

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
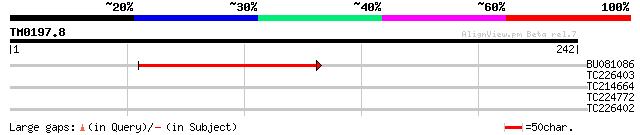
Query= TM0197.8
(242 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza s... 67 9e-12
TC226403 similar to UP|Q9FPW6 (Q9FPW6) POZ/BTB containing-protei... 29 2.2
TC214664 weakly similar to GB|AAL31248.1|16974489|AY061921 At2g2... 27 6.4
TC224772 homologue to UP|O65760 (O65760) Extensin (Fragment), pa... 27 8.3
TC226402 similar to UP|Q9FPW6 (Q9FPW6) POZ/BTB containing-protei... 27 8.3
>BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 426
Score = 66.6 bits (161), Expect = 9e-12
Identities = 34/78 (43%), Positives = 48/78 (60%)
Frame = +2
Query: 56 TT*FLNNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVIT 115
TT F+N+ + + NHK+ LKVD IMLLRN+DQ GLCN TRLI+ + I A +++
Sbjct: 56 TTEFINSLSTSRLPNHKIKLKVDSRIMLLRNLDQNEGLCNSTRLIITMFADHIIEAKIMS 235
Query: 116 GTNANDKVHISIMDLVPS 133
G + V+I + PS
Sbjct: 236 GKGQGNTVYIPRLATSPS 289
>TC226403 similar to UP|Q9FPW6 (Q9FPW6) POZ/BTB containing-protein AtPOB1,
partial (20%)
Length = 548
Score = 28.9 bits (63), Expect = 2.2
Identities = 15/39 (38%), Positives = 22/39 (55%)
Frame = +2
Query: 72 KLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIG 110
+ LL + + + LLR DQ L T+ I+ LGER +G
Sbjct: 119 RALLALPLTMNLLRGQDQQRNLLASTKAIMYSLGERPLG 235
>TC214664 weakly similar to GB|AAL31248.1|16974489|AY061921 At2g27080/T20P8.13
{Arabidopsis thaliana;} , partial (56%)
Length = 1097
Score = 27.3 bits (59), Expect = 6.4
Identities = 11/33 (33%), Positives = 20/33 (60%)
Frame = -1
Query: 186 LEYAQERVLNFEYWNSQANNRPVL*M*CLRKFL 218
+ YAQ ++ + + W + N +* *C+RKF+
Sbjct: 1070 MTYAQNQIKS*DTWRXRINKCATM*H*CIRKFV 972
>TC224772 homologue to UP|O65760 (O65760) Extensin (Fragment), partial (12%)
Length = 1452
Score = 26.9 bits (58), Expect = 8.3
Identities = 16/69 (23%), Positives = 29/69 (41%)
Frame = +2
Query: 174 PSSLMVSYM*LSLEYAQERVLNFEYWNSQANNRPVL*M*CLRKFLEMFDVYNTISFIYFI 233
P+ M + ++ ++ NF +W N +L + C+ F F++F
Sbjct: 773 PTFAMAGALAWQVKNINKKSNNFWFWGRSHGNSGLLPLVCVMFFF----------FLFFF 922
Query: 234 FLLIYYYTI 242
FLL Y Y +
Sbjct: 923 FLLYYLYML 949
>TC226402 similar to UP|Q9FPW6 (Q9FPW6) POZ/BTB containing-protein AtPOB1,
partial (77%)
Length = 1565
Score = 26.9 bits (58), Expect = 8.3
Identities = 14/36 (38%), Positives = 20/36 (54%)
Frame = +3
Query: 72 KLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGER 107
+ LL + + + LLR DQ L T+ I+ LGER
Sbjct: 1152 RALLALPLTMNLLRGQDQQRNLLASTKAIMYSLGER 1259
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.341 0.150 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,591,406
Number of Sequences: 63676
Number of extensions: 113579
Number of successful extensions: 1115
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1103
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1114
length of query: 242
length of database: 12,639,632
effective HSP length: 95
effective length of query: 147
effective length of database: 6,590,412
effective search space: 968790564
effective search space used: 968790564
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0197.8


