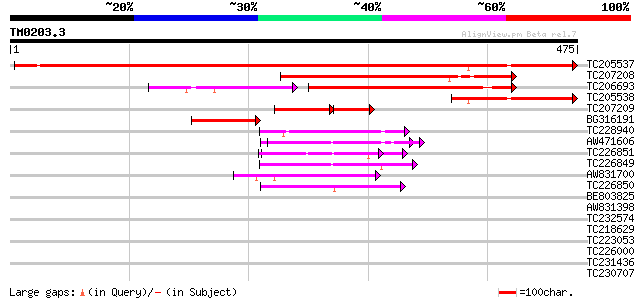
Score E
Sequences producing significant alignments: (bits) Value
TC205537 similar to GB|AAP37774.1|30725504|BT008415 At4g03260 {A... 722 0.0
TC207208 similar to UP|Q9M9E4 (Q9M9E4) F3F9.22, partial (38%) 200 1e-51
TC206693 similar to UP|Q9M9E4 (Q9M9E4) F3F9.22, partial (57%) 150 2e-51
TC205538 140 9e-34
TC207209 similar to UP|Q9M9E4 (Q9M9E4) F3F9.22, partial (29%) 72 2e-23
BG316191 97 2e-20
TC228940 similar to UP|Q8RX50 (Q8RX50) PSR9 (Fragment), partial ... 57 1e-08
AW471606 52 8e-07
TC226851 similar to UP|Q84WJ9 (Q84WJ9) At5g19680, partial (47%) 52 8e-07
TC226849 similar to UP|Q84WJ9 (Q84WJ9) At5g19680, partial (68%) 51 1e-06
AW831700 weakly similar to GP|21703129|gb AT3g15410/MJK13_7 {Ara... 45 5e-05
TC226850 similar to UP|Q84WJ9 (Q84WJ9) At5g19680, partial (40%) 42 8e-04
BE803825 weakly similar to GP|14190425|gb| At1g68400/T2E12_5 {Ar... 40 0.003
AW831398 weakly similar to GP|10241587|emb thymidine kinase (LTK... 38 0.009
TC232574 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2,... 38 0.009
TC218629 38 0.011
TC223053 36 0.043
TC226000 similar to UP|O49469 (O49469) TMV resistance protein N-... 35 0.056
TC231436 weakly similar to UP|Q8RX50 (Q8RX50) PSR9 (Fragment), p... 35 0.073
TC230707 similar to UP|Q12937 (Q12937) BS-84, partial (4%) 35 0.073
>TC205537 similar to GB|AAP37774.1|30725504|BT008415 At4g03260 {Arabidopsis
thaliana;} , partial (33%)
Length = 1678
Score = 722 bits (1863), Expect = 0.0
Identities = 374/476 (78%), Positives = 408/476 (85%), Gaps = 5/476 (1%)
Frame = +1
Query: 5 SLPNIKASTLSPENHAFKHPSSKSRSSDDLHALGMRKKEGFINESEDQIREGQEREDNIG 64
SLPNI+AS LS E AFKH + SRSSDDLHAL M +KE FINE +DQIR QERE+++G
Sbjct: 7 SLPNIRASILSSERDAFKH--ALSRSSDDLHALRMWQKEEFINEFDDQIRADQERENDMG 180
Query: 65 KTEDNHMGEYFDDGFDSYLLSDLEKDWVMPTTDDISEVKTLQGDNSVDCFGEFPNKDFKV 124
K ED M +FDDGFDSYLLS KDWVMP TDD S VKTLQGD+ VDC GEFP KDFK+
Sbjct: 181 KPEDGRMDSFFDDGFDSYLLSGSAKDWVMPITDDTSNVKTLQGDSLVDCAGEFPKKDFKI 360
Query: 125 KRIEDWVVDLQHCGPPVEEINE-LPESVDPVVDINTINGVTAAGVNHKITPGMEAAKRYI 183
KRIEDWV+ LQHCGPP+EE NE LPE ++P++D+NT+NGVTAA V++K+TPGMEAAKRYI
Sbjct: 361 KRIEDWVIGLQHCGPPLEETNEDLPEVIEPLIDVNTVNGVTAASVDNKVTPGMEAAKRYI 540
Query: 184 SSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNN 243
SSL ANA+AAQL NHGLVV+PFLSAFVSLKVLNLAGN+IVRITAGALPRGLH+LNLSRN
Sbjct: 541 SSLGANATAAQLGNHGLVVIPFLSAFVSLKVLNLAGNAIVRITAGALPRGLHALNLSRNK 720
Query: 244 ISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSI 303
IS+IEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKI EVEGLHRLLKLSI
Sbjct: 721 ISTIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKISEVEGLHRLLKLSI 900
Query: 304 LDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQGLLPHLVYYNRQ 363
LDL FNKISTAKCLGQLAANYN+LQAINLDGNP QKNVGDE +KKYLQGLLPHLVYYNRQ
Sbjct: 901 LDLSFNKISTAKCLGQLAANYNTLQAINLDGNPAQKNVGDEHMKKYLQGLLPHLVYYNRQ 1080
Query: 364 AMKVSTLKDGADRLVRLGTN----DRSLRVDRKTTRKGSHGVAAARRPPSTSTHSHRSQT 419
MKVS+LKDGA+R VRLG N DR LR DRKTTRKGS GVAA RRP TSTH+ RS
Sbjct: 1081PMKVSSLKDGAERSVRLGMNSHQFDRGLRADRKTTRKGSQGVAATRRPSITSTHARRS-- 1254
Query: 420 VESPKLSKGKQSHLPPIRTKVSTQSRHYLDAQSKVLNLTSGHSMRKSRSEGTLGAL 475
V+SPKLSKGKQ LPP RTKVSTQSR++ DA K LNL S SMRKSRSEG G L
Sbjct: 1255VDSPKLSKGKQPLLPPTRTKVSTQSRYHFDAPDKALNLVSELSMRKSRSEGNFGVL 1422
>TC207208 similar to UP|Q9M9E4 (Q9M9E4) F3F9.22, partial (38%)
Length = 967
Score = 200 bits (508), Expect = 1e-51
Identities = 108/201 (53%), Positives = 141/201 (69%), Gaps = 4/201 (1%)
Frame = +3
Query: 228 GALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGN 287
G LP+G+ +LNLS+N IS++EGLREL +LR+LDLSYNRI RIG GL+SC+ +KELYL GN
Sbjct: 3 GVLPKGIQTLNLSKNKISTLEGLRELAKLRILDLSYNRISRIGQGLSSCTLIKELYLVGN 182
Query: 288 KIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLK 347
KI +VEGLHRLLKL++LDL FNKI+TAK LGQL AN+NSL+A+NL GN Q N+ D+QL
Sbjct: 183 KISDVEGLHRLLKLTVLDLSFNKITTAKALGQLVANFNSLKALNLLGNSIQSNISDDQLS 362
Query: 348 KYLQGLLPHLVYYNRQAMKV----STLKDGADRLVRLGTNDRSLRVDRKTTRKGSHGVAA 403
K + GLLP +VY N+Q +K L D R LG++ RS +R++ R+ HG +
Sbjct: 363 KAVCGLLPKMVYLNKQPVKAHRTRGILSDSVAR-AALGSSTRS--CNRRSIRRVGHGGSN 533
Query: 404 ARRPPSTSTHSHRSQTVESPK 424
R ST + T + K
Sbjct: 534 LSRGNRRSTSVSQKSTNRTRK 596
>TC206693 similar to UP|Q9M9E4 (Q9M9E4) F3F9.22, partial (57%)
Length = 1947
Score = 150 bits (380), Expect(2) = 2e-51
Identities = 89/177 (50%), Positives = 117/177 (65%), Gaps = 3/177 (1%)
Frame = +3
Query: 251 RELTRLRV-LDLSYNRILR-IGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRF 308
RELTRLRV L NR L+ + GL++C+ +KELYLAGNKI +VEGLHRLLKL++LDL F
Sbjct: 1065 RELTRLRVPLT*VXNRXLKKLDQGLSNCTLVKELYLAGNKISDVEGLHRLLKLTVLDLSF 1244
Query: 309 NKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQGLLPHLVYYNRQAMKVS 368
NKI+T K LGQL ANYNSLQA+NL GNP Q N+ D+QL+K + GLLP LVY N+Q++K
Sbjct: 1245 NKIATTKALGQLVANYNSLQALNLLGNPIQSNISDDQLRKVVCGLLPKLVYLNKQSIKPQ 1424
Query: 369 TLKD-GADRLVRLGTNDRSLRVDRKTTRKGSHGVAAARRPPSTSTHSHRSQTVESPK 424
++ D + + + S R+ +KG + S+S+ HRS S K
Sbjct: 1425 RGREILTDSVAKAALGNSSRNSYRRALKKG------GGQGGSSSSSVHRSSASVSQK 1577
Score = 70.5 bits (171), Expect(2) = 2e-51
Identities = 46/136 (33%), Positives = 74/136 (53%), Gaps = 11/136 (8%)
Frame = +1
Query: 117 FPNKDFKVKRIEDWVVDLQHCGPPVEE-INE-------LPESVDPVVDINTINGVTAAGV 168
F + R+++WV DL+ PP+E+ N+ P S D D ++ TA +
Sbjct: 631 FSTESSSYSRVDEWVKDLEIQQPPLEDDFNDDNIGSIAFPPSPD---DGRSMARSTAQLI 801
Query: 169 NH---KITPGMEAAKRYISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRI 225
H ++ + A + SL ++AA +++ G+ +P LS F SL+ +NL+ N IV I
Sbjct: 802 QHPDANLSKEILNANSVVQSLNPASTAAHISSIGIKAIPSLSHFFSLRCVNLSNNLIVHI 981
Query: 226 TAGALPRGLHSLNLSR 241
T G LP+G+H+LNLSR
Sbjct: 982 TPGFLPKGIHTLNLSR 1029
>TC205538
Length = 601
Score = 140 bits (354), Expect = 9e-34
Identities = 77/109 (70%), Positives = 85/109 (77%), Gaps = 4/109 (3%)
Frame = +1
Query: 371 KDGADRLVRLGTN----DRSLRVDRKTTRKGSHGVAAARRPPSTSTHSHRSQTVESPKLS 426
KDGA+R VRLG N DRSLR DRKTTRKGS GVAA RRP + STH+ RS V+SPKLS
Sbjct: 1 KDGAERSVRLGMNSHQFDRSLRADRKTTRKGSQGVAATRRPSTASTHARRS--VDSPKLS 174
Query: 427 KGKQSHLPPIRTKVSTQSRHYLDAQSKVLNLTSGHSMRKSRSEGTLGAL 475
KGKQ+ LPP RTKVSTQSR++ D+ SKVLNL S SMRKSRSEG G L
Sbjct: 175 KGKQTLLPPTRTKVSTQSRYHFDSPSKVLNLVSERSMRKSRSEGNFGVL 321
>TC207209 similar to UP|Q9M9E4 (Q9M9E4) F3F9.22, partial (29%)
Length = 842
Score = 72.0 bits (175), Expect(2) = 2e-23
Identities = 33/50 (66%), Positives = 42/50 (84%)
Frame = +1
Query: 223 VRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHG 272
V I+ G LP+G+ +LNLS+N IS++EGLREL +LR+LDLSYNRI RIG G
Sbjct: 493 VHISPGVLPKGIQTLNLSKNKISTLEGLRELAKLRILDLSYNRISRIGQG 642
Score = 55.1 bits (131), Expect(2) = 2e-23
Identities = 24/34 (70%), Positives = 30/34 (87%)
Frame = +3
Query: 272 GLASCSSLKELYLAGNKIGEVEGLHRLLKLSILD 305
GL+SC+ +KELYL GNKI +VEGLHRLLK ++LD
Sbjct: 741 GLSSCTLIKELYLVGNKISDVEGLHRLLKATVLD 842
Score = 37.0 bits (84), Expect = 0.019
Identities = 18/34 (52%), Positives = 24/34 (69%)
Frame = +1
Query: 279 LKELYLAGNKIGEVEGLHRLLKLSILDLRFNKIS 312
++ L L+ NKI +EGL L KL ILDL +N+IS
Sbjct: 526 IQTLNLSKNKISTLEGLRELAKLRILDLSYNRIS 627
>BG316191
Length = 177
Score = 96.7 bits (239), Expect = 2e-20
Identities = 46/58 (79%), Positives = 54/58 (92%)
Frame = +2
Query: 153 PVVDINTINGVTAAGVNHKITPGMEAAKRYISSLTANASAAQLANHGLVVVPFLSAFV 210
P+VD+NT+NGVTAA V++K+TP MEAAKRYISSL ANA+AAQL NHGLVV+PFLSAFV
Sbjct: 2 PLVDVNTVNGVTAASVDNKVTPRMEAAKRYISSLGANATAAQLGNHGLVVIPFLSAFV 175
>TC228940 similar to UP|Q8RX50 (Q8RX50) PSR9 (Fragment), partial (46%)
Length = 1110
Score = 57.4 bits (137), Expect = 1e-08
Identities = 42/130 (32%), Positives = 74/130 (56%), Gaps = 4/130 (3%)
Frame = +1
Query: 210 VSLKVLNLAGNSIVRITAG--ALPRGLHSLNLSRNNISSI-EGLRELTRLRVLDLSYNRI 266
+SL L+L GN + + A L R L L+LS N +S++ + + L RL++L++ N I
Sbjct: 34 LSLLYLDLRGNQLTLLPASFSRLVR-LEELDLSSNQLSALPDSIGSLVRLKILNVETNDI 210
Query: 267 LRIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKISTAKCLGQLAANYN 325
+ H + SCSSL+EL + N++ + E + ++ L IL +R+N I K L ++
Sbjct: 211 EELPHSVGSCSSLRELRVDYNRLKALPEAVGKIQSLEILSVRYNNI---KQLPTTMSSLT 381
Query: 326 SLQAINLDGN 335
+L+ +N+ N
Sbjct: 382 NLKELNVSFN 411
Score = 38.5 bits (88), Expect = 0.007
Identities = 37/149 (24%), Positives = 77/149 (50%), Gaps = 8/149 (5%)
Frame = +1
Query: 195 LANHGLVVVP-FLSAFVSLKVLNLAGNSIVRI--TAGALPRGLHSLNLSRNNISSI-EGL 250
L+++ L +P + + V LK+LN+ N I + + G+ L L + N + ++ E +
Sbjct: 124 LSSNQLSALPDSIGSLVRLKILNVETNDIEELPHSVGSCS-SLRELRVDYNRLKALPEAV 300
Query: 251 RELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV-EGL---HRLLKLSILDL 306
++ L +L + YN I ++ ++S ++LKEL ++ N++ V E L L+K++I
Sbjct: 301 GKIQSLEILSVRYNNIKQLPTTMSSLTNLKELNVSFNEL*SVPESLCFATSLVKMNI--- 471
Query: 307 RFNKISTAKCLGQLAANYNSLQAINLDGN 335
N + + L + N L+ +++ N
Sbjct: 472 -GNNFADMRSLPRSIGNLELLEELDISNN 555
>AW471606
Length = 432
Score = 51.6 bits (122), Expect = 8e-07
Identities = 40/131 (30%), Positives = 67/131 (50%)
Frame = +1
Query: 217 LAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASC 276
L+ + +++ A P + +L L+ +S I L L LDL N + + GL SC
Sbjct: 7 LSSDQVLKDNNAADPTSVTTLYLTHKALSDITFLANFINLEKLDLKLNNLTSL-EGLRSC 183
Query: 277 SSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNP 336
+LK L + NK+ +EG+ L KL++L+ NK+ K + Q+ + SL+A+ L+ N
Sbjct: 184 VNLKWLSVVENKLESLEGIQGLTKLTVLNAGKNKL---KSIDQV-MSVVSLRALILNENE 351
Query: 337 CQKNVGDEQLK 347
+QLK
Sbjct: 352 ISSICKLDQLK 384
Score = 46.6 bits (109), Expect = 2e-05
Identities = 38/129 (29%), Positives = 62/129 (47%)
Frame = +1
Query: 211 SLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIG 270
S+ L L ++ IT A L L+L NN++S+EGLR L+ L + N++ +
Sbjct: 55 SVTTLYLTHKALSDITFLANFINLEKLDLKLNNLTSLEGLRSCVNLKWLSVVENKLESL- 231
Query: 271 HGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAI 330
G+ + L L NK+ ++ + ++ L L L N+IS+ L QL L +
Sbjct: 232 EGIQGLTKLTVLNAGKNKLKSIDQVMSVVSLRALILNENEISSICKLDQL----KDLNTL 399
Query: 331 NLDGNPCQK 339
L NP +K
Sbjct: 400 VLSKNPIRK 426
Score = 32.7 bits (73), Expect = 0.36
Identities = 24/65 (36%), Positives = 30/65 (45%)
Frame = +1
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNR 265
+ L VLN N + I L +L L+ N ISSI L +L L L LS N
Sbjct: 238 IQGLTKLTVLNAGKNKLKSIDQVMSVVSLRALILNENEISSICKLDQLKDLNTLVLSKNP 417
Query: 266 ILRIG 270
I +IG
Sbjct: 418 IRKIG 432
>TC226851 similar to UP|Q84WJ9 (Q84WJ9) At5g19680, partial (47%)
Length = 882
Score = 51.6 bits (122), Expect = 8e-07
Identities = 38/128 (29%), Positives = 66/128 (50%), Gaps = 6/128 (4%)
Frame = +1
Query: 212 LKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGH 271
+K ++ N + +T L L LS N I+ +EGL L LRVLD+S N+I +
Sbjct: 148 IKKISFQSNRLTSMTGFEGCVALEELYLSHNGIAKMEGLSSLVNLRVLDVSSNKITSV-D 324
Query: 272 GLASCSSLKELYLAGNKIGEVEGLHRLL-----KLSILDLRFNKIS-TAKCLGQLAANYN 325
+ + + L++L+L N+I +EG+ + KL+ + L+ N + T G L +
Sbjct: 325 DIVNLTKLEDLWLNDNQIASLEGIAEAVSGSKEKLTTIYLKNNPCAKTPNYTGILREIFP 504
Query: 326 SLQAINLD 333
++Q I+ D
Sbjct: 505 NIQQIDSD 528
Score = 47.8 bits (112), Expect = 1e-05
Identities = 33/105 (31%), Positives = 54/105 (51%)
Frame = +1
Query: 209 FVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILR 268
F L++L L N + + L L L +N I + L L ++ + NR+
Sbjct: 10 FHQLQILELGSNKLRVMENLQSLENLQELWLGQNRIKVVN-LCGLKCIKKISFQSNRLTS 186
Query: 269 IGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKIST 313
+ G C +L+ELYL+ N I ++EGL L+ L +LD+ NKI++
Sbjct: 187 MT-GFEGCVALEELYLSHNGIAKMEGLSSLVNLRVLDVSSNKITS 318
>TC226849 similar to UP|Q84WJ9 (Q84WJ9) At5g19680, partial (68%)
Length = 796
Score = 50.8 bits (120), Expect = 1e-06
Identities = 36/139 (25%), Positives = 72/139 (50%), Gaps = 7/139 (5%)
Frame = +1
Query: 210 VSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRI 269
+SL+ ++ +++ +++ L L L N +I + +L V D+++N I +
Sbjct: 220 LSLRQNLISDAAVLPLSSWNALSSLEELVLRDNQFKNIPDVSVFKKLLVFDVAFNEISSL 399
Query: 270 GHGLASCS-SLKELYLAGNKIGEVEGLHRLLKLSILDLRFNK------ISTAKCLGQLAA 322
HGL+ S +LKELY++ N++ +E + +L +L+L NK + + + L +L
Sbjct: 400 -HGLSRVSDTLKELYVSKNEVAMIEEIEHFHQLQLLELGSNKLRVMENLQSLENLQELWL 576
Query: 323 NYNSLQAINLDGNPCQKNV 341
N ++ +NL G C K +
Sbjct: 577 GRNRIKVVNLCGLKCIKKI 633
Score = 40.4 bits (93), Expect = 0.002
Identities = 31/119 (26%), Positives = 57/119 (47%)
Frame = +1
Query: 183 ISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRN 242
+S ++ ++ + + ++ + F L++L L N + + L L L RN
Sbjct: 406 LSRVSDTLKELYVSKNEVAMIEEIEHFHQLQLLELGSNKLRVMENLQSLENLQELWLGRN 585
Query: 243 NISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKL 301
I + L L ++ + L NR+ + G C +L+ELYL+ N I ++EGL L+ L
Sbjct: 586 RIKVVN-LCGLKCIKKISLQSNRLTSM-MGFDGCVALEELYLSHNGIAKMEGLSSLVNL 756
Score = 35.4 bits (80), Expect = 0.056
Identities = 36/130 (27%), Positives = 61/130 (46%), Gaps = 6/130 (4%)
Frame = +1
Query: 207 SAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEGLRELT-RLRVLDLSYNR 265
+A SL+ L L N I ++ + L +++ N ISS+ GL ++ L+ L +S N
Sbjct: 277 NALSSLEELVLRDNQFKNIPDVSVFKKLLVFDVAFNEISSLHGLSRVSDTLKELYVSKNE 456
Query: 266 ILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTA-----KCLGQL 320
+ I + L+ L L NK+ +E L L L L L N+I KC+ ++
Sbjct: 457 VAMIEE-IEHFHQLQLLELGSNKLRVMENLQSLENLQELWLGRNRIKVVNLCGLKCIKKI 633
Query: 321 AANYNSLQAI 330
+ N L ++
Sbjct: 634 SLQSNRLTSM 663
>AW831700 weakly similar to GP|21703129|gb AT3g15410/MJK13_7 {Arabidopsis
thaliana}, partial (23%)
Length = 423
Score = 45.4 bits (106), Expect = 5e-05
Identities = 40/128 (31%), Positives = 64/128 (49%), Gaps = 5/128 (3%)
Frame = +1
Query: 188 ANASAAQLANHGLVVVPF--LSAFVSLKVLNLAGN--SIVRITAGALPRGLHSLNLSRNN 243
++ S +L N+ L +P L++LNL+GN S++ I + L L L +
Sbjct: 19 SSLSCLKLDNNPLKQIPLDGFEVVPXLQILNLSGNAXSLLNIPTFSNLPYLQKLYLKKMK 198
Query: 244 ISSIEG-LRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLS 302
+S + + L L++L+LS N + I L +SLKEL L+ N I + LLK +
Sbjct: 199 LSKVPSDIVGLQXLQILNLSQNSLQSIPVKLKDLTSLKELDLSNNNISVLLPKLSLLKPN 378
Query: 303 ILDLRFNK 310
+ LR NK
Sbjct: 379 LQTLRLNK 402
>TC226850 similar to UP|Q84WJ9 (Q84WJ9) At5g19680, partial (40%)
Length = 507
Score = 41.6 bits (96), Expect = 8e-04
Identities = 33/126 (26%), Positives = 61/126 (48%), Gaps = 5/126 (3%)
Frame = +2
Query: 211 SLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEG-LRELTRLRVLDLSYNRILRI 269
S +L+L+ + + + LP L L+L+ N +S+++ + L+ L+ L L N I
Sbjct: 104 SSTLLDLSSYQLHDLDSVELPPSLTELDLTANRLSTLDPRIGNLSHLKKLSLRQNLITDA 283
Query: 270 G----HGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAKCLGQLAANYN 325
+ SSL+EL L N++ + + KL + D+ FN+IS+ L +++
Sbjct: 284 AVLPLSSWNAISSLEELVLRDNQLKNIPDVTVFKKLLVFDVAFNEISSLHGLSRVSDTLK 463
Query: 326 SLQAIN 331
L N
Sbjct: 464 ELYVSN 481
>BE803825 weakly similar to GP|14190425|gb| At1g68400/T2E12_5 {Arabidopsis
thaliana}, partial (5%)
Length = 409
Score = 39.7 bits (91), Expect = 0.003
Identities = 31/110 (28%), Positives = 52/110 (47%), Gaps = 12/110 (10%)
Frame = +2
Query: 234 LHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAG-NKIGEV 292
+ SLNLS N ++ L L++LDLS+N ++ + G + S+L+ + L+ N V
Sbjct: 38 IQSLNLSYNRFTNSIQLSGFKNLKILDLSHNNLVTLPSGFQNLSNLQHIDLSSCNLQSNV 217
Query: 293 EGLHRLLKLSILDLR-----------FNKISTAKCLGQLAANYNSLQAIN 331
+ + L L LDL F ++T K L N+ S ++N
Sbjct: 218 KPISALHSLHYLDLSNNTFTGNFPYDFPPLTTLKFLNISFNNFTSAISVN 367
>AW831398 weakly similar to GP|10241587|emb thymidine kinase (LTK) {Fagus
sylvatica}, partial (55%)
Length = 432
Score = 38.1 bits (87), Expect = 0.009
Identities = 37/112 (33%), Positives = 56/112 (49%), Gaps = 5/112 (4%)
Frame = +1
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISS--IEGLRELTRLRVLDLSY 263
LS +SL+ L+L+ N I LP L SLN +RNN+S + + L L+LS
Sbjct: 55 LSDLMSLRELDLSDNKIHDTIPYQLPPNLTSLNFARNNLSGNLPYSISAMVSLNYLNLSN 234
Query: 264 NRI-LRIGHGLASCSSLKELYLAGNKI-GEV-EGLHRLLKLSILDLRFNKIS 312
N + + +G AS L L L+ N G++ L LS L L+ N+++
Sbjct: 235 NALSMTVGDIFASLQDLGTLDLSFNNFSGDLPPSFVALANLSSLFLQKNQLT 390
>TC232574 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2, partial
(10%)
Length = 852
Score = 38.1 bits (87), Expect = 0.009
Identities = 32/89 (35%), Positives = 47/89 (51%), Gaps = 9/89 (10%)
Frame = +3
Query: 210 VSLKVLNLAGNSIVRITAGALPR------GLHSLNLSRNNISS--IEGLRELTRLRVLDL 261
+ LK ++L+ N++ G +P+ GL SLNLSRNN+S + L L LDL
Sbjct: 21 LELKSIDLSSNNLT----GEIPKEVGYLLGLVSLNLSRNNLSGEIPSRIGNLRSLESLDL 188
Query: 262 SYNRI-LRIGHGLASCSSLKELYLAGNKI 289
S N I RI L+ L++L L+ N +
Sbjct: 189 SRNHISRRIPSSLSEIDDLQKLDLSHNSL 275
>TC218629
Length = 1306
Score = 37.7 bits (86), Expect = 0.011
Identities = 27/94 (28%), Positives = 41/94 (42%)
Frame = +3
Query: 243 NISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLS 302
NI + L LR+L + Y C L+E+Y + N+ E G +KL
Sbjct: 510 NIIMPSTIANLPNLRILSIKY------------CFELEEIYGSNNESDEPLGEIAFMKLE 653
Query: 303 ILDLRFNKISTAKCLGQLAANYNSLQAINLDGNP 336
L L+ + T+ C G + N+ SLQ + L P
Sbjct: 654 ELTLKSLRSLTSFCQGSYSFNFPSLQKVQLKDCP 755
>TC223053
Length = 1024
Score = 35.8 bits (81), Expect = 0.043
Identities = 30/77 (38%), Positives = 40/77 (50%), Gaps = 5/77 (6%)
Frame = +2
Query: 234 LHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKI-GEV 292
L SLNL I+ L LT LR LDL NR+ L +C+SL+ YL+ N GE+
Sbjct: 650 LPSLNLR----GPIDSLSTLTYLRFLDLHENRLNGTVSPLLNCTSLEIKYLSRNDFSGEI 817
Query: 293 E----GLHRLLKLSILD 305
+ L LL++ I D
Sbjct: 818 QPALSSLRLLLRVDISD 868
>TC226000 similar to UP|O49469 (O49469) TMV resistance protein N-like,
partial (4%)
Length = 938
Score = 35.4 bits (80), Expect = 0.056
Identities = 24/79 (30%), Positives = 38/79 (47%)
Frame = +3
Query: 256 LRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKISTAK 315
+R LDLS+ ++ I + S L+ L L+GN + L +L KL L L+ K K
Sbjct: 558 MRELDLSFCNLVEIPDAIGIMSCLERLDLSGNNFATLPNLKKLSKLVCLKLQHCK--QLK 731
Query: 316 CLGQLAANYNSLQAINLDG 334
L +L + N+ + G
Sbjct: 732 SLPELPSRINNFDGLRQAG 788
>TC231436 weakly similar to UP|Q8RX50 (Q8RX50) PSR9 (Fragment), partial (35%)
Length = 836
Score = 35.0 bits (79), Expect = 0.073
Identities = 40/141 (28%), Positives = 68/141 (47%), Gaps = 4/141 (2%)
Frame = +1
Query: 211 SLKVLNLAGNSIVRI--TAGALPRGLHSLNLSRNNISSI-EGLRELTRLRVLDLSYNRIL 267
SLK LN+ N + + T G L L L N + ++ E + +L L +L L YNR+
Sbjct: 4 SLKRLNVETNELEELPYTIGNCS-SLSVLKLDLNQLKALPEAIGKLECLEILTLHYNRVK 180
Query: 268 RIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKISTAKCLGQLAANYNS 326
R+ + + +LKEL ++ N++ V E L L L+L N L L A+ +
Sbjct: 181 RLPSTMDNLCNLKELDVSFNELEFVPESLCFATNLKKLNLGKNFAD----LRALPASIGN 348
Query: 327 LQAINLDGNPCQKNVGDEQLK 347
L+ + + ++ D+Q+K
Sbjct: 349 LEMLE------ELDISDDQIK 393
>TC230707 similar to UP|Q12937 (Q12937) BS-84, partial (4%)
Length = 990
Score = 35.0 bits (79), Expect = 0.073
Identities = 38/142 (26%), Positives = 67/142 (46%), Gaps = 12/142 (8%)
Frame = +3
Query: 183 ISSLTANASAAQLANHGLVVVPF-LSAFVSLKVLNLAGNSIVRITAGALP------RGLH 235
IS LT + N+ +P +S +L+ L L NS +G +P + L
Sbjct: 3 ISQLTQLTTLDLADNNFFGPIPSSISLLSNLQTLTLRSNSF----SGTIPPSITTLKSLL 170
Query: 236 SLNLSRNNISSI--EGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIG--- 290
SL+L+ N++S + LT LR LDLS+N++ G S+L EL + N +
Sbjct: 171 SLDLAHNSLSGYLPNSMNSLTTLRRLDLSFNKL--TGSIPKLPSNLLELAIKANSLSGPL 344
Query: 291 EVEGLHRLLKLSILDLRFNKIS 312
+ + + +L +++L N ++
Sbjct: 345 QKQSFEGMNQLEVVELSENALT 410
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.132 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,669,745
Number of Sequences: 63676
Number of extensions: 239446
Number of successful extensions: 1066
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 1028
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1049
length of query: 475
length of database: 12,639,632
effective HSP length: 101
effective length of query: 374
effective length of database: 6,208,356
effective search space: 2321925144
effective search space used: 2321925144
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0203.3


