

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0195a.11
(377 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
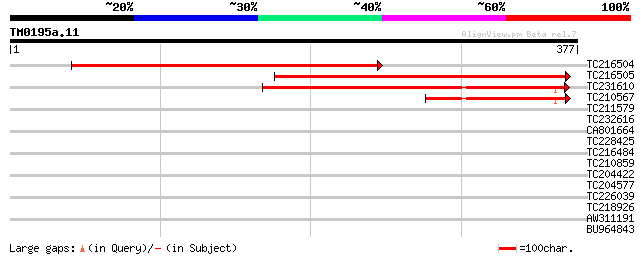
Score E
Sequences producing significant alignments: (bits) Value
TC216504 260 8e-70
TC216505 221 4e-58
TC231610 similar to UP|Q9ASQ7 (Q9ASQ7) AT4g27970/T13J8_80, parti... 184 7e-47
TC210567 67 2e-11
TC211579 similar to UP|Q8S8K1 (Q8S8K1) Predicted protein, partia... 36 0.033
TC232616 similar to GB|AAD25756.1|4587525|T5I8 Contains the PF|0... 32 0.61
CA801664 30 2.3
TC228425 similar to UP|O81447 (O81447) Calcineurin B-like protei... 29 3.0
TC216484 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabin... 29 3.0
TC210859 29 4.0
TC204422 28 5.2
TC204577 weakly similar to UP|EXL1_ARATH (Q9LZT4) Expansin-like ... 28 5.2
TC226039 similar to UP|O48618 (O48618) Cytochome b5 (Fragment), ... 28 5.2
TC218926 similar to UP|CAZ_DROME (Q27294) RNA-binding protein ca... 28 5.2
AW311191 similar to SP|Q9BBN9|NU1C_ NAD(P)H-quinone oxidoreducta... 28 6.8
BU964843 weakly similar to GP|21553602|gb| glycine-rich RNA-bind... 28 8.9
>TC216504
Length = 843
Score = 260 bits (664), Expect = 8e-70
Identities = 123/207 (59%), Positives = 157/207 (75%)
Frame = +3
Query: 42 VLTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLA 101
+LTKLHAGYF I LS G QALLWK+L D TL H +++PS AFL+LW +AL T
Sbjct: 222 ILTKLHAGYFFICLSFGAQALLWKSLSKHNKDSQTLWHGFNLMPSIAFLLLWCVALLTAT 401
Query: 102 LLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWV 161
LSLLY+L+C+FH +MVK EF HHVGVN ++APWISW L+LQSAP + T+ Y VL
Sbjct: 402 TLSLLYVLKCIFHLDMVKEEFSHHVGVNCMYAPWISWLLMLQSAPIIVHSTSCYGVLCLA 581
Query: 162 FAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFS 221
F+ +++LD+K+YGQWFT KRFLS ANPTS +SVIGNLV AQ A +GWKESA+ + S
Sbjct: 582 FSFVILLLDIKLYGQWFTTKKRFLSVVANPTSLVSVIGNLVSAQVVAEIGWKESALLMVS 761
Query: 222 LGMVHYLVLFVTLYQRLSGGNRLPVLL 248
LG+V+YL++FVTLYQRL+ G++ P +L
Sbjct: 762 LGLVYYLIIFVTLYQRLTSGSQFPTVL 842
>TC216505
Length = 760
Score = 221 bits (563), Expect = 4e-58
Identities = 110/197 (55%), Positives = 139/197 (69%)
Frame = +1
Query: 177 WFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVLFVTLYQ 236
WF K LS TS +SVIGN V AQ A +G ESA+ +FSLG+V+YL++FVTLYQ
Sbjct: 1 WFXTKKXXLSVVXXXTSLVSVIGNXVSAQVVAEIGXXESALLMFSLGLVYYLIIFVTLYQ 180
Query: 237 RLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPT 296
RL+ GN+ P +LRP + LFFAAP +ASLAW SI G F T SKML FLSLFLFMS CRP
Sbjct: 181 RLTSGNQFPTVLRPTYLLFFAAPSMASLAWKSITGAFVTPSKMLLFLSLFLFMSQACRPA 360
Query: 297 LFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVSLALTVF 356
+F++SMKR NV WWAYSFP+T L +A +YA+EVK ++ LMLV+ +SVLV +AL +
Sbjct: 361 MFKKSMKRLNVTWWAYSFPLTFLGLACAEYAQEVKSGMASWLMLVICMVSVLVFIALMII 540
Query: 357 TFINSKMLLPDDDPIAT 373
T + + L + PI T
Sbjct: 541 TVLKIERLFHKNAPIMT 591
>TC231610 similar to UP|Q9ASQ7 (Q9ASQ7) AT4g27970/T13J8_80, partial (40%)
Length = 863
Score = 184 bits (466), Expect = 7e-47
Identities = 96/207 (46%), Positives = 131/207 (62%), Gaps = 3/207 (1%)
Frame = +3
Query: 169 LDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYL 228
L++KIYGQW + G R LS ANPT+ LSV+GN VGA A MG KE + +++G+ HY+
Sbjct: 3 LELKIYGQWMSGGTRRLSKVANPTNHLSVVGNFVGALLGASMGLKEGPIFFYAIGLAHYI 182
Query: 229 VLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLF 288
V+FVTLYQRL LP L PVFF+F A P VAS+AW SI FD S++ +F+ LFL+
Sbjct: 183 VVFVTLYQRLPTNETLPKELHPVFFMFVAPPVVASMAWASIQDSFDYGSRISYFIGLFLY 362
Query: 289 MSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVL 348
SL R FR F++AWWAY+FP+ A A+ YA EV ++ L ++L +S L
Sbjct: 363 FSLAVRVNFFRGF--TFSLAWWAYTFPMASTASATVRYANEVTNVVTKTLSVILSLMSTL 536
Query: 349 VSLALTVFTFINS---KMLLPDDDPIA 372
+ A+ V T +++ K L P+D IA
Sbjct: 537 IIAAVLVSTILHALVFKDLFPNDHAIA 617
>TC210567
Length = 545
Score = 66.6 bits (161), Expect = 2e-11
Identities = 33/100 (33%), Positives = 61/100 (61%), Gaps = 3/100 (3%)
Frame = +2
Query: 277 SKMLFFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISH 336
S++ +F ++FL++SL R LFR +F+++WWAY+FP+T A+A+ Y +V ++
Sbjct: 5 SRIFYFTAMFLYISLAVRVNLFRGF--KFSLSWWAYTFPMTAAAIATITYTNQVTNVLTQ 178
Query: 337 VLMLVLLALSVLVSLALTVFTFINS---KMLLPDDDPIAT 373
L ++L ++ A+ V T +++ + L P+D IAT
Sbjct: 179 ALSVILSLIATFTVTAVLVSTIVHAFVLRDLFPNDLAIAT 298
>TC211579 similar to UP|Q8S8K1 (Q8S8K1) Predicted protein, partial (20%)
Length = 742
Score = 35.8 bits (81), Expect = 0.033
Identities = 17/49 (34%), Positives = 23/49 (46%)
Frame = +3
Query: 112 LFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWW 160
LFHF F++ +G F W++WF Q P + YLVL W
Sbjct: 396 LFHF------FIYELGFGSCFV*WVAWFFKFQRGPCLGTCLYIYLVLSW 524
>TC232616 similar to GB|AAD25756.1|4587525|T5I8 Contains the PF|00650
CRAL/TRIO phosphatidyl-inositol-transfer protein domain.
ESTs gb|T76582, gb|N06574 and gb|Z25700 come from this
gene. {Arabidopsis thaliana;} , partial (48%)
Length = 1214
Score = 31.6 bits (70), Expect = 0.61
Identities = 15/42 (35%), Positives = 22/42 (51%)
Frame = -2
Query: 98 FTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWF 139
FT L L +R + F +V+ FLH G ++ W+SWF
Sbjct: 313 FTQGLSFKLLEIRKVVAFFLVRRAFLHRFGFGFIH**WLSWF 188
>CA801664
Length = 439
Score = 29.6 bits (65), Expect = 2.3
Identities = 17/68 (25%), Positives = 36/68 (52%)
Frame = +1
Query: 75 STLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAP 134
+TLR ++ + FL+ + + L + L SLL L+ +F+F + F+ ++ +
Sbjct: 106 NTLRSLIKKLKKEIFLLKYKIILLMIFLFSLLILIINIFNF*FL--IFIINISIYKSICH 279
Query: 135 WISWFLLL 142
+ WF++L
Sbjct: 280 VLIWFVVL 303
>TC228425 similar to UP|O81447 (O81447) Calcineurin B-like protein 3, complete
Length = 1542
Score = 29.3 bits (64), Expect = 3.0
Identities = 16/48 (33%), Positives = 26/48 (53%)
Frame = +1
Query: 85 PSSAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLF 132
P+ AFL+ + ++ +SLLYL L M+K + +V NY+F
Sbjct: 1396 PTLAFLLDRKIMPLSIQFISLLYLPLFLARLLMIKNQLAAYVFCNYIF 1539
>TC216484 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (58%)
Length = 1213
Score = 29.3 bits (64), Expect = 3.0
Identities = 15/46 (32%), Positives = 19/46 (40%), Gaps = 7/46 (15%)
Frame = -1
Query: 127 GVNYLFAPWISWFLLLQSAP-------FVAPKTATYLVLWWVFAVP 165
G N F PW S L + P ++ A LV+WW F P
Sbjct: 460 GANATFPPWRSVMLTKPADPGAVPVAWYIVAARAVPLVIWWSFLAP 323
>TC210859
Length = 1210
Score = 28.9 bits (63), Expect = 4.0
Identities = 18/43 (41%), Positives = 26/43 (59%)
Frame = -2
Query: 68 IGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLR 110
+GP KSTLR + + PSSA L S +L +AL++ +L R
Sbjct: 339 LGPWAQKSTLRFMRTLGPSSAALAQSSWSLLLMALVTGPFLGR 211
>TC204422
Length = 1820
Score = 28.5 bits (62), Expect = 5.2
Identities = 17/37 (45%), Positives = 21/37 (55%), Gaps = 2/37 (5%)
Frame = -3
Query: 68 IGPTHDKSTLRHVVHMVPSSAFLV--LWSLALFTLAL 102
+GP KSTLR + + PSSA L WSL L L +
Sbjct: 369 LGPWAQKSTLRFMRTLGPSSAALAQSSWSLLLMVLVI 259
>TC204577 weakly similar to UP|EXL1_ARATH (Q9LZT4) Expansin-like 1 precursor
(At-EXPL1) (Ath-ExpBeta-2.1), partial (8%)
Length = 1025
Score = 28.5 bits (62), Expect = 5.2
Identities = 16/57 (28%), Positives = 25/57 (43%), Gaps = 1/57 (1%)
Frame = -2
Query: 38 SLNSVLTKLHAGYFRISLSLGGQALLWKTLIG-PTHDKSTLRHVVHMVPSSAFLVLW 93
+L ++ L G+F L+LGG ++ + G H +RH H P F W
Sbjct: 868 TLVPIICNLFLGFFLFKLNLGGISISRLPVTGYDNHGVHPIRHATHPKP*PDFATWW 698
>TC226039 similar to UP|O48618 (O48618) Cytochome b5 (Fragment), partial
(62%)
Length = 775
Score = 28.5 bits (62), Expect = 5.2
Identities = 21/68 (30%), Positives = 34/68 (49%)
Frame = +3
Query: 224 MVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFL 283
+V L+LF++L Q+ GNRL P+ FLFF + +L + G + +FF+
Sbjct: 528 IVLVLLLFISL*QQCPSGNRLV----PLLFLFFLGHHLKALHYLKYHAGIN----FVFFM 683
Query: 284 SLFLFMSL 291
L + L
Sbjct: 684 CCGLLLKL 707
>TC218926 similar to UP|CAZ_DROME (Q27294) RNA-binding protein cabeza
(Sarcoma-associated RNA-binding fly homolog) (P19),
partial (5%)
Length = 906
Score = 28.5 bits (62), Expect = 5.2
Identities = 10/35 (28%), Positives = 21/35 (59%)
Frame = -2
Query: 106 LYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFL 140
+++L + FN++ + HV +N+ F ++ WFL
Sbjct: 902 IHVLEKIS*FNIITLQVRSHVELNFFFGLFVKWFL 798
>AW311191 similar to SP|Q9BBN9|NU1C_ NAD(P)H-quinone oxidoreductase chain 1
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase chain
1), partial (25%)
Length = 318
Score = 28.1 bits (61), Expect = 6.8
Identities = 17/51 (33%), Positives = 28/51 (54%)
Frame = +2
Query: 268 SIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTV 318
+I+G F TL+K LFLF+S+ R TL R + + W + P+++
Sbjct: 98 TIIGIFITLAKTY----LFLFLSIATRWTLPRLRIDQLLNLGWKFLLPISL 238
>BU964843 weakly similar to GP|21553602|gb| glycine-rich RNA-binding
protein-like {Arabidopsis thaliana}, partial (27%)
Length = 435
Score = 27.7 bits (60), Expect = 8.9
Identities = 19/55 (34%), Positives = 27/55 (48%), Gaps = 1/55 (1%)
Frame = +1
Query: 150 PKTATYLVLWWVFAVPVVVLDVKIYGQWFTKGKRF-LSTAANPTSQLSVIGNLVG 203
P AT + VF + +LDVKI T+GK + T NP S + I ++ G
Sbjct: 97 PYDATEETIRTVFNLYGAILDVKIINDPRTRGKCYCFVTFTNPRSAIDAINDMNG 261
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.330 0.141 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,527,015
Number of Sequences: 63676
Number of extensions: 328444
Number of successful extensions: 2584
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 2555
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2580
length of query: 377
length of database: 12,639,632
effective HSP length: 99
effective length of query: 278
effective length of database: 6,335,708
effective search space: 1761326824
effective search space used: 1761326824
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0195a.11


