

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
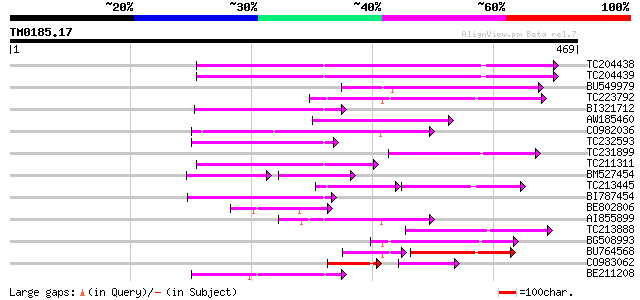
Query= TM0185.17
(469 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 166 2e-41
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 164 1e-40
BU549979 102 5e-22
TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polypro... 92 6e-19
BI321712 92 6e-19
AW185460 89 5e-18
CO982036 85 8e-17
TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (F... 78 1e-14
TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Frag... 75 8e-14
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 74 1e-13
BM527454 weakly similar to GP|27901709|gb| gag-pol polyprotein {... 48 2e-13
TC213445 50 7e-13
BI787454 weakly similar to GP|21434|emb|CA ORF4 {Solanum tuberos... 70 3e-12
BE802806 67 2e-11
AI855899 similar to GP|2244960|emb| retrotransposon like protein... 65 8e-11
TC213888 similar to UP|Q9SFE1 (Q9SFE1) T26F17.17, partial (11%) 64 1e-10
BG508993 62 4e-10
BU764568 53 1e-09
CO983062 47 3e-09
BE211208 58 1e-08
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 166 bits (420), Expect = 2e-41
Identities = 99/302 (32%), Positives = 164/302 (53%), Gaps = 2/302 (0%)
Frame = +1
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSHYIEK 214
+YVDD++ G + + + F+M +GE LG+++ + + I LSQS Y +
Sbjct: 3781 IYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSKYAKN 3960
Query: 215 VLKKFDHFDCKSVSTPFDQNTKLQPHKR-CPVAQLEYSKAIRCLMYAMTCTRPDIAYVVG 273
++KKF + TP + KL + V Q Y I L+Y +T +RPDI Y VG
Sbjct: 3961 IVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLY-LTASRPDITYAVG 4137
Query: 274 RLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIY-NGYPSVLEGYTYASWVTCVKDHAST 332
+RY NP H + V R+LKY+NGT +Y ++Y + S+L GY A W D ST
Sbjct: 4138 VCARYQANPKISHLNQVKRILKYVNGTSDYGIMYCHCSDSMLVGYCDADWAGSADDRKST 4317
Query: 333 SG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKLMRQ 392
SG F LG +SW SKKQ C++ ST AE+IA + + V ++ +L E + +M
Sbjct: 4318 SGGCFYLGTNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVM-- 4491
Query: 393 VSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKESCRSFDER 452
+++CD+ ++ + + V + +++HI +RH +++DL+ + VI++ V T ++ F +
Sbjct: 4492 -TLYCDNMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLEHVDTEEQIADIFTKA 4668
Query: 453 SD 454
D
Sbjct: 4669 LD 4674
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 164 bits (414), Expect = 1e-40
Identities = 98/302 (32%), Positives = 162/302 (53%), Gaps = 2/302 (0%)
Frame = +1
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSHYIEK 214
+YVDD++ G + + + F+M +GE LG+++ + + I LSQS Y +
Sbjct: 3778 IYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSRYAKN 3957
Query: 215 VLKKFDHFDCKSVSTPFDQNTKLQPHKR-CPVAQLEYSKAIRCLMYAMTCTRPDIAYVVG 273
++KKF + TP + KL + V Q Y I L+Y +T +RPDI Y VG
Sbjct: 3958 IVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLY-LTASRPDITYAVG 4134
Query: 274 RLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPS-VLEGYTYASWVTCVKDHAST 332
+RY NP H V R+LKY+NGT +Y ++Y + +L GY A W D ST
Sbjct: 4135 VCARYQANPKISHLTQVKRILKYVNGTSDYGIMYCHCSNPMLVGYCDADWAGSADDRKST 4314
Query: 333 SG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKLMRQ 392
SG F LG +SW SKKQ C++ ST AE+IA + + V ++ +L E + +M
Sbjct: 4315 SGGCFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVM-- 4488
Query: 393 VSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKESCRSFDER 452
+++CD+ ++ + + V + +++HI +RH +++DL+ + VI++ V T ++ F +
Sbjct: 4489 -TLYCDNMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLKHVDTEEQIADIFTKA 4665
Query: 453 SD 454
D
Sbjct: 4666 LD 4671
>BU549979
Length = 615
Score = 102 bits (253), Expect = 5e-22
Identities = 56/169 (33%), Positives = 89/169 (52%), Gaps = 2/169 (1%)
Frame = -1
Query: 275 LSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSVLE--GYTYASWVTCVKDHAST 332
L RY NP DHW +V++YL GT +Y L+Y + LE GY+ + + CV ST
Sbjct: 612 LGRYQSNPGIDHWKTAKKVMRYLQGTKDYMLMYK-QTNCLEVIGYSDSDFAGCVDSRRST 436
Query: 333 SG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKLMRQ 392
SG IF L G VSW S KQT IA STM EF+ + V L++ + + + + R
Sbjct: 435 SGYIFMLADGVVSWRSSKQTLIATSTMEVEFVPCFEATSHGVWLKSFMSSLRVVDSISRP 256
Query: 393 VSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
+ ++CD+ + A + +S+HI +++ +++ + + I V T
Sbjct: 255 LKLYCDNFAAVFMAKNNKSGNRSKHIDIKYLVIRERVKEKKVVIEHVNT 109
>TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein,
partial (7%)
Length = 804
Score = 91.7 bits (226), Expect = 6e-19
Identities = 57/201 (28%), Positives = 106/201 (52%), Gaps = 5/201 (2%)
Frame = +1
Query: 249 EYSKAIRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIY- 307
E+ + I L Y +RP+I + V +SR+ P H RVL+ + GTI +++
Sbjct: 19 EFRRLIGSLRYLCN-SRPNICFAVSLISRFMKRPRLSHMQAAKRVLRLIKGTIGSGVLFP 195
Query: 308 ----NGYPSVLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEF 363
+G P +L GYT + W + ST G +F V+ SKKQ IA ST AE+
Sbjct: 196 FKAKSGKPDLL-GYTDSDWKRDPEQEKSTGGYLFMYNDAPVA*SSKKQDVIALSTCEAEY 372
Query: 364 IALAACSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHS 423
+A + + + V + NLL E+ + + + V++ D++ ++ A +G+S+HI LR
Sbjct: 373 VAASLGACQAVWMMNLLEELKL--RERKPVNLLIDNKSAINLAKHPTLHGRSKHIELRFH 546
Query: 424 HVKDLITNGVISIVFVRTVKE 444
+++D ++ G +++ + + ++
Sbjct: 547 YIRDQVSKGNVTVEYCKAEEQ 609
>BI321712
Length = 399
Score = 91.7 bits (226), Expect = 6e-19
Identities = 52/126 (41%), Positives = 75/126 (59%), Gaps = 1/126 (0%)
Frame = -3
Query: 154 CLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSHYIE 213
CLYVDD++ G + + E+ K +SN F+M D+G LGI++ ++ + I ++Q Y +
Sbjct: 382 CLYVDDLIFTGNNPSMFEEFKKDMSNEFEMTDMGLMAYYLGIEVKQEDKGIFITQEGYAK 203
Query: 214 KVLKKFDHFDCKSVSTPFDQNTKLQPHKRCP-VAQLEYSKAIRCLMYAMTCTRPDIAYVV 272
+VLKKF D V TP + +KL H++ V Y I L Y +TCTRPDI YVV
Sbjct: 202 EVLKKFKMDDANPVGTPMECGSKLSKHEKGENVDPTLYKSLIGSLRY-LTCTRPDILYVV 26
Query: 273 GRLSRY 278
G +SRY
Sbjct: 25 GVVSRY 8
>AW185460
Length = 411
Score = 88.6 bits (218), Expect = 5e-18
Identities = 50/118 (42%), Positives = 65/118 (54%), Gaps = 1/118 (0%)
Frame = +2
Query: 251 SKAIRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGY 310
SK+ L TRPDI Y LSR+ +PS+ H+ R+L+YL GT + + Y
Sbjct: 53 SKSEPTLYIKSQATRPDIMYATSLLSRFMQSPSQIHFGAGKRILRYLQGTKAFGIWYTTE 232
Query: 311 P-SVLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALA 367
S L GYT + W D STSG F+LG G SW SKKQ +A ST AE++A+A
Sbjct: 233 TNSELLGYTDSDWAGSTDDMKSTSGYAFSLGSGMFSWASKKQATVAQSTAEAEYVAVA 406
>CO982036
Length = 674
Score = 84.7 bits (208), Expect = 8e-17
Identities = 54/205 (26%), Positives = 99/205 (47%), Gaps = 4/205 (1%)
Frame = -2
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSH 210
V +YVD ++I G+ ++ S L++SF +K LG+ D + I++ +++ ++
Sbjct: 652 VYLLVYVD-IIITGSSCTLIQNLTSKLNSSFPLKLLGKLDYFVEIEVKSMPDLLFSLRTS 476
Query: 211 YIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLEYSKAIRCLMYAMTCTRPDIAY 270
E +K + +S+P KL + + +++ + T RP+I++
Sbjct: 475 IFEIFCRK-PR*QAQPISSPMTTTCKLSKSDSDLFSGPTFYRSVVGALQYTTVIRPEISF 299
Query: 271 VVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSL----IYNGYPSVLEGYTYASWVTCV 326
V ++ ++ NP HW V R+L+YL G+++Y L + P + G+ A W + V
Sbjct: 298 AVNKVCQFMSNPLDSHWTEVKRILRYLKGSLSYGL*LKPAISSQPLPIRGFCDADWASAV 119
Query: 327 KDHASTSG*IFNLGGGAVSWGSKKQ 351
D STSG LG +SW KQ
Sbjct: 118 DDKRSTSGAAVFLGPNLISWWXXKQ 44
>TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (Fragment),
partial (16%)
Length = 562
Score = 77.8 bits (190), Expect = 1e-14
Identities = 45/123 (36%), Positives = 68/123 (54%), Gaps = 1/123 (0%)
Frame = +1
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSH 210
++ LYVDD+L+ D VE+ K + +F+M +LG LGI+I + + + Q
Sbjct: 193 LIVSLYVDDLLVTRDDARLVEEFKQEMMQAFEMTNLGLMTYFLGIEIKQSQNKVLICQRK 372
Query: 211 YIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRC-PVAQLEYSKAIRCLMYAMTCTRPDIA 269
Y +++LKKF +CKSVSTP +Q K + + Y I CLMY +T TRPDI
Sbjct: 373 YAKEILKKFQMEECKSVSTPMNQKEKFNKVDGADKIDEGYYRSLIGCLMY-LTATRPDIL 549
Query: 270 YVV 272
+ +
Sbjct: 550 FAI 558
>TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Fragment),
partial (30%)
Length = 687
Score = 74.7 bits (182), Expect = 8e-14
Identities = 41/126 (32%), Positives = 70/126 (55%)
Frame = +2
Query: 314 LEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKET 373
L GY A W C D STSG +GG VSW SKKQT +A S+ AE+ ++A + E
Sbjct: 17 LSGYCDADWAGCPMDRRSTSGYCVFIGGNLVSWKSKKQTVVARSSAEAEYRSMAMVTCEL 196
Query: 374 V*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGV 433
+ ++ L E+ +L Q+ ++CD+Q L A + V++ +++HI + +++ + +
Sbjct: 197 MWIKQFLQELRFCEEL--QMKLYCDNQAALHIASNPVFHERTKHIEIDCHFIREKLLSKE 370
Query: 434 ISIVFV 439
I F+
Sbjct: 371 IVTEFI 388
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 73.9 bits (180), Expect = 1e-13
Identities = 48/152 (31%), Positives = 79/152 (51%), Gaps = 1/152 (0%)
Frame = +3
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSHYIEK 214
+YVDD++ T ++ + + F+ GE +LG++II+ I + Q Y +
Sbjct: 501 IYVDDIIFGATSKRMCKEFFELMKDGFETSMKGELKFLLGLQIIQKVYGIFIHQEKYTKS 680
Query: 215 VLKKFDHFDCKSVSTPFDQNTKL-QPHKRCPVAQLEYSKAIRCLMYAMTCTRPDIAYVVG 273
LK+F + K ++TP ++T + + K + EYS I L Y +T +RPDI +VV
Sbjct: 681 HLKRFRMDEAKPMATPMHRSTIIDKDEKGNHTS*KEYSGMIDSLSY-LTSSRPDIVFVVC 857
Query: 274 RLSRYTCNPSKDHWHVVNRVLKYLNGTINYSL 305
+R+ P H V R+L+YL GT N+ L
Sbjct: 858 LCARFQSYPKISHVTAVKRILRYLVGTTNHCL 953
>BM527454 weakly similar to GP|27901709|gb| gag-pol polyprotein {Vitis
vinifera}, partial (19%)
Length = 437
Score = 47.8 bits (112), Expect(2) = 2e-13
Identities = 24/70 (34%), Positives = 40/70 (56%)
Frame = +2
Query: 147 SGKAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGL 206
S + V +YVDD++I G D ++ + K L + F KDLG+ + LGI++ + + I +
Sbjct: 23 SSRCVYLMVYVDDIVITGNDQGKIAQLKGHLFSHFQTKDLGKFEYFLGIEVAQSKDGIII 202
Query: 207 SQSHYIEKVL 216
SQ Y +L
Sbjct: 203 SQRKYALDIL 232
Score = 45.8 bits (107), Expect(2) = 2e-13
Identities = 21/64 (32%), Positives = 38/64 (58%)
Frame = +1
Query: 223 DCKSVSTPFDQNTKLQPHKRCPVAQLEYSKAIRCLMYAMTCTRPDIAYVVGRLSRYTCNP 282
DC+ + + D N KL P++ P + E + + + +T TRP+I++VVG +S++ +P
Sbjct: 241 DCRPIDSLMDPNKKLLPNQGKPYSDSERYRILVGKLIYLTITRPNISFVVGVVSQFMQSP 420
Query: 283 SKDH 286
DH
Sbjct: 421 HNDH 432
>TC213445
Length = 705
Score = 49.7 bits (117), Expect(2) = 7e-13
Identities = 27/102 (26%), Positives = 51/102 (49%)
Frame = +1
Query: 325 CVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEIL 384
C D STS +G VSW SKKQ + ST AE+I+ + + +R L++
Sbjct: 397 CKTDRESTSDTCHFIGSALVSWHSKKQNSVVLSTAEAEYISARSYYAQIFWMRQQLFD-- 570
Query: 385 IWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVK 426
+ + + I CD+ ++ + + + +++HI +RH ++
Sbjct: 571 -YGLKLDHIPIRCDNTSAINLSKNHILYSRTKHIEIRHHFLR 693
Score = 42.0 bits (97), Expect(2) = 7e-13
Identities = 26/71 (36%), Positives = 36/71 (50%), Gaps = 1/71 (1%)
Frame = +2
Query: 254 IRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV 313
I +Y T +RP I + V RY NP + H V+ R+++YL G IN L Y S
Sbjct: 200 IESFLYLST-SRPHIMFSVCMCVRYQANPKESHLSVIKRIMRYLLGIINLGLWYPKNSSY 376
Query: 314 -LEGYTYASWV 323
L GY+ A +
Sbjct: 377 NLVGYSDAKLI 409
>BI787454 weakly similar to GP|21434|emb|CA ORF4 {Solanum tuberosum}, partial
(21%)
Length = 421
Score = 69.7 bits (169), Expect = 3e-12
Identities = 41/124 (33%), Positives = 67/124 (53%), Gaps = 1/124 (0%)
Frame = +2
Query: 148 GKAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLS 207
GK V +YVDD++I D ++ + K L N F KDL LGI++ + G+ + +S
Sbjct: 50 GKCVYLMVYVDDIMITKKDATKIVQLKEHLFNHFQTKDLRYLKYFLGIEVAQSGDGVVIS 229
Query: 208 QSHYIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQLE-YSKAIRCLMYAMTCTRP 266
Q Y +L++ +C+ V +P D N KL ++ E Y + + L+Y +T TRP
Sbjct: 230 QRKYALDILEETGMQNCRLVDSPMDPNLKLMAYQSEVYPDPERYRRLVGKLIY-LTITRP 406
Query: 267 DIAY 270
DI++
Sbjct: 407 DISF 418
>BE802806
Length = 285
Score = 67.0 bits (162), Expect = 2e-11
Identities = 41/95 (43%), Positives = 55/95 (57%), Gaps = 10/95 (10%)
Frame = -2
Query: 183 MKDLGEADVILGIKIIRD---GEIIGLSQSHYIEKVLKKFDHFDCKSVSTPFDQNTKLQ- 238
MKDLG A ILGI I RD GE+ LSQS+Y++KV+++F K VSTP +TKL
Sbjct: 284 MKDLGSARRILGIDIHRDRAKGELF-LSQSNYLKKVVERFRMHQSKPVSTPLGHHTKLSV 108
Query: 239 ------PHKRCPVAQLEYSKAIRCLMYAMTCTRPD 267
+R + Q Y+ + +MY M C+RPD
Sbjct: 107 TQAPETAEERSKMNQTPYANGVGSIMYGMVCSRPD 3
>AI855899 similar to GP|2244960|emb| retrotransposon like protein
{Arabidopsis thaliana}, partial (18%)
Length = 418
Score = 64.7 bits (156), Expect = 8e-11
Identities = 43/135 (31%), Positives = 66/135 (48%), Gaps = 6/135 (4%)
Frame = +1
Query: 223 DCKSVSTPFDQNTKLQPH--KRCPVAQLEYSKAIRCLMYAMTCTRPDIAYVVGRLSRYTC 280
DC +STP + KL + P A +Y + L Y +T TRP+IAY V ++S +
Sbjct: 19 DCNGISTPMVSSYKLSKFGSELLPNAH-QYRDIVGALQY-VTLTRPNIAYNVNKVSEFMS 192
Query: 281 NPSKDHWHVVNRVLKYLNGTINYSLI----YNGYPSVLEGYTYASWVTCVKDHASTSG*I 336
+P + + V R+L+YL+GT+ L+ + L Y W + + STSG
Sbjct: 193 SPLQSY*LTVKRILRYLSGTVTQGLLLQPAHMDAKISLRAYNDLDWGSDPAEMRSTSGSC 372
Query: 337 FNLGGGAVSWGSKKQ 351
G ++W SKKQ
Sbjct: 373 IFSGSNLIAWSSKKQ 417
>TC213888 similar to UP|Q9SFE1 (Q9SFE1) T26F17.17, partial (11%)
Length = 493
Score = 64.3 bits (155), Expect = 1e-10
Identities = 36/122 (29%), Positives = 68/122 (55%)
Frame = +3
Query: 328 DHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWP 387
D ST+G +F +G A +W SKKQ + ST AE++A +C + LRNLL E+ +
Sbjct: 12 DRKSTTGFVFFMGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELKMPQ 191
Query: 388 KLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKESCR 447
+ ++ + D++ L+ A + V++ KS+HI R+ +++ I + + +V + ++
Sbjct: 192 EEPMEICV--DNKSALALAKNPVFHEKSKHIDTRYHFIRECIEKKEVKLKYVMSQDQAAD 365
Query: 448 SF 449
F
Sbjct: 366 IF 371
>BG508993
Length = 374
Score = 62.4 bits (150), Expect = 4e-10
Identities = 40/126 (31%), Positives = 65/126 (50%), Gaps = 3/126 (2%)
Frame = +1
Query: 299 GTINYSLIY---NGYPSVLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIA 355
GTI++ L Y N Y V G+ + + V D ST+G +F +G +W SKKQ +
Sbjct: 7 GTIDFGLFYSPSNNYKLV--GFCDSDFAGDVDDRKSTTGFVFFMGDCVFTWSSKKQGIVT 180
Query: 356 DSTMAAEFIALAACSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKS 415
T AE++A +C+ + LR LL E+ + K I+ D++ A + V++ +S
Sbjct: 181 LFTCEAEYVAATSCTCHAIWLRRLLEELQLLQK--ESTKIYVDNRSAQELAKNSVFHERS 354
Query: 416 RHIVLR 421
+HI R
Sbjct: 355 KHIDTR 372
>BU764568
Length = 420
Score = 53.1 bits (126), Expect(2) = 1e-09
Identities = 30/87 (34%), Positives = 53/87 (60%)
Frame = +3
Query: 332 TSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKLMR 391
TSG +GG +SW SKKQ+ +A S+ AE+ A+A + E + L+ LL E+ +
Sbjct: 165 TSGYCVLIGGNLISWKSKKQSVVAKSSAEAEYRAMALVTCELIWLKQLL*ELKF--EEDT 338
Query: 392 QVSIHCDSQCTLSKAYSQVYNGKSRHI 418
Q+++ CD+Q L A + +++ +++HI
Sbjct: 339 QMTLICDNQAALHIASNPIFH*RTKHI 419
Score = 27.3 bits (59), Expect(2) = 1e-09
Identities = 17/55 (30%), Positives = 28/55 (50%), Gaps = 2/55 (3%)
Frame = +1
Query: 276 SRYTCNPSKDHWHVVNRVLKYLNGTINYSLIY--NGYPSVLEGYTYASWVTCVKD 328
S++ +P +DHW+ V+ +LK LIY G+ ++ GY+ A V D
Sbjct: 1 SQFLNSPCQDHWNAVS*ILK*TKSAPGKGLIYEDKGHSQII-GYSDAD*VGSPSD 162
>CO983062
Length = 390
Score = 47.0 bits (110), Expect(2) = 3e-09
Identities = 20/44 (45%), Positives = 29/44 (65%)
Frame = +2
Query: 264 TRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIY 307
TRP+I YVV ++ +Y NP HW VV R+L+YL + Y ++Y
Sbjct: 44 TRPEINYVVNKVXQYMTNPLDSHWAVVKRILRYLK--VPYFMVY 169
Score = 32.3 bits (72), Expect(2) = 3e-09
Identities = 19/51 (37%), Positives = 24/51 (46%)
Frame = +3
Query: 322 WVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKE 372
W + + D STS G +SW SKKQ A S+ AE+ L S E
Sbjct: 228 WASNIDDRRSTSRAAIFPGRNLISWWSKKQKVTARSSTEAEYQILVQTSIE 380
>BE211208
Length = 413
Score = 57.8 bits (138), Expect = 1e-08
Identities = 38/131 (29%), Positives = 69/131 (52%), Gaps = 3/131 (2%)
Frame = +2
Query: 151 VMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKI--IRDGEIIGLSQ 208
V +YVDD++I G ++ L+++F +K LG+ D LGI++ G ++ L+Q
Sbjct: 26 VYLLVYVDDIIITGRSNYLIQSLVHHLNSNFSLKQLGQLDYFLGIEVHHTPTGSVL-LTQ 202
Query: 209 SHYIEKVLKKFDHFDCKSVSTPFDQNTKLQPHKRCPVAQ-LEYSKAIRCLMYAMTCTRPD 267
S YI +L K D + K +S+P N +L + ++ Y + L Y T TRP+
Sbjct: 203 SKYICDLLHKTDMAEAKPISSPMVTNLRLSKNGDDLLSDPTMYRSVVGALQYP-TITRPE 379
Query: 268 IAYVVGRLSRY 278
I++ ++ ++
Sbjct: 380 ISFAANKVCQF 412
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.351 0.154 0.546
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,451,847
Number of Sequences: 63676
Number of extensions: 338869
Number of successful extensions: 2813
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 2767
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2788
length of query: 469
length of database: 12,639,632
effective HSP length: 101
effective length of query: 368
effective length of database: 6,208,356
effective search space: 2284675008
effective search space used: 2284675008
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0185.17


