

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0191.15
(398 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
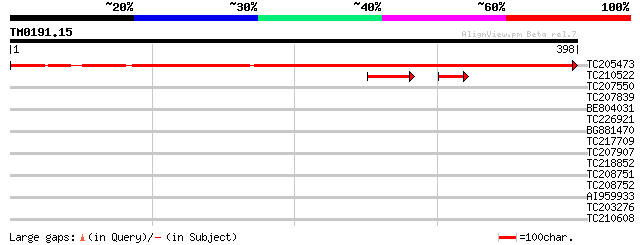
Score E
Sequences producing significant alignments: (bits) Value
TC205473 UP|HEM6_SOYBN (P35055) Coproporphyrinogen III oxidase, ... 702 0.0
TC210522 similar to UP|HEM6_ARATH (Q9LR75) Coproporphyrinogen II... 62 4e-11
TC207550 31 0.86
TC207839 31 1.1
BE804031 weakly similar to GP|15624060|dbj B1088C09.21 {Oryza sa... 30 1.9
TC226921 similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein p... 30 2.5
BG881470 30 2.5
TC217709 29 3.3
TC207907 similar to UP|Q96313 (Q96313) AT.I.24-13 protein, parti... 29 4.3
TC218852 similar to GB|AAP37807.1|30725570|BT008448 At2g29660 {A... 28 5.6
TC208751 similar to UP|Q84Q60 (Q84Q60) At5g49350, partial (6%) 28 7.3
TC208752 similar to UP|Q84Q60 (Q84Q60) At5g49350, partial (6%) 28 7.3
AI959933 weakly similar to GP|27461187|gb type IV collagen alpha... 28 9.5
TC203276 similar to GB|AAL15273.1|16323077|AY057642 AT5g36230/T3... 28 9.5
TC210608 weakly similar to PIR|E96527|E96527 protein F27J15.22 [... 28 9.5
>TC205473 UP|HEM6_SOYBN (P35055) Coproporphyrinogen III oxidase, chloroplast
precursor (Coproporphyrinogenase) (Coprogen oxidase) ,
complete
Length = 1621
Score = 702 bits (1812), Expect = 0.0
Identities = 340/398 (85%), Positives = 360/398 (90%)
Frame = +1
Query: 1 MPSCATIVSPPSYAIPFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVS 60
M CA+IVS PSYA PF + ++ST+ PTA L+KR W P +KG VRA VS
Sbjct: 61 MMHCASIVSAPSYAFPFRSGSASTT-PTAISLTKRSWKPPPSM-------AKGPVRATVS 216
Query: 61 IEKETPEAERPETFLRGGDTAEALSSSSSVRARFEKMIREAQDSVCDAIEAADGGAKFKE 120
IEKETPEA RPETFLRG D A+ SS+SVRARFEKMIREAQD+VC A+EAADGGA+FKE
Sbjct: 217 IEKETPEANRPETFLRGVDEAQ---SSTSVRARFEKMIREAQDTVCSALEAADGGAQFKE 387
Query: 121 DVWSRPGGGGGISRVLQDGAVWEKAGVNVSVVYGVMPPDAYRAAKGAAASPDEKPGPVPF 180
DVWSRPGGGGGISRVLQDGAVWEKAGVNVSVVYGVMPPDAYRAAKG D+KPGPVPF
Sbjct: 388 DVWSRPGGGGGISRVLQDGAVWEKAGVNVSVVYGVMPPDAYRAAKGVPT--DQKPGPVPF 561
Query: 181 FAAGISSVLHPKNPFAPTMHFNYRYFETDAPKDAPGAPKQWWFGGGTDLTPAYIFEEDVK 240
FAAGISSVLHPKNPFAPT+HFNYRYFETDAPKDAPGAP+QWWFGGGTDLTPAYIFEEDVK
Sbjct: 562 FAAGISSVLHPKNPFAPTLHFNYRYFETDAPKDAPGAPRQWWFGGGTDLTPAYIFEEDVK 741
Query: 241 HFHSIQKQACDKFDPTFYPRFKKWCDDYFYIKHRGERRGLGGIFFDDLNDYDQEMLLSFA 300
HFHSIQKQACDKF+PTFYPRFKKWCDDYFYIKHRGERRGLGGIFFDDLNDYDQEMLLSFA
Sbjct: 742 HFHSIQKQACDKFEPTFYPRFKKWCDDYFYIKHRGERRGLGGIFFDDLNDYDQEMLLSFA 921
Query: 301 TECANSVVPSYIPIIEKRKDTPFTDHQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIE 360
TECANSV+P+Y+PIIEKRKD PF DHQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIE
Sbjct: 922 TECANSVIPAYLPIIEKRKDLPFNDHQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRIE 1101
Query: 361 SILVSLPLTSRWEYDHKPEEGSEEWKLLDACINPKEWI 398
SILVSLPLT+RWEYDHKPEEGSEEWKLLDACINPKEWI
Sbjct: 1102SILVSLPLTARWEYDHKPEEGSEEWKLLDACINPKEWI 1215
>TC210522 similar to UP|HEM6_ARATH (Q9LR75) Coproporphyrinogen III oxidase,
chloroplast precursor (Coproporphyrinogenase) (Coprogen
oxidase) , partial (9%)
Length = 762
Score = 62.4 bits (150), Expect(2) = 4e-11
Identities = 26/33 (78%), Positives = 28/33 (84%)
Frame = +3
Query: 252 KFDPTFYPRFKKWCDDYFYIKHRGERRGLGGIF 284
KFDPTFYP +KKWCDDY YI+HR ERRGL IF
Sbjct: 351 KFDPTFYP*YKKWCDDYCYIEHRDERRGLREIF 449
Score = 23.1 bits (48), Expect(2) = 4e-11
Identities = 9/21 (42%), Positives = 14/21 (65%)
Frame = +2
Query: 302 ECANSVVPSYIPIIEKRKDTP 322
E A +P+++P+ EKR D P
Sbjct: 428 ERATRNIPAWVPVGEKRWDCP 490
>TC207550
Length = 775
Score = 31.2 bits (69), Expect = 0.86
Identities = 22/75 (29%), Positives = 36/75 (47%)
Frame = +1
Query: 16 PFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVSIEKETPEAERPETFL 75
P +S SS+S+PT + NP+ ++SL +KG +S E E+P E
Sbjct: 139 PASSSVSSSSAPTLNAAKNNKLNPI--ANLLSSLVAKGL----ISAETESPTTVPSEAPK 300
Query: 76 RGGDTAEALSSSSSV 90
D E +++S S+
Sbjct: 301 GSKDQTEIITTSCSL 345
>TC207839
Length = 785
Score = 30.8 bits (68), Expect = 1.1
Identities = 23/91 (25%), Positives = 37/91 (40%)
Frame = -2
Query: 4 CATIVSPPSYAIPFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVSIEK 63
C T+ PP A+ FF S + ++ +P S G+ G + +
Sbjct: 529 CRTLAVPPPPALLFFASLPAEAT-----------SPFRGLSLSLSAGAGGVTSSEIDPSD 383
Query: 64 ETPEAERPETFLRGGDTAEALSSSSSVRARF 94
AE FLR G TA A++ +S + + F
Sbjct: 382 GLHSAEPILIFLREGTTATAVAGTSDLSSSF 290
>BE804031 weakly similar to GP|15624060|dbj B1088C09.21 {Oryza sativa
(japonica cultivar-group)}, partial (37%)
Length = 413
Score = 30.0 bits (66), Expect = 1.9
Identities = 26/107 (24%), Positives = 45/107 (41%), Gaps = 7/107 (6%)
Frame = +1
Query: 2 PSCATIVSPPSYAIPFFTSTSSTSSPTAKLLSK----RRWNPLIQT---KTMTSLGSKGA 54
PSCA SP + A T+ S+TSS + + R ++ + T T ++ ++ A
Sbjct: 58 PSCAASASPSTSATSPSTTASATSSTPSSAAAATSPCRAYSSVASTLAELTTSASSTRAA 237
Query: 55 VRAGVSIEKETPEAERPETFLRGGDTAEALSSSSSVRARFEKMIREA 101
G S P P D+ A+S++ + ++ K EA
Sbjct: 238 SSTGSSNASRAPTRTTPAILAEASDS*CAMSATEATKSSQRKTDLEA 378
>TC226921 similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein precursor,
partial (5%)
Length = 641
Score = 29.6 bits (65), Expect = 2.5
Identities = 17/40 (42%), Positives = 20/40 (49%)
Frame = -3
Query: 101 AQDSVCDAIEAADGGAKFKEDVWSRPGGGGGISRVLQDGA 140
A+D AA GG D + R GGGGG+ L DGA
Sbjct: 339 AEDGCFREGAAARGGVSVGVDDFQRVGGGGGLVADLVDGA 220
>BG881470
Length = 382
Score = 29.6 bits (65), Expect = 2.5
Identities = 21/75 (28%), Positives = 37/75 (49%)
Frame = +2
Query: 16 PFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVSIEKETPEAERPETFL 75
P ++ SS+S+PT + NP+ + ++SL +KG +S E E+P E
Sbjct: 26 PASSNISSSSAPTLNTAKNNKLNPI--SNLLSSLVAKGL----ISAETESPTMVPSEVPK 187
Query: 76 RGGDTAEALSSSSSV 90
D E +++S S+
Sbjct: 188GSKDQTEIITTSCSL 232
>TC217709
Length = 795
Score = 29.3 bits (64), Expect = 3.3
Identities = 21/77 (27%), Positives = 40/77 (51%)
Frame = -1
Query: 13 YAIPFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLGSKGAVRAGVSIEKETPEAERPE 72
++IPF +S + ++ + S +PLI ++T T+ +K A + G++ + T P
Sbjct: 693 HSIPF*SSVAGCTAGSVVARSST*SSPLI*SRTSTNPMAKAASKPGLTQRENT---SFPS 523
Query: 73 TFLRGGDTAEALSSSSS 89
T L G T +S++ S
Sbjct: 522 TTLVVGSTLRRISATDS 472
>TC207907 similar to UP|Q96313 (Q96313) AT.I.24-13 protein, partial (80%)
Length = 858
Score = 28.9 bits (63), Expect = 4.3
Identities = 30/131 (22%), Positives = 50/131 (37%), Gaps = 7/131 (5%)
Frame = +2
Query: 245 IQKQACDKFDPTFYPRFKKWCDDYFYIKHRGERRGLGGIFFDDLNDYDQEMLLSFATECA 304
+ K C K D YP FK + D +++G R D E + +F E A
Sbjct: 338 MDKAVCSKVDIHSYPTFKVFYDGEEVARYQGTR--------------DVESMKTFVLEEA 475
Query: 305 NSVVPSYIPIIEKRKDTPFTDHQKAWQQLRRGRYVEFNLVYDRG-------TTFGLKTGG 357
EK +++ Q LR Y+++N ++ G TFG +
Sbjct: 476 -----------EKAAAKALESNKEL*QLLRCSHYLDYN*LFTVGGAHPR*VITFGFRLRH 622
Query: 358 RIESILVSLPL 368
S+L+++ L
Sbjct: 623 SFFSVLINICL 655
>TC218852 similar to GB|AAP37807.1|30725570|BT008448 At2g29660 {Arabidopsis
thaliana;} , partial (54%)
Length = 1201
Score = 28.5 bits (62), Expect = 5.6
Identities = 13/31 (41%), Positives = 17/31 (53%)
Frame = -1
Query: 127 GGGGGISRVLQDGAVWEKAGVNVSVVYGVMP 157
GGGGG+ L G VW AG+ ++ MP
Sbjct: 127 GGGGGLRGGLGGGGVWGFAGLGGEEMFDEMP 35
>TC208751 similar to UP|Q84Q60 (Q84Q60) At5g49350, partial (6%)
Length = 635
Score = 28.1 bits (61), Expect = 7.3
Identities = 22/65 (33%), Positives = 31/65 (46%), Gaps = 2/65 (3%)
Frame = -3
Query: 70 RPETFLRGGDTAEALSSSSSVRARFEKMIREAQDSVCDAIEAADGGA--KFKEDVWSRPG 127
R TFL A + S +++ F+ ++ DS + GGA K E W RPG
Sbjct: 399 RTTTFL----FANSPSRNNATTFLFDSEEHQSPDS*KLV*MSGGGGAFGKKGESPWLRPG 232
Query: 128 GGGGI 132
GGGG+
Sbjct: 231 GGGGL 217
>TC208752 similar to UP|Q84Q60 (Q84Q60) At5g49350, partial (6%)
Length = 356
Score = 28.1 bits (61), Expect = 7.3
Identities = 22/65 (33%), Positives = 31/65 (46%), Gaps = 2/65 (3%)
Frame = -1
Query: 70 RPETFLRGGDTAEALSSSSSVRARFEKMIREAQDSVCDAIEAADGGA--KFKEDVWSRPG 127
R TFL A + S +++ F+ ++ DS + GGA K E W RPG
Sbjct: 347 RTTTFL----FANSPSRNNATTFLFDSEEHQSPDS*KLV*MSGGGGAFGKKGESPWLRPG 180
Query: 128 GGGGI 132
GGGG+
Sbjct: 179 GGGGL 165
>AI959933 weakly similar to GP|27461187|gb type IV collagen alpha 5 chain
{Canis familiaris}, partial (2%)
Length = 380
Score = 27.7 bits (60), Expect = 9.5
Identities = 31/101 (30%), Positives = 40/101 (38%), Gaps = 3/101 (2%)
Frame = -2
Query: 127 GGGGGISRVLQDGAVWEKAGVNVSVVYGVMPPDAYRAAKGAAASPDEKPGPVPFFA-AGI 185
GGG +R W+ G + +G PP A P +PFF+ AG
Sbjct: 316 GGGFFFTRGAPGFPPWKGGGGPPPIFWGGPPPWA--------------PPGLPFFSRAGF 179
Query: 186 SSVLHPKNPFAPTMHFNYRYFETDAPKD--APGAPKQWWFG 224
L NPF P F+ +F P APGA Q +FG
Sbjct: 178 WGGLGGGNPFPPGTPFSLFWFFFPPPLGGWAPGA--QIFFG 62
>TC203276 similar to GB|AAL15273.1|16323077|AY057642 AT5g36230/T30G6_9
{Arabidopsis thaliana;} , complete
Length = 2082
Score = 27.7 bits (60), Expect = 9.5
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = +2
Query: 15 IPFFTSTSSTSSPTAKLLSKRRWNPLI 41
IPFF + +P A RRWNPL+
Sbjct: 1235 IPFFIGSEKEQTPRAGKPL*RRWNPLL 1315
>TC210608 weakly similar to PIR|E96527|E96527 protein F27J15.22 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(29%)
Length = 589
Score = 27.7 bits (60), Expect = 9.5
Identities = 13/29 (44%), Positives = 22/29 (75%)
Frame = +3
Query: 2 PSCATIVSPPSYAIPFFTSTSSTSSPTAK 30
PSC++ +PPS+A PF+ S+SS S +++
Sbjct: 162 PSCSS*RAPPSWA-PFWASSSSRRSSSSR 245
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.135 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,766,786
Number of Sequences: 63676
Number of extensions: 296254
Number of successful extensions: 1761
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 1741
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1755
length of query: 398
length of database: 12,639,632
effective HSP length: 99
effective length of query: 299
effective length of database: 6,335,708
effective search space: 1894376692
effective search space used: 1894376692
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0191.15


