

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0193.7
(848 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
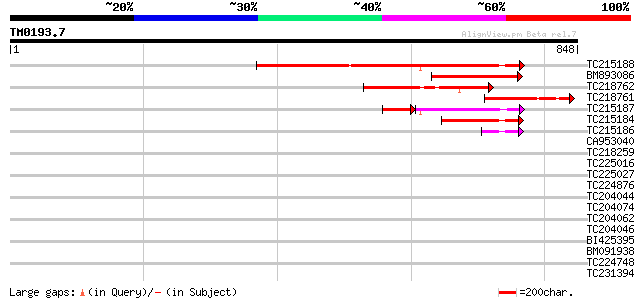
Score E
Sequences producing significant alignments: (bits) Value
TC215188 homologue to UP|Q9XIS5 (Q9XIS5) Starch branching enzyme... 509 e-144
BM893086 homologue to GP|5441248|dbj branching enzyme 3 {Phaseol... 260 2e-69
TC218762 homologue to UP|Q9XIS4 (Q9XIS4) Starch branching enzyme... 218 9e-57
TC218761 similar to UP|Q9XIS4 (Q9XIS4) Starch branching enzyme ... 154 1e-37
TC215187 similar to UP|Q9XIS5 (Q9XIS5) Starch branching enzyme ... 124 2e-28
TC215184 homologue to UP|Q9XIS5 (Q9XIS5) Starch branching enzyme... 120 3e-27
TC215186 homologue to UP|Q9XIS5 (Q9XIS5) Starch branching enzyme... 49 1e-05
CA953040 homologue to PIR|B85091|B85 isoamylase-like protein [im... 35 0.18
TC218259 31 2.6
TC225016 similar to UP|Q39835 (Q39835) Extensin, partial (32%) 30 4.5
TC225027 similar to UP|Q39835 (Q39835) Extensin, partial (49%) 30 4.5
TC224876 weakly similar to UP|Q39835 (Q39835) Extensin, partial ... 30 4.5
TC204044 homologue to UP|Q09083 (Q09083) Hydroxyproline-rich gly... 30 5.8
TC204074 homologue to UP|Q39835 (Q39835) Extensin, partial (26%) 30 5.8
TC204062 homologue to UP|Q09083 (Q09083) Hydroxyproline-rich gly... 30 5.8
TC204046 UP|Q39835 (Q39835) Extensin, complete 30 5.8
BI425395 similar to PIR|T10863|T108 extensin precursor - kidney ... 30 5.8
BM091938 30 5.8
TC224748 similar to UP|Q9FFD8 (Q9FFD8) Gb|AAF26981.1 (At5g16470)... 30 5.8
TC231394 similar to UP|Q9LW11 (Q9LW11) Zinc finger protein-like;... 29 7.6
>TC215188 homologue to UP|Q9XIS5 (Q9XIS5) Starch branching enzyme , partial
(48%)
Length = 1878
Score = 509 bits (1311), Expect = e-144
Identities = 243/412 (58%), Positives = 302/412 (72%), Gaps = 11/412 (2%)
Frame = +3
Query: 370 YFHTGDRGYHKLWDSRLFNYANWEVLRFLLSNLRWWLEEFKFDGFRFDGVTSMLYHHHGV 429
YFH G RGYH +WDSRLFNY +WEVLR+LLSN RWWL+E+KFDGFRFDGVTSM+Y HHG+
Sbjct: 3 YFHPGSRGYHWMWDSRLFNYGSWEVLRYLLSNARWWLDEYKFDGFRFDGVTSMMYTHHGL 182
Query: 430 NIAFSGDYNEYFSEATDVDAVVYLMLANSLIHNILPDATVIAEDVSGMPGLGRPISEVGI 489
+AF+G+YNEYF ATDVDAVVYLML N +IH + P+A I EDVSGMP P + GI
Sbjct: 183 EVAFTGNYNEYFGFATDVDAVVYLMLTNDVIHGLFPEAVTIGEDVSGMPTFCLPTQDGGI 362
Query: 490 GFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGDKT 549
GFDYRL MAI DKWI+ LK K D +W M +I +LTNRR+ EKCV+YAESHDQ++VGDKT
Sbjct: 363 GFDYRLHMAIADKWIEILK-KNDEDWKMGDIIHTLTNRRWLEKCVAYAESHDQALVGDKT 539
Query: 550 FSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFGHPEWI 609
+F LMD+++Y M+ ++P I+RGIALHKMI ITM LGGEGYLNFMGNEFGHPEWI
Sbjct: 540 IAFWLMDKDMYDFMALDRPSTPIIDRGIALHKMIRLITMGLGGEGYLNFMGNEFGHPEWI 719
Query: 610 DFPR-----------EGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLA 658
DFPR GN S++KCRR++ L D D+LRY+ M FD+AM L++KF F+
Sbjct: 720 DFPRGDQHLPNGVVVPGNNNSFDKCRRRFDLGDADYLRYQGMQEFDQAMQHLEEKFGFMT 899
Query: 659 STKQIVSSTNEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFG 718
+ Q +S NE DK+IVFERG+L+FVFNFH +Y Y+VGC PGKY++ LDSD FG
Sbjct: 900 AEHQYISRKNEGDKIIVFERGNLIFVFNFHWTNSYSDYRVGCSTPGKYKIVLDSDDALFG 1079
Query: 719 GHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVYYRVDESQ 770
G R+ H ++FT+ E +++RP SF I +P RT VVY DE++
Sbjct: 1080GFSRLNHAAEYFTS--------EGWYDDRPRSFLIYAPSRTAVVYALADEAE 1211
>BM893086 homologue to GP|5441248|dbj branching enzyme 3 {Phaseolus
vulgaris}, partial (16%)
Length = 408
Score = 260 bits (665), Expect = 2e-69
Identities = 120/136 (88%), Positives = 130/136 (95%)
Frame = +1
Query: 632 TDHLRYKFMNAFDKAMNLLDDKFSFLASTKQIVSSTNEEDKVIVFERGDLVFVFNFHPET 691
TDHLRYKFMNAFD+AMNLLDDKFSFLASTKQIVSS +++DKVIVFERGDL+FVFNFHPE
Sbjct: 1 TDHLRYKFMNAFDRAMNLLDDKFSFLASTKQIVSSADDDDKVIVFERGDLIFVFNFHPEN 180
Query: 692 TYEGYKVGCDLPGKYRVALDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSF 751
TYEGYKVGCDLPGKYRVALDSDA EFGG GRVGH+VDHFT+PEGIPGVPE+NFNNRPNSF
Sbjct: 181 TYEGYKVGCDLPGKYRVALDSDAWEFGGRGRVGHDVDHFTSPEGIPGVPETNFNNRPNSF 360
Query: 752 KILSPPRTCVVYYRVD 767
K+LSP RTCV YYRV+
Sbjct: 361 KVLSPARTCVAYYRVE 408
>TC218762 homologue to UP|Q9XIS4 (Q9XIS4) Starch branching enzyme , partial
(23%)
Length = 909
Score = 218 bits (555), Expect = 9e-57
Identities = 117/203 (57%), Positives = 140/203 (68%), Gaps = 9/203 (4%)
Frame = +3
Query: 530 SEKCVSYAESHDQSIVGDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMS 589
+EKCVSYAESHDQ++VGDKT +FLLMD E+YSG SCL +ASP +E+GI L KMIHFITM+
Sbjct: 9 AEKCVSYAESHDQAVVGDKTVAFLLMD*EMYSGTSCLVEASPIVEQGIYLQKMIHFITMA 188
Query: 590 LGGEGYLNFMGNEFGHPEWIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNL 649
LGGEGYLNFMGNEFGHPEWID + + + R + R + + ++
Sbjct: 189 LGGEGYLNFMGNEFGHPEWIDSQEK----AMDGVMRSAGVSGIGGYR-----SLEIQVHE 341
Query: 650 LDDKFSFLASTKQIVSSTNEED---------KVIVFERGDLVFVFNFHPETTYEGYKVGC 700
+ + LA ++ N+ D IVFERGDL+FVFNFHPE TYEGYKVGC
Sbjct: 342 CI*QSNELA***LLIPCINKTDCEQCR*R*QGYIVFERGDLIFVFNFHPENTYEGYKVGC 521
Query: 701 DLPGKYRVALDSDAREFGGHGRV 723
DLPGKYRV LDSDA EFGGHGRV
Sbjct: 522 DLPGKYRVVLDSDAWEFGGHGRV 590
>TC218761 similar to UP|Q9XIS4 (Q9XIS4) Starch branching enzyme , partial
(12%)
Length = 780
Score = 154 bits (390), Expect = 1e-37
Identities = 85/136 (62%), Positives = 99/136 (72%)
Frame = +2
Query: 710 LDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVYYRVDES 769
LD DA EFG RV +VDHFT+PEGIPGVPE+NFNNRPNSFK+LSP RTCV YYRV+ES
Sbjct: 2 LDXDAWEFGXRXRVXXDVDHFTSPEGIPGVPETNFNNRPNSFKVLSPARTCVAYYRVEES 181
Query: 770 QEENSISNLVGVQETSTAADIVANIPDGSSASKEREVSNFNWTMETLAAANADVAKIPDE 829
QE++ ++LVGV+ETS AAD VA IPD SAS E E + ETLAA ADVAKIPDE
Sbjct: 182 QEDDDNNSLVGVEETSAAAD-VAKIPD-ESASTESEDIKLDGVKETLAA--ADVAKIPDE 349
Query: 830 LVPAAENEVFQDEVED 845
P + D V++
Sbjct: 350 SAPLESEDSNLDVVKE 397
>TC215187 similar to UP|Q9XIS5 (Q9XIS5) Starch branching enzyme , partial
(25%)
Length = 817
Score = 124 bits (310), Expect = 2e-28
Identities = 63/175 (36%), Positives = 102/175 (58%), Gaps = 11/175 (6%)
Frame = +2
Query: 607 EWIDFPR-----------EGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFS 655
EWIDFPR GN S++KC+R+++L + D+L+Y+ M F++A+ L++ F
Sbjct: 272 EWIDFPRGDQHLPNGVVVPGNNNSFDKCKRKFNLGNADYLKYQGMQEFNQAIXHLEEXFD 451
Query: 656 FLASTKQIVSSTNEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAR 715
F+ + Q +S NE +K+IVF++G+L+F+FNFH +Y Y+V P KY++ L+SD
Sbjct: 452 FITAKHQYISQKNESNKIIVFKKGNLIFIFNFH*TNSYSDYRVNYSTPRKYKIILNSDNT 631
Query: 716 EFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVYYRVDESQ 770
F + H +FT+ + ++N+P SF I +P RT Y DES+
Sbjct: 632 LFNNFR*LNHAAKYFTSGK--------*YDNQPQSFLIYTPSRTAXGYTLADESK 772
Score = 73.2 bits (178), Expect = 5e-13
Identities = 33/49 (67%), Positives = 39/49 (79%)
Frame = +3
Query: 558 EIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFGHP 606
++Y M+ ++P I+RGIALHKMI ITM LGGEGYLNFMGNEFGHP
Sbjct: 3 DMYDFMALDRPSTPIIDRGIALHKMIRLITMGLGGEGYLNFMGNEFGHP 149
>TC215184 homologue to UP|Q9XIS5 (Q9XIS5) Starch branching enzyme , partial
(16%)
Length = 1220
Score = 120 bits (301), Expect = 3e-27
Identities = 57/122 (46%), Positives = 78/122 (63%)
Frame = +2
Query: 647 MNLLDDKFSFLASTKQIVSSTNEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDLPGKY 706
M L++KF F+ + Q +S NE DK+IVFERG+L+FVFNFH +Y Y+VGC PGKY
Sbjct: 209 MQHLEEKFGFMTAEHQYISRKNEGDKIIVFERGNLIFVFNFHWNNSYSDYRVGCSTPGKY 388
Query: 707 RVALDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVYYRV 766
++ LDSD FGG R+ H ++FT+ E +++RP SF I +P RT VVY
Sbjct: 389 KIVLDSDDALFGGFSRLNHTAEYFTS--------EGWYDDRPRSFLIYAPSRTAVVYALA 544
Query: 767 DE 768
D+
Sbjct: 545 DD 550
>TC215186 homologue to UP|Q9XIS5 (Q9XIS5) Starch branching enzyme , partial
(6%)
Length = 589
Score = 48.5 bits (114), Expect = 1e-05
Identities = 24/63 (38%), Positives = 36/63 (57%)
Frame = +3
Query: 706 YRVALDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVYYR 765
+++ LDSD FGG R+ H ++FT+ E +++RP SF I +P RT VVY
Sbjct: 402 FQIVLDSDDALFGGFSRLNHTAEYFTS--------EGWYDDRPRSFLIYAPSRTAVVYAL 557
Query: 766 VDE 768
D+
Sbjct: 558 ADD 566
>CA953040 homologue to PIR|B85091|B85 isoamylase-like protein [imported] -
Arabidopsis thaliana, partial (19%)
Length = 421
Score = 34.7 bits (78), Expect = 0.18
Identities = 24/83 (28%), Positives = 38/83 (44%), Gaps = 3/83 (3%)
Frame = +1
Query: 338 LHVLMDVVHSHASNNVTDGLNGFDVGQVSQESYFHTGDRGYHKLWDSRLFNYANWE---V 394
+ V++DVV++H +N D + ++ D L S N N V
Sbjct: 10 MQVILDVVYNH-TNEADDAFPYTTSFRGIDNKVYYMLDNNGQLLNFSGCGNTLNCNHPVV 186
Query: 395 LRFLLSNLRWWLEEFKFDGFRFD 417
+ +L +LR W+ E+ DGFRFD
Sbjct: 187 MELILDSLRHWVTEYHVDGFRFD 255
>TC218259
Length = 1295
Score = 30.8 bits (68), Expect = 2.6
Identities = 24/84 (28%), Positives = 40/84 (47%), Gaps = 10/84 (11%)
Frame = +2
Query: 141 IVYREWAPAAQEAQIIGDFNEWNGSNHPMEK------NQFGV-WSIKIPDVAGNPAIPHN 193
I+YR W+ +EA + NG P +K N+FG+ W I + +G+P H
Sbjct: 56 IIYR-WSQLNKEANL----KRKNGRARPQQK*KEPKQNKFGLFWRISLALTSGSPPSLHA 220
Query: 194 SRVKFRFRHGGVWADRI---PAWI 214
++K + GV+A + P W+
Sbjct: 221 FQLKVSLANLGVFASALVSRPLWM 292
>TC225016 similar to UP|Q39835 (Q39835) Extensin, partial (32%)
Length = 973
Score = 30.0 bits (66), Expect = 4.5
Identities = 15/38 (39%), Positives = 19/38 (49%)
Frame = +2
Query: 231 VYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEP 268
V++ PP S + +KYP PP P P Y H S P
Sbjct: 443 VHYSPPPSPKKPYKYPSPPPPVHPP-YTPHPVYHSPPP 553
>TC225027 similar to UP|Q39835 (Q39835) Extensin, partial (49%)
Length = 747
Score = 30.0 bits (66), Expect = 4.5
Identities = 15/38 (39%), Positives = 19/38 (49%)
Frame = +2
Query: 231 VYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEP 268
V++ PP S + +KYP PP P P Y H S P
Sbjct: 611 VHYSPPPSPKKPYKYPSPPPPVHPP-YTPHPVYHSPPP 721
>TC224876 weakly similar to UP|Q39835 (Q39835) Extensin, partial (66%)
Length = 1340
Score = 30.0 bits (66), Expect = 4.5
Identities = 25/97 (25%), Positives = 42/97 (42%), Gaps = 6/97 (6%)
Frame = +3
Query: 231 VYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPR------INSYKEFADDILPRI 284
V++ PP S + +KYP PP P P + ++ S P + Y+E+ I
Sbjct: 1035 VHYSPPPSPKKPYKYPSPPPP--PPTNKPYIYASPPPPYHY*LA*LYGYEEYVQRIKESD 1208
Query: 285 RANNYNTVQLMAVMEHSYYASFGYHVTNFFAVSSRSG 321
R + + E ++ SFG V++FF + G
Sbjct: 1209 RKGRRSASR--CTKEKNWKQSFG--VSDFFLLLMNIG 1307
Score = 29.3 bits (64), Expect = 7.6
Identities = 17/48 (35%), Positives = 20/48 (41%)
Frame = +1
Query: 221 PTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEP 268
P + PY Y PP + +KYP PP P P Y H S P
Sbjct: 505 PPPYKKPY--YYHSPPPPPKKPYKYPSPPPPVHPP-YTPHPVYHSPPP 639
>TC204044 homologue to UP|Q09083 (Q09083) Hydroxyproline-rich glycoprotein
precursor, partial (38%)
Length = 1080
Score = 29.6 bits (65), Expect = 5.8
Identities = 13/36 (36%), Positives = 17/36 (47%)
Frame = +2
Query: 216 YATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKP 251
Y + P K + P Y PP + +KYP PP P
Sbjct: 5 YHSPPPPKHSPPPPYYYKSPPPPPKEPYKYPSPPPP 112
>TC204074 homologue to UP|Q39835 (Q39835) Extensin, partial (26%)
Length = 366
Score = 29.6 bits (65), Expect = 5.8
Identities = 13/36 (36%), Positives = 17/36 (47%)
Frame = +1
Query: 216 YATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKP 251
Y + P K + P Y PP + +KYP PP P
Sbjct: 34 YHSPPPPKHSPPPPYYYKSPPPPPKEPYKYPSPPPP 141
>TC204062 homologue to UP|Q09083 (Q09083) Hydroxyproline-rich glycoprotein
precursor, partial (34%)
Length = 722
Score = 29.6 bits (65), Expect = 5.8
Identities = 13/36 (36%), Positives = 17/36 (47%)
Frame = +1
Query: 216 YATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKP 251
Y + P K + P Y PP + +KYP PP P
Sbjct: 100 YHSPPPPKHSPPPPYYYKSPPPPPKKPYKYPSPPPP 207
>TC204046 UP|Q39835 (Q39835) Extensin, complete
Length = 1627
Score = 29.6 bits (65), Expect = 5.8
Identities = 13/36 (36%), Positives = 17/36 (47%)
Frame = +1
Query: 216 YATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKP 251
Y + P K + P Y PP + +KYP PP P
Sbjct: 574 YHSPPPPKHSPPPPYYYKSPPPPPKKPYKYPSPPPP 681
>BI425395 similar to PIR|T10863|T108 extensin precursor - kidney bean,
partial (13%)
Length = 421
Score = 29.6 bits (65), Expect = 5.8
Identities = 13/36 (36%), Positives = 17/36 (47%)
Frame = +3
Query: 216 YATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKP 251
Y + P K + P Y PP + +KYP PP P
Sbjct: 6 YHSPPPPKHSPPPPYYYKSPPPPPKEPYKYPSPPPP 113
>BM091938
Length = 421
Score = 29.6 bits (65), Expect = 5.8
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = +2
Query: 231 VYWDPPLSERYQFKYPRPPKPKAP 254
V++ PP S + +KYP PP P P
Sbjct: 317 VHYSPPPSPKKPYKYPSPPPPVHP 388
>TC224748 similar to UP|Q9FFD8 (Q9FFD8) Gb|AAF26981.1 (At5g16470), partial
(88%)
Length = 881
Score = 29.6 bits (65), Expect = 5.8
Identities = 22/65 (33%), Positives = 31/65 (46%)
Frame = -2
Query: 214 IKYATVDPTKFAAPYDGVYWDPPLSERYQFKYPRPPKPKAPRIYEAHVGMSSSEPRINSY 273
I A PT+F Y PP R + PRPP P++P SS+ P ++SY
Sbjct: 352 ISRARPSPTRFGPSY------PPSRRRGSWS-PRPPSPRSP-----SPCASSASPSLSSY 209
Query: 274 KEFAD 278
+ F+D
Sbjct: 208 R-FSD 197
>TC231394 similar to UP|Q9LW11 (Q9LW11) Zinc finger protein-like; Ser/Thr
protein kinase-like protein, partial (62%)
Length = 689
Score = 29.3 bits (64), Expect = 7.6
Identities = 20/59 (33%), Positives = 25/59 (41%)
Frame = +2
Query: 701 DLPGKYRVALDSDAREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRT 759
DL G Y + +R FGG GR G + G E NF +R FK +P T
Sbjct: 230 DLAGSYNSSDILRSRAFGGSGRPGWKSGDWICTRS--GCNEHNFASRMECFKCSAPRDT 400
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,286,965
Number of Sequences: 63676
Number of extensions: 577902
Number of successful extensions: 2918
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 2617
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2839
length of query: 848
length of database: 12,639,632
effective HSP length: 105
effective length of query: 743
effective length of database: 5,953,652
effective search space: 4423563436
effective search space used: 4423563436
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0193.7


