

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
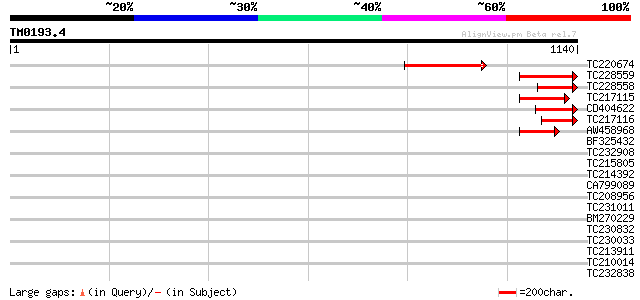
Query= TM0193.4
(1140 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC220674 similar to GB|CAD41408.1|21741286|OSJN00110 OSJNBb0078D... 272 5e-73
TC228559 similar to GB|CAD41408.1|21741286|OSJN00110 OSJNBb0078D... 180 4e-45
TC228558 similar to UP|Q9SHN9 (Q9SHN9) F7F22.1, partial (4%) 141 1e-33
TC217115 similar to GB|CAD41408.1|21741286|OSJN00110 OSJNBb0078D... 132 1e-30
CD404622 119 6e-27
TC217116 similar to UP|Q9SHN9 (Q9SHN9) F7F22.1, partial (4%) 107 4e-23
AW458968 56 8e-08
BF325432 39 0.017
TC232908 similar to UP|Q9FZ97 (Q9FZ97) F3H9.11 protein, partial ... 33 0.71
TC215805 33 0.71
TC214392 similar to GB|AAP31966.1|30387601|BT006622 At3g19640 {A... 32 2.1
CA799089 31 2.7
TC208956 GB|AAP47439.1|32331823|AY164865 RTNLB18w {Glycine max;}... 31 3.5
TC231011 weakly similar to UP|Q8GTR4 (Q8GTR4) Pullulanase-like p... 31 3.5
BM270229 GP|16648779|gb At1g69160/F4N2_9 {Arabidopsis thaliana},... 31 3.5
TC230832 weakly similar to UP|Q9LIC4 (Q9LIC4) Gb|AAD32917.1, par... 30 6.0
TC230033 weakly similar to UP|IF34_YEAST (Q04067) Eukaryotic tra... 30 6.0
TC213911 similar to UP|Q9LW67 (Q9LW67) Ankyrin-like protein, par... 30 6.0
TC210014 similar to UP|Q9FMH3 (Q9FMH3) Similarity to chloroplast... 30 7.9
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 30 7.9
>TC220674 similar to GB|CAD41408.1|21741286|OSJN00110 OSJNBb0078D11.16 {Oryza
sativa (japonica cultivar-group);} , partial (11%)
Length = 505
Score = 272 bits (696), Expect = 5e-73
Identities = 127/164 (77%), Positives = 143/164 (86%)
Frame = +1
Query: 795 VKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFVV 854
VKI+VIA+F AF LASIAL TRIEPGLEQ+I LPRDSYLQGYF+N+SEYLR+GPPLYFVV
Sbjct: 1 VKILVIAVFAAFTLASIALCTRIEPGLEQQIALPRDSYLQGYFSNISEYLRVGPPLYFVV 180
Query: 855 KNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPE 914
K+YNYS ES HTNQLCSIS C+S+SLLNEIS+ASLVP +SYIAKPAASWLDDFLVWISPE
Sbjct: 181 KDYNYSLESKHTNQLCSISHCDSNSLLNEISRASLVPTSSYIAKPAASWLDDFLVWISPE 360
Query: 915 AFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRH 958
AF CCRKFTN SYCPPDDQPPCC P++ C G + C ++H
Sbjct: 361 AFSCCRKFTNDSYCPPDDQPPCCLPDEGPCGLGG-EVCKXLYKH 489
>TC228559 similar to GB|CAD41408.1|21741286|OSJN00110 OSJNBb0078D11.16 {Oryza
sativa (japonica cultivar-group);} , partial (10%)
Length = 1037
Score = 180 bits (456), Expect = 4e-45
Identities = 95/116 (81%), Positives = 103/116 (87%)
Frame = +3
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYF 1084
V+ V S+ A EF ++ S VASGD+DQR KEALGTMGASVFSGITLTKLVGVIVL F
Sbjct: 114 VNLVMSVGIAVEFCVHMTHSFTVASGDRDQRAKEALGTMGASVFSGITLTKLVGVIVLCF 293
Query: 1085 SRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPPSRCTIIEQEENRSSTSS 1140
SRTEVFVIYYF+MYLSLVLLGFLHGLVFLPVVLSIFGPPSRC+IIEQEE+RSSTSS
Sbjct: 294 SRTEVFVIYYFRMYLSLVLLGFLHGLVFLPVVLSIFGPPSRCSIIEQEEDRSSTSS 461
>TC228558 similar to UP|Q9SHN9 (Q9SHN9) F7F22.1, partial (4%)
Length = 562
Score = 141 bits (356), Expect = 1e-33
Identities = 71/79 (89%), Positives = 77/79 (96%)
Frame = +3
Query: 1062 TMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFG 1121
TMGASVFSGITLTKLVGVIVL FS+TEVFVIYYF+MYLSLVLLGFLHGLVFLPV+LS+FG
Sbjct: 3 TMGASVFSGITLTKLVGVIVLCFSKTEVFVIYYFRMYLSLVLLGFLHGLVFLPVLLSVFG 182
Query: 1122 PPSRCTIIEQEENRSSTSS 1140
PPSRC+IIEQ E+RSSTSS
Sbjct: 183 PPSRCSIIEQGEDRSSTSS 239
>TC217115 similar to GB|CAD41408.1|21741286|OSJN00110 OSJNBb0078D11.16 {Oryza
sativa (japonica cultivar-group);} , partial (8%)
Length = 777
Score = 132 bits (331), Expect = 1e-30
Identities = 66/101 (65%), Positives = 81/101 (79%)
Frame = +1
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYF 1084
V+ + S+ A EF + + V+ GD+ QR K AL TMGASVFSGITLTKLVGV+VL F
Sbjct: 124 VNLIMSIGIAVEFCVHIVHAFMVSLGDRSQRAKTALCTMGASVFSGITLTKLVGVLVLCF 303
Query: 1085 SRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPPSR 1125
S +E+FV+YYFQMYL+LV++GFLHGLVFLPVVLS+FGPP R
Sbjct: 304 STSEIFVVYYFQMYLALVIIGFLHGLVFLPVVLSLFGPPLR 426
>CD404622
Length = 552
Score = 119 bits (299), Expect = 6e-27
Identities = 62/85 (72%), Positives = 73/85 (84%), Gaps = 1/85 (1%)
Frame = -2
Query: 1057 KEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVV 1116
K AL TMGASVF+GITLT LVGV+VL FS +++FV+YYFQMYL+LVL+GFLHGLVFLPVV
Sbjct: 539 KTALCTMGASVFTGITLTILVGVLVLCFSTSQIFVVYYFQMYLALVLIGFLHGLVFLPVV 360
Query: 1117 LSIFGPPSRCTII-EQEENRSSTSS 1140
LS+FGPP R T+I EQ E+ S SS
Sbjct: 359 LSLFGPPLRYTVIKEQVEDMPSASS 285
>TC217116 similar to UP|Q9SHN9 (Q9SHN9) F7F22.1, partial (4%)
Length = 1213
Score = 107 bits (266), Expect = 4e-23
Identities = 54/73 (73%), Positives = 64/73 (86%), Gaps = 1/73 (1%)
Frame = +1
Query: 1069 SGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPPSRCTI 1128
SGITLTKLVGV+VL FS +++FV+YYFQMYL+LVL+GFLHGLVFLPVVLS+FGPP R T+
Sbjct: 721 SGITLTKLVGVLVLCFSTSQIFVVYYFQMYLALVLIGFLHGLVFLPVVLSLFGPPLRYTV 900
Query: 1129 I-EQEENRSSTSS 1140
I EQ E+ S SS
Sbjct: 901 IKEQLEDMPSASS 939
>AW458968
Length = 409
Score = 56.2 bits (134), Expect = 8e-08
Identities = 35/81 (43%), Positives = 51/81 (62%)
Frame = +2
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYF 1084
V+ + S+ A EF + ++ V+ D+ QR K AL TM +SVFSGITLTKL G L F
Sbjct: 167 VNLMMSIGMAGEFCEHIVHAVTVS*RDRSQRSKTALRTMRSSVFSGITLTKLDGEHDLCF 346
Query: 1085 SRTEVFVIYYFQMYLSLVLLG 1105
S +++ V+ QMY ++ L+G
Sbjct: 347 STSQMSVVDD*QMYTAVGLIG 409
>BF325432
Length = 414
Score = 38.5 bits (88), Expect = 0.017
Identities = 14/23 (60%), Positives = 17/23 (73%), Gaps = 1/23 (4%)
Frame = +1
Query: 933 QPPCCAPEDDSC-VSGACKDCTT 954
QPPCC P++ C + G CKDCTT
Sbjct: 199 QPPCCLPDEGPCGLGGVCKDCTT 267
>TC232908 similar to UP|Q9FZ97 (Q9FZ97) F3H9.11 protein, partial (39%)
Length = 725
Score = 33.1 bits (74), Expect = 0.71
Identities = 25/84 (29%), Positives = 42/84 (49%)
Frame = -2
Query: 852 FVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWI 911
F+ + +++T T CSIS+ ++ S+ N + + L+P T+ P + +D I
Sbjct: 430 FITSKFPNMADATTTPSHCSISKRSAPSMSNPVLRPGLIPLTN----PQCTVIDS---RI 272
Query: 912 SPEAFGCCRKFTNGSYCPPDDQPP 935
S E+F C F G PP +PP
Sbjct: 271 SKESFLC---FKVG--YPPKSEPP 215
>TC215805
Length = 783
Score = 33.1 bits (74), Expect = 0.71
Identities = 21/78 (26%), Positives = 34/78 (42%), Gaps = 5/78 (6%)
Frame = +2
Query: 919 CRKFTNGSYCPPDDQPPC-----CAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRD 973
CR+ SY PP D+P CA + +SC++G C R+ + + ++
Sbjct: 242 CRQPLPESYAPPADEPWMTGIFGCAEDRESCLTGLFCPCVLFGRNVE---------RLKE 394
Query: 974 KLPWFLSALPSADCAKGG 991
PW + A C +GG
Sbjct: 395 DTPWTGPCICHAICIEGG 448
>TC214392 similar to GB|AAP31966.1|30387601|BT006622 At3g19640 {Arabidopsis
thaliana;} , partial (67%)
Length = 1330
Score = 31.6 bits (70), Expect = 2.1
Identities = 30/139 (21%), Positives = 58/139 (41%), Gaps = 7/139 (5%)
Frame = -1
Query: 375 FFDSHLAPFYRIEQLILATVPDHMNSTSPRIVSADNIRFLFEVQKKVDAIRANYSGLMVS 434
F + HL+ + + + +L +PD +N+T SA N+ F+++ G+ +
Sbjct: 670 FREIHLSHIFVVIEQVLQFIPDLLNTTGDGD*SALNLTNAFQIESTDLGGELVQGGMSLL 491
Query: 435 LQDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVEHLNYCFQQYSSADQCMSAFK---- 490
LQ + D + Q+ L+ +++ +N D V + D C+ K
Sbjct: 490 LQGFSLVFQDATSSLQTGLEGNELEGKNLDGVVGVGVGVSISLVVVADDPCLELMKEGGD 311
Query: 491 ---APLDPSTVLGGFSGKD 506
A ++ +LGG G D
Sbjct: 310 GGVARVEEEELLGGDDGLD 254
>CA799089
Length = 435
Score = 31.2 bits (69), Expect = 2.7
Identities = 23/79 (29%), Positives = 35/79 (44%), Gaps = 4/79 (5%)
Frame = +1
Query: 81 ELTCSPNQSLFINVTSVDKAGGNSTVGGIDYFVSDAF----GEGLYESCKDVKFGSMNSR 136
+ TCSPNQ++ +++ K G N Y V+D GEG+ C G + S
Sbjct: 133 DFTCSPNQTVELSINLTMKGGDN-------YLVTDPIVTVSGEGINLLC----LGVLKSN 279
Query: 137 AIQFIGAGAQNFKEWFAFI 155
+ IG QNF + +
Sbjct: 280 NVNIIG---QNFMTGYRIV 327
>TC208956 GB|AAP47439.1|32331823|AY164865 RTNLB18w {Glycine max;} , complete
Length = 1256
Score = 30.8 bits (68), Expect = 3.5
Identities = 35/117 (29%), Positives = 55/117 (46%), Gaps = 2/117 (1%)
Frame = +2
Query: 552 AQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSK 611
A RN E + EE+ + A + + ++ +++ A+ +T+G F I K
Sbjct: 470 ASLRNKQPPTLPELVVSEEMVN-NVAASFRVKINNMLLIAH-DITIG-----KDFRIFFK 628
Query: 612 VLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVG--VDNMCILVHA 666
V+ + L +LSV+GSV F L TL +M IP L G VD C L+H+
Sbjct: 629 VV-----ICLWLLSVIGSVFSFFTLAYIGTL-MMITIPALYHKYGGYVDKCCGLIHS 781
>TC231011 weakly similar to UP|Q8GTR4 (Q8GTR4) Pullulanase-like protein
(Starch debranching enzyme), partial (12%)
Length = 749
Score = 30.8 bits (68), Expect = 3.5
Identities = 11/24 (45%), Positives = 17/24 (70%)
Frame = +2
Query: 936 CCAPEDDSCVSGACKDCTTCFRHS 959
CCA +D+SCV GA ++ + CF +
Sbjct: 317 CCASKDNSCVCGAKENVSHCFNQT 388
>BM270229 GP|16648779|gb At1g69160/F4N2_9 {Arabidopsis thaliana}, partial
(3%)
Length = 435
Score = 30.8 bits (68), Expect = 3.5
Identities = 13/28 (46%), Positives = 15/28 (53%)
Frame = +1
Query: 186 NVSAYSCSDTSLGCSCGDCPSSSVCSNS 213
NV +C LGCSCG C S + C S
Sbjct: 349 NVMQMACVLNLLGCSCGTCASRAACWES 432
>TC230832 weakly similar to UP|Q9LIC4 (Q9LIC4) Gb|AAD32917.1, partial (4%)
Length = 719
Score = 30.0 bits (66), Expect = 6.0
Identities = 13/39 (33%), Positives = 21/39 (53%)
Frame = -1
Query: 188 SAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANSCSI 226
S +S T G S PSS++CS+S+ ++ CS+
Sbjct: 350 SLHSSVRTKTGSSSSSSPSSTICSSSSEIRASEEKGCSL 234
>TC230033 weakly similar to UP|IF34_YEAST (Q04067) Eukaryotic translation
initiation factor 3 RNA-binding subunit (eIF-3
RNA-binding subunit) (eIF3 p33) (Translation initiation
factor eIF3, p33 subunit), partial (9%)
Length = 491
Score = 30.0 bits (66), Expect = 6.0
Identities = 17/69 (24%), Positives = 27/69 (38%)
Frame = +1
Query: 142 GAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSSGMKPMNVSAYSCSDTSLGCSC 201
GAG ++ F F GR + P+ R + S + +++C C C
Sbjct: 61 GAGRSRWRR*FFFQGRASLPHRTS-------R*TPARRSSCQTRVPISFNCRRNKKYCKC 219
Query: 202 GDCPSSSVC 210
CPS + C
Sbjct: 220 RSCPSKAFC 246
>TC213911 similar to UP|Q9LW67 (Q9LW67) Ankyrin-like protein, partial (29%)
Length = 537
Score = 30.0 bits (66), Expect = 6.0
Identities = 18/60 (30%), Positives = 24/60 (40%)
Frame = +2
Query: 901 ASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSD 960
AS + ++W+ P C K PD PP C+P DD K R+SD
Sbjct: 314 ASKRNQTVIWVKPLTNDCYLKRE------PDTHPPLCSPSDDPDAVWGVKMKACITRYSD 475
>TC210014 similar to UP|Q9FMH3 (Q9FMH3) Similarity to chloroplast nucleoid
DNA-binding protein, partial (12%)
Length = 576
Score = 29.6 bits (65), Expect = 7.9
Identities = 39/162 (24%), Positives = 63/162 (38%), Gaps = 15/162 (9%)
Frame = +1
Query: 473 NYCFQQYS-SADQCMSAFKAPLDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGN--- 528
NY F+ C+ F DP+T+LGG ++ +V Y D E +
Sbjct: 40 NYLFRHSKVRGAYCLGVFSNGNDPTTLLGGIVVRN------TLVMY------DREHSKIG 183
Query: 529 --ETAKAVAWEKAFIQLVKDELL-PMAQSRNLTLAFS-----SESSIEEELKRESTADAI 580
+T + WE+ + L+ P ++ NLT AF S S +L A I
Sbjct: 184 FWKTNCSELWERLHVSNAPPPLMPPKSEGTNLTKAFKPSVAPSPSQYNLQLGELQIAQLI 363
Query: 581 TIL---VSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGV 619
++ +SY+ + YI+ G H S L+ S +
Sbjct: 364 VVISFNISYMDIKPYITELTGLIAHELDVNTSQVHLMNFSSL 489
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 29.6 bits (65), Expect = 7.9
Identities = 19/52 (36%), Positives = 28/52 (53%)
Frame = -2
Query: 1067 VFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLS 1118
+F + + V VIVL+F VF+I+ Y F+HGLVF V+L+
Sbjct: 400 IFLVVIVVSFVVVIVLFF----VFIIFCVVYY-------FIHGLVFFVVLLN 278
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 52,794,908
Number of Sequences: 63676
Number of extensions: 816159
Number of successful extensions: 4428
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 4347
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4420
length of query: 1140
length of database: 12,639,632
effective HSP length: 108
effective length of query: 1032
effective length of database: 5,762,624
effective search space: 5947027968
effective search space used: 5947027968
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0193.4


