

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0199.9
(56 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
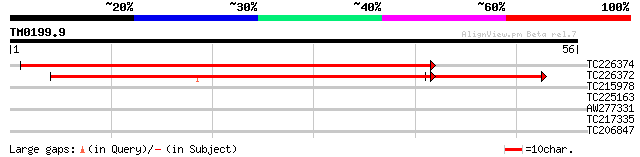
TC226374 similar to UP|VATC_ARATH (Q9SDS7) Vacuolar ATP synthase... 74 2e-14
TC226372 similar to UP|VATC_ARATH (Q9SDS7) Vacuolar ATP synthase... 71 1e-13
TC215978 weakly similar to UP|Q8S1K2 (Q8S1K2) PDI-like protein, ... 26 3.7
TC225163 emb|X02559.1|MIOBRN26 Oenothera berteriana mitochondria... 25 6.3
AW277331 25 6.3
TC217335 similar to UP|GIC2_YEAST (Q06648) GTPase-interacting co... 25 8.3
TC206847 similar to GB|AAF76441.1|8569096|F2J10 ESTs gb|AI994059... 25 8.3
>TC226374 similar to UP|VATC_ARATH (Q9SDS7) Vacuolar ATP synthase subunit C
(V-ATPase C subunit) (Vacuolar proton pump C subunit) ,
partial (66%)
Length = 1048
Score = 73.6 bits (179), Expect = 2e-14
Identities = 32/41 (78%), Positives = 36/41 (87%)
Frame = +2
Query: 2 AVVWWVVSLPVHNSASELWNRLQQNVSKHSFDTPLYRFNIP 42
A +WVVSLPV NSAS LWN+LQ+ +SKHSFDTPLYRFNIP
Sbjct: 134 ATRYWVVSLPVQNSASTLWNKLQEQISKHSFDTPLYRFNIP 256
>TC226372 similar to UP|VATC_ARATH (Q9SDS7) Vacuolar ATP synthase subunit C
(V-ATPase C subunit) (Vacuolar proton pump C subunit)
, partial (45%)
Length = 577
Score = 70.9 bits (172), Expect(2) = 1e-13
Identities = 31/39 (79%), Positives = 36/39 (91%), Gaps = 1/39 (2%)
Frame = +2
Query: 5 WWVVSLPVHNSAS-ELWNRLQQNVSKHSFDTPLYRFNIP 42
+W VSLPVHNSAS +LWN+LQ+ +SKHSFDTPLYRFNIP
Sbjct: 32 YWAVSLPVHNSASSQLWNQLQERISKHSFDTPLYRFNIP 148
Score = 20.4 bits (41), Expect(2) = 1e-13
Identities = 9/12 (75%), Positives = 10/12 (83%)
Frame = +3
Query: 42 PISGSEP*TLSA 53
P S SEP*TLS+
Sbjct: 147 PTSASEP*TLSS 182
>TC215978 weakly similar to UP|Q8S1K2 (Q8S1K2) PDI-like protein, partial (34%)
Length = 1554
Score = 26.2 bits (56), Expect = 3.7
Identities = 17/49 (34%), Positives = 26/49 (52%)
Frame = +1
Query: 8 VSLPVHNSASELWNRLQQNVSKHSFDTPLYRFNIPISGSEP*TLSAMIS 56
V++ SASELWN + S+ TPL I ++ S P +++M S
Sbjct: 1225 VAMKCTQSASELWNAMTM-----SWWTPLVASRI*LTNSIPLAINSMSS 1356
>TC225163 emb|X02559.1|MIOBRN26 Oenothera berteriana mitochondrial gene for
26S ribosomal RNA, partial (83%)
Length = 3184
Score = 25.4 bits (54), Expect = 6.3
Identities = 7/27 (25%), Positives = 14/27 (50%)
Frame = -2
Query: 6 WVVSLPVHNSASELWNRLQQNVSKHSF 32
W P+HN ++E W + ++ + F
Sbjct: 399 WFFHHPIHNKSNETWRKRSEHFAAEEF 319
>AW277331
Length = 276
Score = 25.4 bits (54), Expect = 6.3
Identities = 11/25 (44%), Positives = 15/25 (60%)
Frame = +1
Query: 24 QQNVSKHSFDTPLYRFNIPISGSEP 48
QQN+ + FD L +F+ S SEP
Sbjct: 91 QQNIDRREFDDDLKKFSSLPSNSEP 165
>TC217335 similar to UP|GIC2_YEAST (Q06648) GTPase-interacting component 2,
partial (5%)
Length = 1795
Score = 25.0 bits (53), Expect = 8.3
Identities = 8/18 (44%), Positives = 12/18 (66%)
Frame = +2
Query: 4 VWWVVSLPVHNSASELWN 21
+WW V++P +S LWN
Sbjct: 329 IWWGVAIPPSSSICCLWN 382
>TC206847 similar to GB|AAF76441.1|8569096|F2J10 ESTs gb|AI994059, gb|T43740
come from this gene. {Arabidopsis thaliana;} , partial
(46%)
Length = 1571
Score = 25.0 bits (53), Expect = 8.3
Identities = 13/48 (27%), Positives = 24/48 (49%)
Frame = +2
Query: 6 WVVSLPVHNSASELWNRLQQNVSKHSFDTPLYRFNIPISGSEP*TLSA 53
W++ P + + W RLQ ++ + YRF + ++P TL+A
Sbjct: 755 WLIIRPDGDGTWKPWGRLQAWREPNNSNAVGYRFEVLPGTADPVTLAA 898
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.135 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,919,968
Number of Sequences: 63676
Number of extensions: 31588
Number of successful extensions: 245
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 243
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 244
length of query: 56
length of database: 12,639,632
effective HSP length: 32
effective length of query: 24
effective length of database: 10,602,000
effective search space: 254448000
effective search space used: 254448000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0199.9


