

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0190.3
(329 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
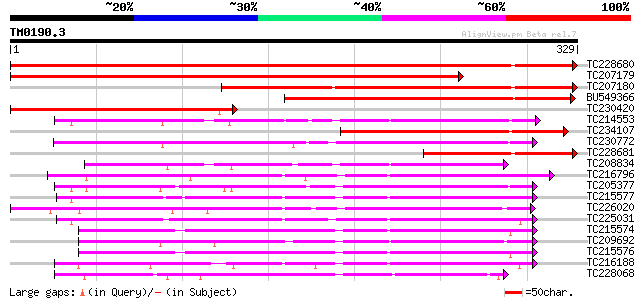
Score E
Sequences producing significant alignments: (bits) Value
TC228680 similar to UP|G2O1_PEA (Q9SQ80) Gibberellin 2-beta-diox... 535 e-153
TC207179 similar to UP|G2O1_PEA (Q9SQ80) Gibberellin 2-beta-diox... 429 e-121
TC207180 similar to GB|AAD45425.1|5579094|AF100955 gibberellin 2... 347 6e-96
BU549366 homologue to GP|4678586|emb GA 2-oxidase {Phaseolus coc... 248 2e-66
TC230420 similar to UP|G2OX_PHACN (Q9XG83) Gibberellin 2-beta-di... 151 3e-37
TC214553 similar to UP|Q39224 (Q39224) SRG1 protein (F6I1.30/F6I... 150 6e-37
TC234107 similar to UP|G2O2_PEA (Q9XHM5) Gibberellin 2-beta-diox... 148 3e-36
TC230772 similar to UP|O24648 (O24648) Gibberellin 3 beta-hydrox... 147 5e-36
TC228681 similar to UP|O04162 (O04162) Dioxygenase, partial (27%) 145 3e-35
TC208834 similar to UP|Q39224 (Q39224) SRG1 protein (F6I1.30/F6I... 141 5e-34
TC216796 similar to GB|AAQ65160.1|34365697|BT010537 At4g10500 {A... 140 6e-34
TC205377 137 5e-33
TC215577 homologue to UP|Q41681 (Q41681) 1-aminocylopropane-1-ca... 135 3e-32
TC226020 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, ... 129 2e-30
TC225031 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-c... 127 7e-30
TC215574 similar to UP|O04076 (O04076) ACC-oxidase, partial (98%) 124 4e-29
TC209692 weakly similar to UP|Q39224 (Q39224) SRG1 protein (F6I1... 124 6e-29
TC215576 similar to UP|O04076 (O04076) ACC-oxidase, partial (97%) 123 1e-28
TC216188 similar to UP|Q84L58 (Q84L58) 1-aminocyclopropane-1-car... 122 2e-28
TC228068 anthocyanidin synthase [Glycine max] 116 2e-26
>TC228680 similar to UP|G2O1_PEA (Q9SQ80) Gibberellin 2-beta-dioxygenase 1
(Gibberellin 2-beta-hydroxylase 1) (Gibberellin
2-oxidase 1) (GA 2-oxidase 1) (SLENDER protein) ,
complete
Length = 1226
Score = 535 bits (1379), Expect = e-153
Identities = 264/329 (80%), Positives = 290/329 (87%)
Frame = +1
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MVLLSK TEQYSY+KNCM T S TIP+VDLSKPDAK++IVKACEEFGFFKVINHGVSM
Sbjct: 163 MVLLSKATTEQYSYIKNCMPTKFSSTIPIVDLSKPDAKTLIVKACEEFGFFKVINHGVSM 342
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 120
EAIS+LE EA FFSM + EKEK G NPFGYG+K+IGHNGDVGW+EYLLL TNQE N
Sbjct: 343 EAISELEYEAFKFFSMSLNEKEKVGPPNPFGYGSKKIGHNGDVGWIEYLLLNTNQEHNFS 522
Query: 121 VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNH 180
V+ +NPEKFRC+L+ Y+ +VR M CEIL+LMAEGLKIQ K+V SKLLMDKQSDS+FRVNH
Sbjct: 523 VYGKNPEKFRCLLNSYMSSVRKMACEILELMAEGLKIQQKDVFSKLLMDKQSDSIFRVNH 702
Query: 181 YPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINV 240
Y ACPEM L+ +NLIGFGEHTDPQIISLLRSNNTSGLQI LRDG+WISVPPD SFFINV
Sbjct: 703 YAACPEMTLNDQNLIGFGEHTDPQIISLLRSNNTSGLQIYLRDGNWISVPPDDKSFFINV 882
Query: 241 GDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKE 300
GDSLQVMTNGRFRSV+HRVLANGF+SRLSMIYFGGPPLSEKI PL SL+KG +ESLYKE
Sbjct: 883 GDSLQVMTNGRFRSVRHRVLANGFKSRLSMIYFGGPPLSEKIPPLSSLMKG--KESLYKE 1056
Query: 301 FTWFEYKNSSYGSRLADNRLGHFERIAAS 329
FTWFEYK S YGSRL+ NRL HFERIAAS
Sbjct: 1057FTWFEYKKSIYGSRLSKNRLEHFERIAAS 1143
>TC207179 similar to UP|G2O1_PEA (Q9SQ80) Gibberellin 2-beta-dioxygenase 1
(Gibberellin 2-beta-hydroxylase 1) (Gibberellin
2-oxidase 1) (GA 2-oxidase 1) (SLENDER protein) ,
partial (80%)
Length = 1097
Score = 429 bits (1103), Expect = e-121
Identities = 208/263 (79%), Positives = 230/263 (87%)
Frame = +3
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MVLLSK TEQYSY+KN M T S TIP+VDLSKPDAK++IVKACEEFGFFKVINHGV M
Sbjct: 168 MVLLSKATTEQYSYIKNYMPTAFSSTIPIVDLSKPDAKTLIVKACEEFGFFKVINHGVPM 347
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 120
E ISQLESEA FFSMP+ EKEK G P+GYG+K+IGHNGDVGWVEYLLL TNQE N
Sbjct: 348 ETISQLESEAFKFFSMPLNEKEKAGPPKPYGYGSKKIGHNGDVGWVEYLLLNTNQEHNFS 527
Query: 121 VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNH 180
+ +N EKFRC+L+ Y+ +VR M CEIL+LMAEGLKIQ KNV SKLLMDKQSDSV RVNH
Sbjct: 528 FYGKNAEKFRCLLNSYMSSVRKMACEILELMAEGLKIQQKNVFSKLLMDKQSDSVLRVNH 707
Query: 181 YPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINV 240
YPACPE+ ++G+NLIGFGEHTDPQIISLLRSNNTSGLQI LRDG+WISVPPD+ SFFINV
Sbjct: 708 YPACPELAVNGQNLIGFGEHTDPQIISLLRSNNTSGLQIFLRDGNWISVPPDHKSFFINV 887
Query: 241 GDSLQVMTNGRFRSVKHRVLANG 263
GDSLQVMTNGRFRSV+HRVLANG
Sbjct: 888 GDSLQVMTNGRFRSVRHRVLANG 956
>TC207180 similar to GB|AAD45425.1|5579094|AF100955 gibberellin 2-oxidase
{Pisum sativum;} , partial (61%)
Length = 867
Score = 347 bits (889), Expect = 6e-96
Identities = 170/206 (82%), Positives = 188/206 (90%)
Frame = +3
Query: 124 QNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPA 183
+N EKFRC+L+ Y+ +VR M CEIL+LMAEGLKIQ KNV SKLLMDK+SDSVFRVNHYPA
Sbjct: 3 KNAEKFRCLLNSYMSSVRKMACEILELMAEGLKIQQKNVFSKLLMDKESDSVFRVNHYPA 182
Query: 184 CPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDS 243
CPE+ ++G+N+IGFGEHTDPQIISLLRSNNTSGLQI LRDG+WISVPPD+ SFFINVGDS
Sbjct: 183 CPEL-VNGQNMIGFGEHTDPQIISLLRSNNTSGLQIFLRDGNWISVPPDHKSFFINVGDS 359
Query: 244 LQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTW 303
LQVMTNGRFRSVKHRVL NGF+SRLSMIYFGGPPLSEKI PL SL+KG +ESLYKEFTW
Sbjct: 360 LQVMTNGRFRSVKHRVLTNGFKSRLSMIYFGGPPLSEKIVPLSSLMKG--KESLYKEFTW 533
Query: 304 FEYKNSSYGSRLADNRLGHFERIAAS 329
FEYKN +Y SRLADNRLGHFERI AS
Sbjct: 534 FEYKNLTYASRLADNRLGHFERIVAS 611
>BU549366 homologue to GP|4678586|emb GA 2-oxidase {Phaseolus coccineus},
partial (51%)
Length = 622
Score = 248 bits (634), Expect = 2e-66
Identities = 123/170 (72%), Positives = 146/170 (85%), Gaps = 1/170 (0%)
Frame = -2
Query: 160 KNVLSKLLMDKQSDSVFRVNHYPACPEM-GLDGENLIGFGEHTDPQIISLLRSNNTSGLQ 218
+N LS+LL D++SDS FR+N P CPE+ L+G NL+GFGEHTDPQIIS+LRSN+TSGLQ
Sbjct: 618 RNALSRLLKDEKSDSCFRLNXXPPCPEVQALNGRNLVGFGEHTDPQIISVLRSNSTSGLQ 439
Query: 219 ISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPL 278
I L DG+W+SVPPD +SFFINVGD+LQVMTNGRF+SVKHRVLA+ +SRLSMIYFGG PL
Sbjct: 438 ICLTDGTWVSVPPDQTSFFINVGDTLQVMTNGRFKSVKHRVLADPTKSRLSMIYFGGAPL 259
Query: 279 SEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLADNRLGHFERIAA 328
SEKI+PLPSL+ G EES YKEFTW+EYK ++Y SRLADNRL FE+ AA
Sbjct: 258 SEKISPLPSLMLKG-EESFYKEFTWWEYKKAAYASRLADNRLAPFEKSAA 112
>TC230420 similar to UP|G2OX_PHACN (Q9XG83) Gibberellin 2-beta-dioxygenase
(Gibberellin 2-beta-hydroxylase) (Gibberellin 2-oxidase)
(GA 2-oxidase) , partial (41%)
Length = 572
Score = 151 bits (382), Expect = 3e-37
Identities = 73/136 (53%), Positives = 97/136 (70%), Gaps = 4/136 (2%)
Frame = +2
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MV+LS+ A Q+ +K C TP IPVVDL+ PDAK+ IVKAC +FGFFK++NHGV +
Sbjct: 164 MVVLSQPALNQFFLLKTCKPTPLFSGIPVVDLTDPDAKTHIVKACRDFGFFKLVNHGVPL 343
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 120
E ++ LE+E L FF P ++K++ G +PFGYG+KRIG NGDVGWVEYLLL TN +V P
Sbjct: 344 EFMANLENETLRFFKKPQSDKDRAGPPDPFGYGSKRIGPNGDVGWVEYLLLNTNPDVISP 523
Query: 121 ----VHSQNPEKFRCV 132
+ ++P+ FR V
Sbjct: 524 KSQFIFRESPQNFRVV 571
>TC214553 similar to UP|Q39224 (Q39224) SRG1 protein (F6I1.30/F6I1.30)
(At1g17020/F6I1.30), partial (52%)
Length = 1808
Score = 150 bits (380), Expect = 6e-37
Identities = 100/298 (33%), Positives = 158/298 (52%), Gaps = 16/298 (5%)
Frame = +1
Query: 27 IPVVDLSK-----PDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
+P++DL + P + AC+E+GFF++INHGV + ++ FF++P+ EK
Sbjct: 166 VPIIDLHQLLSEDPSELEKLDHACKEWGFFQLINHGVDPPVVENMKIGVQEFFNLPMEEK 345
Query: 82 EKTGLA--NPFGYGNKRI-GHNGDVGWVEYLLLKTNQEVNVPVHSQNP-------EKFRC 131
+K + G+G + + W + T P+HS+NP + FR
Sbjct: 346 QKFWQTPEDMQGFGQLFVVSEEQKLEWADMFYAHT-----FPLHSRNPHLIPKIPQPFRE 510
Query: 132 VLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDG 191
L++Y +R M I+ LM + LKI+ N LS+L D R+N+YP CP+
Sbjct: 511 NLENYCLELRKMCITIIGLMKKALKIKT-NELSELFEDPSQG--IRMNYYPPCPQP---- 669
Query: 192 ENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNG 250
E +IG H+D ++ LL+ N GLQI +DG WI V P ++F INVGD L+++TNG
Sbjct: 670 ERVIGINPHSDSGALTILLQVNEVEGLQIR-KDGKWIPVKPLSNAFVINVGDMLEILTNG 846
Query: 251 RFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKN 308
+RS++HR + N + R+S+ F P +S I P PSL+ + +L+K +Y N
Sbjct: 847 IYRSIEHRGIVNSEKERISIAMFHRPQMSRVIGPAPSLVT-PERPALFKRIGVADYLN 1017
>TC234107 similar to UP|G2O2_PEA (Q9XHM5) Gibberellin 2-beta-dioxygenase 2
(Gibberellin 2-beta-hydroxylase 2) (Gibberellin
2-oxidase 2) (GA 2-oxidase 2) , partial (40%)
Length = 687
Score = 148 bits (374), Expect = 3e-36
Identities = 72/132 (54%), Positives = 93/132 (69%)
Frame = +3
Query: 193 NLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRF 252
N IG EH+DPQI++++RSNN GLQIS DG WI VPPD + + VGD+LQV+TNGRF
Sbjct: 39 NNIGXXEHSDPQILTIMRSNNVDGLQISTHDGLWIPVPPDPNEXXVMVGDALQVLTNGRF 218
Query: 253 RSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYG 312
SV+HRVL N ++R+SM+YF PPL+ I PLP ++ SLYK FTW +YK ++Y
Sbjct: 219 ASVRHRVLTNTTKARMSMMYFAAPPLNRWITPLPMMVT-PHNPSLYKPFTWAQYKQAAYS 395
Query: 313 SRLADNRLGHFE 324
RL D RL F+
Sbjct: 396 LRLGDARLDLFK 431
>TC230772 similar to UP|O24648 (O24648) Gibberellin 3 beta-hydroxylase
(2-oxoglutarate-dependent dioxygenase), partial (85%)
Length = 1264
Score = 147 bits (372), Expect = 5e-36
Identities = 94/288 (32%), Positives = 154/288 (52%), Gaps = 7/288 (2%)
Frame = +3
Query: 26 TIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTG 85
++PV+DL+ P+A +I AC +G ++V+NHG+ M + ++ FS+P +K K
Sbjct: 96 SVPVIDLNDPNASKLIHHACTTWGAYQVLNHGIPMSLLQDIQWVGETLFSLPSHQKHKAA 275
Query: 86 LA--NPFGYGNKRIGHN-GDVGWVE-YLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVR 141
+ GYG RI + W E + ++ + E + Q+ +K+ + Y A++
Sbjct: 276 RSPDGVDGYGLARISSFFPKLMWSEGFTIVGSPLEHFRQLWPQDYDKYCDFVMQYDEAMK 455
Query: 142 NMGCEILDLMAEGLKIQPKNVL---SKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFG 198
+ +++ LM + L I +++ SK +K + ++N YP CP D + +G
Sbjct: 456 KLVGKLMLLMLDSLGITKEDLKWAGSKGQFEKTC-AALQLNSYPTCP----DPDRAMGLA 620
Query: 199 EHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHR 258
HTD ++++L NN SGLQ+ + W++VPP INVGD L +++NG + SV HR
Sbjct: 621 AHTDSTLLTILYQNNISGLQVHRKGVGWVTVPPLSGGLVINVGDLLHILSNGLYPSVLHR 800
Query: 259 VLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
VL N Q RLS+ Y GPP + +I P L+ G + LYK TW EY
Sbjct: 801 VLVNRIQRRLSVAYLCGPPPNVEICPHAKLV-GPNKPPLYKAVTWNEY 941
>TC228681 similar to UP|O04162 (O04162) Dioxygenase, partial (27%)
Length = 664
Score = 145 bits (365), Expect = 3e-35
Identities = 74/89 (83%), Positives = 79/89 (88%)
Frame = +2
Query: 241 GDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKE 300
G S VMTNGRFRSV+HRVLANGF+SRLSMIYFGGPPLSEKIAPL SL+KG +ESLYKE
Sbjct: 230 GAST*VMTNGRFRSVRHRVLANGFKSRLSMIYFGGPPLSEKIAPLSSLMKG--KESLYKE 403
Query: 301 FTWFEYKNSSYGSRLADNRLGHFERIAAS 329
FTWFEYK S YGSRL+ NRL HFERIAAS
Sbjct: 404 FTWFEYKKSIYGSRLSKNRLEHFERIAAS 490
>TC208834 similar to UP|Q39224 (Q39224) SRG1 protein (F6I1.30/F6I1.30)
(At1g17020/F6I1.30), partial (54%)
Length = 1070
Score = 141 bits (355), Expect = 5e-34
Identities = 84/257 (32%), Positives = 139/257 (53%), Gaps = 11/257 (4%)
Frame = +2
Query: 44 ACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGLANPF--GYGNKRI-GHN 100
AC+E+GFF+++NHGV+ + ++ E +FF++P++EK+K G+G + +
Sbjct: 44 ACKEWGFFQLVNHGVNSSLVEKVRLETQDFFNLPMSEKKKFWQTPQHMEGFGQAFVVSED 223
Query: 101 GDVGWVEYLLLKTNQEVNVPVHSQNPE-------KFRCVLDDYLCAVRNMGCEILDLMAE 153
+ W + + T +P HS+ P FR L+ Y ++++ I+ LM +
Sbjct: 224 QKLDWADLYYMTT-----LPKHSRMPHLFPQLPLPFRDTLEAYSREIKDLAIVIIGLMGK 388
Query: 154 GLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEHTDP-QIISLLRSN 212
LKIQ + + + + + R+N+YP CPE E +IG H+D + LL+ N
Sbjct: 389 ALKIQEREIRE---LFEDGIQLMRMNYYPPCPEP----EKVIGLTPHSDGIGLAILLQLN 547
Query: 213 NTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIY 272
GLQI +DG W+ V P ++F +NVGD L+++TNG +RS++HR NG + RLS
Sbjct: 548 EVEGLQIR-KDGLWVPVKPLINAFIVNVGDILEIITNGIYRSIEHRATVNGEKERLSFAT 724
Query: 273 FGGPPLSEKIAPLPSLL 289
F P + P PSL+
Sbjct: 725 FYSPSSDGVVGPAPSLI 775
>TC216796 similar to GB|AAQ65160.1|34365697|BT010537 At4g10500 {Arabidopsis
thaliana;} , partial (72%)
Length = 1454
Score = 140 bits (354), Expect = 6e-34
Identities = 91/304 (29%), Positives = 155/304 (50%), Gaps = 10/304 (3%)
Frame = +1
Query: 23 CSFTIPVVDLSKPDAKSIIVK---ACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVT 79
C I + DL P+ II + AC+ +GFF+V NHGV I ++ FF +P +
Sbjct: 211 CIPLIDLQDLHGPNRSHIIQQIDQACQNYGFFQVTNHGVPEGVIEKIMKVTREFFGLPES 390
Query: 80 EKEKTGLANPFGYGNKRIGHNGDV----GWVEYLLLKTNQ-EVNVPVHSQNPEKFRCVLD 134
EK K+ +PF N + W ++L L + E + NP R +
Sbjct: 391 EKLKSYSTDPFKASRLSTSFNVNSEKVSSWRDFLRLHCHPIEDYIKEWPSNPPSLREDVA 570
Query: 135 DYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDK--QSDSVFRVNHYPACPEMGLDGE 192
+Y +R + ++++ ++E L ++ ++ +++++ K Q +N+YPACPE L
Sbjct: 571 EYCRKMRGVSLKLVEAISESLGLE-RDYINRVVGGKKGQEQQHLAMNYYPACPEPELT-- 741
Query: 193 NLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRF 252
G HTDP +I++L + GLQ+ L+DG W++V P ++F +NVGD +QV++N ++
Sbjct: 742 --YGLPGHTDPTVITILLQDEVPGLQV-LKDGKWVAVNPIPNTFVVNVGDQIQVISNDKY 912
Query: 253 RSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYG 312
+SV HR + N + R+S+ F P I P P L+ Y FT+ EY + +
Sbjct: 913 KSVLHRAVVNCNKDRISIPTFYFPSNDAIIGPAPQLIHHHHHPPQYNNFTYNEYYQNFWN 1092
Query: 313 SRLA 316
L+
Sbjct: 1093RGLS 1104
>TC205377
Length = 1345
Score = 137 bits (346), Expect = 5e-33
Identities = 96/311 (30%), Positives = 151/311 (47%), Gaps = 31/311 (9%)
Frame = +3
Query: 27 IPVVDLSK--------PDAKSIIVK----ACEEFGFFKVINHGVSMEAISQLESEALNFF 74
IP++DLS P A +VK AC E+GFF+V NHGV + +E + FF
Sbjct: 165 IPIIDLSPITNHRVSDPSAIEGLVKEIGSACNEWGFFQVTNHGVPLTLRQNIEKASKLFF 344
Query: 75 SMPVTEKEKTGL--ANPFGYGNKRIGHNGDV-GWVEYLLLKTNQEVNVPVHS-------- 123
+ EK K ++P GY + H +V W E + +PV S
Sbjct: 345 AQSAEEKRKVSRNESSPAGYYDTE--HTKNVRDWKEVFDFLAKEPTFIPVTSDEHDDRVN 518
Query: 124 ----QNPE---KFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF 176
Q+PE FR V +Y+ + + +IL+L+A L ++ K V + D+ S
Sbjct: 519 QWTNQSPEYPLNFRVVTQEYIQEMEKLSFKILELIALSLGLEAKRVEEFFIKDQTS--FI 692
Query: 177 RVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLR-DGSWISVPPDYSS 235
R+NHYP CP L +G G H DP +++L + GL++ + D WI V P +
Sbjct: 693 RLNHYPPCPYPDL----ALGVGRHKDPGALTILAQDEVGGLEVRRKADQEWIRVKPTPDA 860
Query: 236 FFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEE 295
+ IN+GD++QV +N + SV HRV+ N + R S+ +F P ++ PL L+ +
Sbjct: 861 YIINIGDTVQVWSNDAYESVDHRVVVNSEKERFSIPFFFFPAHDTEVKPLEELI-NEQNP 1037
Query: 296 SLYKEFTWFEY 306
S Y+ + W ++
Sbjct: 1038SKYRPYKWGKF 1070
>TC215577 homologue to UP|Q41681 (Q41681) 1-aminocylopropane-1-carboxylate
oxidase homolog , complete
Length = 1382
Score = 135 bits (339), Expect = 3e-32
Identities = 92/288 (31%), Positives = 152/288 (51%), Gaps = 9/288 (3%)
Frame = +2
Query: 28 PVVDLSK------PDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
PVVD+ K P A II ACE +GFF+++NHG+S+E + +E + + ++
Sbjct: 62 PVVDMGKLNTEERPAAMEIIKDACENWGFFELVNHGISIELMDTVEKLTKEHYKKTMEQR 241
Query: 82 EKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVR 141
K + + G + + N D+ W L+ NV ++ + +R + + +
Sbjct: 242 FKEMVTSK-GLESVQSEIN-DLDWESTFFLRHLPLSNVSDNADLDQDYRKTMKKFALELE 415
Query: 142 NMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSV-FRVNHYPACPEMGLDGENLIGFGEH 200
+ ++LDL+ E L ++ K L K+ + + +V++YP CP L + G H
Sbjct: 416 KLAEQLLDLLCENLGLE-KGYLKKVFYGSKGPNFGTKVSNYPPCPTPDL----IKGLRAH 580
Query: 201 TDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRV 259
TD II L + + SGLQ+ L+D WI VPP S IN+GD L+V+TNG+++SV HRV
Sbjct: 581 TDAGGIILLFQDDKVSGLQL-LKDDQWIDVPPMRHSIVINLGDQLEVITNGKYKSVMHRV 757
Query: 260 LANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES-LYKEFTWFEY 306
+A +R+S+ F P I+P P+L+K E S +Y +F + +Y
Sbjct: 758 IAQADDTRMSIASFYNPGDDAVISPAPALVKELDETSQVYPKFVFDDY 901
>TC226020 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, partial
(93%)
Length = 1441
Score = 129 bits (323), Expect = 2e-30
Identities = 90/325 (27%), Positives = 165/325 (50%), Gaps = 20/325 (6%)
Frame = +3
Query: 1 MVLLSKQATEQYSYVKNCMATP------CSFTIPVVDLSKPDAKS--------IIVKACE 46
+ L+++ T + S+V++ P S IPV+ L+ D IV+ACE
Sbjct: 84 LTYLAQEKTLESSFVRDEEERPKVAYNEFSDEIPVISLAGIDEVDGRRREICEKIVEACE 263
Query: 47 EFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGLANPFGYG---NKRIGHNGDV 103
+G F+V++HGV + ++++ A FF++P EK + ++ G + +
Sbjct: 264 NWGIFQVVDHGVDQQLVAEMTRLAKEFFALPPDEKLRFDMSGAKKGGFIVSSHLQGESVQ 443
Query: 104 GWVEYLLLKT--NQEVNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKN 161
W E + + +E + PE +R V ++Y V + C+++++++E + ++ K
Sbjct: 444 DWREIVTYFSYPKRERDYSRWPDTPEGWRSVTEEYSDKVMGLACKLMEVLSEAMGLE-KE 620
Query: 162 VLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISL 221
LSK +D V VN+YP CP+ L +G HTDP I+LL + GLQ +
Sbjct: 621 GLSKACVDMDQKVV--VNYYPKCPQPDLT----LGLKRHTDPGTITLLLQDQVGGLQATR 782
Query: 222 RDG-SWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSE 280
+G +WI+V P ++F +N+GD ++NGRF++ H+ + N SRLS+ F P +
Sbjct: 783 DNGKTWITVQPVEAAFVVNLGDHAHYLSNGRFKNADHQAVVNSNHSRLSIATFQNPAPNA 962
Query: 281 KIAPLPSLLKGGKEESLYKEFTWFE 305
+ PL ++ G++ + + T+ E
Sbjct: 963 TVYPLK--IREGEKPVMEEPITFAE 1031
>TC225031 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-carboxylic
acid oxidase, complete
Length = 1244
Score = 127 bits (319), Expect = 7e-30
Identities = 89/290 (30%), Positives = 148/290 (50%), Gaps = 11/290 (3%)
Frame = +1
Query: 28 PVVDLSK------PDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
P+++L K D I ACE +GFF+++NHG+ + + +E + + E+
Sbjct: 124 PLINLEKLSGEERNDTMEKIKDACENWGFFELVNHGIPHDILDTVERLTKEHYRKCMEER 303
Query: 82 EKTGLANPFGYGNKRIGHN-GDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAV 140
K +A+ G + D+ W L+ E N+ +++R V+ D+ +
Sbjct: 304 FKELVASK---GLDAVQTEVKDMDWESTFHLRHLPESNISEIPDLIDEYRKVMKDFALRL 474
Query: 141 RNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSV-FRVNHYPACPEMGLDGENLIGFGE 199
+ ++LDL+ E L ++ K L K + + +V +YP CP + E + G
Sbjct: 475 EKLAEQLLDLLCENLGLE-KGYLKKAFYGSRGPTFGTKVANYPPCP----NPELVKGLRP 639
Query: 200 HTDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHR 258
HTD II L + + SGLQ+ L+DG W+ VPP S +N+GD L+V+TNG+++SV+HR
Sbjct: 640 HTDAGGIILLFQDDKVSGLQL-LKDGQWVDVPPMRHSIVVNIGDQLEVITNGKYKSVEHR 816
Query: 259 VLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEE--SLYKEFTWFEY 306
V+A +R+S+ F P I P P LL+ EE LY +F + +Y
Sbjct: 817 VIAQTDGTRMSIASFYNPGSDAVIYPAPELLEKEAEEKNQLYPKFVFEDY 966
>TC215574 similar to UP|O04076 (O04076) ACC-oxidase, partial (98%)
Length = 1169
Score = 124 bits (312), Expect = 4e-29
Identities = 77/269 (28%), Positives = 140/269 (51%), Gaps = 3/269 (1%)
Frame = +3
Query: 41 IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHN 100
I AC+ +GFF+++NHG+ +E + +E + + ++ K +++ +
Sbjct: 108 IEDACQNWGFFELVNHGIPLELLDTVERLTKEHYRKCMEKRFKEAVSSKGLEAEVK---- 275
Query: 101 GDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPK 160
D+ W L+ N+ +++R + ++ + + E+LDL+ E L ++
Sbjct: 276 -DMDWESTFFLRHLPTSNISEIPDLSQEYRDAMKEFAQKLEKLAEELLDLLCENLGLEKG 452
Query: 161 NVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEHTDPQ-IISLLRSNNTSGLQI 219
+ + + + +V +YPACP+ L + G HTD II LL+ + SGLQ+
Sbjct: 453 YLKNAFYGSRGPNFGTKVANYPACPKPEL----VKGLRAHTDAGGIILLLQDDKVSGLQL 620
Query: 220 SLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLS 279
L++G W+ VPP S +N+GD ++V+TNGR++SV+HRV+A +R+S+ F P
Sbjct: 621 -LKNGQWVDVPPMRHSIVVNLGDQIEVITNGRYKSVEHRVIAQTNGTRMSVASFYNPASD 797
Query: 280 EKIAPLPSLL--KGGKEESLYKEFTWFEY 306
I P P+LL K E +Y +F + +Y
Sbjct: 798 ALIYPAPALLEQKAEDTEQVYPKFVFEDY 884
>TC209692 weakly similar to UP|Q39224 (Q39224) SRG1 protein (F6I1.30/F6I1.30)
(At1g17020/F6I1.30), partial (36%)
Length = 1229
Score = 124 bits (311), Expect = 6e-29
Identities = 79/271 (29%), Positives = 138/271 (50%), Gaps = 5/271 (1%)
Frame = +2
Query: 41 IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGL--ANPFGYGNKRIG 98
+ AC ++GFF+++ HG+S + LE E FF +P+ EK K + + GYG
Sbjct: 200 LTSACRDWGFFQLVEHGISSVVMKTLEDEVEGFFMLPMEEKMKYKVRPGDVEGYGTVIGS 379
Query: 99 HNGDVGWVEYLLLKTNQEV--NVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLK 156
+ + W + L +K N N + + P R +L+ Y+ ++N+ ++ L+ + LK
Sbjct: 380 EDQKLDWGDRLFMKINXRSIRNPHLFPELPSSLRNILELYIEELQNLAMILMGLLGKTLK 559
Query: 157 IQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRS-NNTS 215
I+ + +L + + R+ +YP CP+ L ++G H+D I++L N +
Sbjct: 560 IEKR----ELEVFEDGIQNMRMTYYPPCPQPEL----VMGLSAHSDATGITILNQMNGVN 715
Query: 216 GLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGG 275
GLQI +DG WI V + +N+GD +++M+NG ++SV+HR N + R+S+ F
Sbjct: 716 GLQIK-KDGVWIPVNVISEALVVNIGDIIEIMSNGAYKSVEHRATVNSEKERISVAMFFL 892
Query: 276 PPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
P +I P S L + L+K EY
Sbjct: 893 PKFQSEIGPAVS-LTNPEHPPLFKRIVVEEY 982
>TC215576 similar to UP|O04076 (O04076) ACC-oxidase, partial (97%)
Length = 1151
Score = 123 bits (309), Expect = 1e-28
Identities = 78/269 (28%), Positives = 139/269 (50%), Gaps = 3/269 (1%)
Frame = +2
Query: 41 IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHN 100
I AC+ +GFF+++NHG+ +E + +E + + + K +A+ +
Sbjct: 59 IEDACQNWGFFELVNHGIPLELLDTVERLTKEHYRKCMENRFKEAVASKALEVEVK---- 226
Query: 101 GDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPK 160
D+ W L+ N+ +++R + ++ + + E+LDL+ E L ++
Sbjct: 227 -DMDWESTFFLRHLPTSNISEIPDLSQEYRDAMKEFAKKLEKLAEELLDLLCENLGLEKG 403
Query: 161 NVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEHTDPQ-IISLLRSNNTSGLQI 219
+ + K + +V +YPACP+ L + G HTD II LL+ + SGLQ+
Sbjct: 404 YLKNAFYGSKGPNFGTKVANYPACPKPEL----VKGLRAHTDAGGIILLLQDDKVSGLQL 571
Query: 220 SLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLS 279
L+D W+ VPP S +N+GD ++V+TNGR++SV+HRV+A +R+S+ F P
Sbjct: 572 -LKDDQWVDVPPMRHSIVVNLGDQIEVITNGRYKSVEHRVVARTDGTRMSVASFYNPAND 748
Query: 280 EKIAPLPSLL--KGGKEESLYKEFTWFEY 306
I P P+LL + + E +Y +F + +Y
Sbjct: 749 AVIYPAPALLEKEAQETEQVYPKFVFEDY 835
>TC216188 similar to UP|Q84L58 (Q84L58) 1-aminocyclopropane-1-carboxylic acid
oxidase, complete
Length = 1362
Score = 122 bits (306), Expect = 2e-28
Identities = 87/297 (29%), Positives = 147/297 (49%), Gaps = 17/297 (5%)
Frame = +3
Query: 27 IPVVDLSKPDAK------SIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTE 80
+PV+D SK + + + I CEE+GFF++INHG+ E + +++ A F+ + E
Sbjct: 147 VPVIDFSKLNGEERTKTMAQIANGCEEWGFFQLINHGIPEELLERVKKVASEFYKLEREE 326
Query: 81 --KEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEK---FRCVLDD 135
K T + K+ V W + + L + E PEK FR + +
Sbjct: 327 NFKNSTSVKLLSDSVEKKSSEMEHVDWEDVITLLDDNEW--------PEKTPGFRETMAE 482
Query: 136 YLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF---RVNHYPACPEMGLDGE 192
Y ++ + +++++M E L + K + K L ++ F +V+HYP CP L
Sbjct: 483 YRAELKKLAEKLMEVMDENLGLT-KGYIKKALNGGDGENAFFGTKVSHYPPCPHPEL--- 650
Query: 193 NLIGFGEHTDPQ-IISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGR 251
+ G HTD +I L + + GLQ+ L++G WI V P ++ IN GD ++V++NGR
Sbjct: 651 -VKGLRAHTDAGGVILLFQDDKVGGLQM-LKEGQWIDVQPLPNAIVINTGDQIEVLSNGR 824
Query: 252 FRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKE--ESLYKEFTWFEY 306
++S HRVLA +R S+ F P I P P L++ + + Y +F + +Y
Sbjct: 825 YKSCWHRVLATPDGNRRSIASFYNPSFKATICPAPQLVEKEDQQVDETYPKFVFGDY 995
>TC228068 anthocyanidin synthase [Glycine max]
Length = 1231
Score = 116 bits (290), Expect = 2e-26
Identities = 79/283 (27%), Positives = 146/283 (50%), Gaps = 20/283 (7%)
Frame = +3
Query: 27 IPVVDLSKPDAKSIIV---------KACEEFGFFKVINHGVSMEAISQLESEALNFFSMP 77
+P +DL + D++ +V KA EE+G ++NHG+ E I +++ FF +
Sbjct: 141 VPTIDLREIDSEDEVVRGKCREKLKKAAEEWGVMHLVNHGIPDELIERVKKAGETFFGLA 320
Query: 78 VTEKEKTGLANPF------GYGNKRIGH-NGDVGWVEYL--LLKTNQEVNVPVHSQNPEK 128
V EKEK AN GYG+K + +G + W +Y L + ++ + P
Sbjct: 321 VEEKEK--YANDLESGKIQGYGSKLANNASGQLEWEDYFFHLAFPEDKRDLSFWPKKPAD 494
Query: 129 FRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMG 188
+ V +Y +R + +IL+ ++ GL ++ + + ++ ++ ++N+YP CP+
Sbjct: 495 YIEVTSEYAKRLRGLATKILEALSIGLGLEGRRLEKEVGGMEELLLQLKINYYPICPQPE 674
Query: 189 LDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMT 248
L +G HTD ++ L N GLQ+ + G W++ +S +++GD++++++
Sbjct: 675 L----ALGVEAHTDVSSLTFLLHNMVPGLQLFYQ-GQWVTAKCVPNSILMHIGDTIEILS 839
Query: 249 NGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKI--APLPSLL 289
NG+++S+ HR L N + R+S F PP EKI PLP L+
Sbjct: 840 NGKYKSILHRGLVNKEKVRISWAVFCEPP-KEKIILQPLPELV 965
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,438,575
Number of Sequences: 63676
Number of extensions: 194319
Number of successful extensions: 1088
Number of sequences better than 10.0: 206
Number of HSP's better than 10.0 without gapping: 985
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 990
length of query: 329
length of database: 12,639,632
effective HSP length: 98
effective length of query: 231
effective length of database: 6,399,384
effective search space: 1478257704
effective search space used: 1478257704
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0190.3


