

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.7
(1813 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
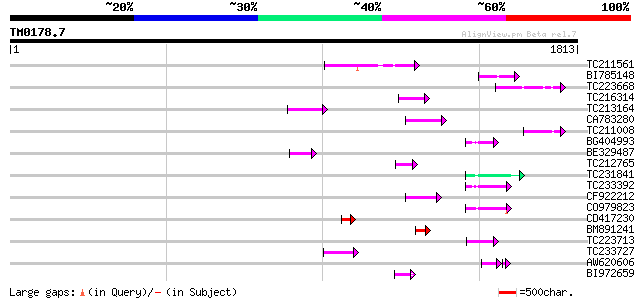
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 118 3e-26
BI785148 homologue to GP|25166165|dbj murF {Wigglesworthia brevi... 95 2e-19
TC223668 80 1e-14
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 75 2e-13
TC213164 74 6e-13
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 72 2e-12
TC211008 68 4e-11
BG404993 60 7e-09
BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase... 60 1e-08
TC212765 60 1e-08
TC231841 59 1e-08
TC233392 58 3e-08
CF922212 58 4e-08
CO979823 58 4e-08
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 58 4e-08
BM891241 57 6e-08
TC223713 57 7e-08
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 57 1e-07
AW620606 50 2e-07
BI972659 55 2e-07
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase (Fragment)
, partial (59%)
Length = 1077
Score = 118 bits (295), Expect = 3e-26
Identities = 97/320 (30%), Positives = 152/320 (47%), Gaps = 15/320 (4%)
Frame = +1
Query: 1006 MDNAFLAQEIIHHMGASMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIM 1065
+ +A +A E+I A + K L FK+D EKAYDSVSW FL + + F + ++ I
Sbjct: 1 LHSALIANEVIDE--AKRSNKSCLVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIE 174
Query: 1066 RCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLF--VLCSLHGAI-------VY*YS 1116
CV +++S+L NGS F P RGLR+GDPL+P+LF V L+G + +Y
Sbjct: 175 ECVKSASISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRRAVEENLYKPY 354
Query: 1117 *VGGNRSLEAN*NFPKGTTYF---ASLFCR*RALVLSSIIFSSAIGGGDS*TILCLFGVK 1173
VG N + + T +F A ++L S S + + + +FGV
Sbjct: 355 LVGANGVPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKSCFGVFGVT 534
Query: 1174 DKYEQI*SDCFQGSTKSCERGDPEYC---PYPFCAKPWQVSWLEEGFQEVDSITYWIIFK 1230
D+++Q E + +C +PF + DSI +
Sbjct: 535 DQWKQ-------------EAANYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKC--E 669
Query: 1231 KKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQ 1290
+KLS WK ++ G+V L KSV+++IP Y F +P+S+ K+ + R F+W Q
Sbjct: 670 RKLSKWK*RHISFGGRVTLIKSVLTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGGGPDQ 849
Query: 1291 RGWHRVNWKTLTQPKEHGGL 1310
+ + W+ L PKE GG+
Sbjct: 850 KKISWIRWEKLCLPKERGGI 909
>BI785148 homologue to GP|25166165|dbj murF {Wigglesworthia brevipalpis},
partial (2%)
Length = 422
Score = 95.1 bits (235), Expect = 2e-19
Identities = 47/131 (35%), Positives = 73/131 (54%)
Frame = -2
Query: 1500 PEKL*VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELWGKL 1559
P+K+ + WL+LH++L N+ R+ L S CS+P ED++HCLRD PH++E+W
Sbjct: 418 PKKIRIMFWLILHNSLSKNSTRFHRNLVNSSAFPHCSNPREDSLHCLRDFPHAKEIWLGF 239
Query: 1560 GAWGWSTFRLPDLKA*VKTQLQGSNNIRFLAGLWGVWKWRNNVVLDPLPWTLDGAWQKLR 1619
G + LP + + +K+QL GS F +W V KW NN + D W + Q +
Sbjct: 238 G-FAPHLSSLPLM*SGLKSQLNGSKVFLFTTIVWWV*KWGNNCIFDDEKWDIQRVVQSIC 62
Query: 1620 HDHDELLQFSS 1630
HD+L+ +S
Sbjct: 61 FYHDDLVSTNS 29
>TC223668
Length = 866
Score = 79.7 bits (195), Expect = 1e-14
Identities = 73/230 (31%), Positives = 97/230 (41%), Gaps = 4/230 (1%)
Frame = -3
Query: 1552 SQELWGKLGAWGWSTFRLPDLKA*VKTQLQGSNNIRFLAGLWGVWKWRNNVVLDPLPWTL 1611
++ELW ++ S +L A L ++ + W WKW N V+ PL W+
Sbjct: 852 TRELWLRVSNGNGSLLSHTNLFAXXXAVLSNNDCAYKVLEAWWAWKWCN*VIFSPLIWSF 673
Query: 1612 DGAWQKLRHDHDELLQFS--SGDLLDEHHFLINLWKPPPPEF--VKLNTDGSFQDTASSM 1667
+ K H LQ S S DL+ W PPP VKLN DGS M
Sbjct: 672 Q--FIKKIHC*GYCLQSSLVSKDLVSSKRLT---WTSPPPLLNEVKLNVDGSGHYEPHVM 508
Query: 1668 GGGGLIRDAKGAWLGGFMSHTSIGNPFLAEVLALRDGLCLAWVQGHRRIIREVDCAGLSD 1727
GGLIRD G WL F H +GNP L+E+ AL L + H + C +
Sbjct: 507 EVGGLIRDGVGRWLFRFARHCGLGNPLLSELFALEASL-----RSHI-V*TWWSCLLIGQ 346
Query: 1728 ILRFEDRCRLHDHAGVLQEIRELMERPWRCSVHWTNRDSNASADFLARRG 1777
+ R C L ++ IR M+R W S+ N N+ DFLA G
Sbjct: 345 MSR*A--CMLMQI--LIHSIRSFMKRDWSFSLCHVNTKVNSVVDFLAEWG 208
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 75.5 bits (184), Expect = 2e-13
Identities = 37/99 (37%), Positives = 57/99 (57%)
Frame = +2
Query: 1243 MVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQRGWHRVNWKTLT 1302
M G+V L KSV+SA+P + F +P+ I K+ R+F+W V W +
Sbjct: 2 MGGRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADIC 181
Query: 1303 QPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQV 1341
PK GGL IKD+ +FN +L G+ +W LA+N ++LW ++
Sbjct: 182 NPKIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWARL 298
>TC213164
Length = 446
Score = 73.9 bits (180), Expect = 6e-13
Identities = 38/128 (29%), Positives = 69/128 (53%)
Frame = +3
Query: 889 KEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLI 948
K+EV + + + GPDG + F K++W+++ DL + + + + G+ +
Sbjct: 36 KKEVRATMWDRGNAKSPGPDGLNFKFIKEFWDILKTDLLRFLDEFYANGVFLKRSNAFFL 215
Query: 949 VLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDN 1008
LIPKV P ++ E++PISL +YK++ K++ R + +L II Q F+ G + N
Sbjct: 216 ALIPKVHDP*SLNEYKPISLIGCIYKIVAKLLSGRHKKVLPTIIDERQTVFMEGRHTLHN 395
Query: 1009 AFLAQEII 1016
+A EI+
Sbjct: 396 VVIANEIM 419
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase homolog
T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 72.0 bits (175), Expect = 2e-12
Identities = 48/139 (34%), Positives = 68/139 (48%), Gaps = 7/139 (5%)
Frame = +2
Query: 1265 FWLPRSINHKINQEMRRFIWLKNSGQRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLG 1324
F LP I K+ RRF+W + R VNWKT+ PK GGL IKD+ FNT+LLG
Sbjct: 11 FSLP*KIIAKLESLQRRFLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTTLLG 190
Query: 1325 KVVWLLANNSSKLWVQVLAHKYLHGESIFSVHRRANASPIWQGILKARDQLQEGFQ---- 1380
K W L + W +VL KY ++ + S W+ ++K + QLQ
Sbjct: 191 KWRWDLFYIQQEPWAKVLQSKYGGWRALEEGSSGSKDSAWWKDLIKTQ-QLQRNIPLKRE 367
Query: 1381 --FRMGDGSSSVWYHD-WT 1396
+++G G ++ D WT
Sbjct: 368 TIWKVGGGDRIKFWEDLWT 424
>TC211008
Length = 516
Score = 67.8 bits (164), Expect = 4e-11
Identities = 47/139 (33%), Positives = 73/139 (51%), Gaps = 5/139 (3%)
Frame = +1
Query: 1644 WKPPPPEFVKLNTDGS-FQDTASSMGG---GGLIRDAKGAWLGGFMSHTSIGNPFLAEVL 1699
W+ P +V LNTDGS F++ + G GGL+RD+ G +LGGF + + LAE+
Sbjct: 64 WRCPXNGWVCLNTDGSVFENHRNGCSGSACGGLVRDSSGCYLGGFTVNLGNTSVTLAELW 243
Query: 1700 ALRDGLCLAWVQGHRRIIREVDCAGLSDILRFEDRCRLHDHA-GVLQEIRELMERPWRCS 1758
+ GL LAW G +++ ++D ++R +D A ++ EI EL+ + W
Sbjct: 244 GVVHGLKLAWDLGCKKVKVDIDSGNALGLVRHGPVA--NDPAFALVSEINELVRKEWLVE 417
Query: 1759 VHWTNRDSNASADFLARRG 1777
R+SN +AD LA G
Sbjct: 418 FSHVFRESNRAADKLAHLG 474
>BG404993
Length = 412
Score = 60.5 bits (145), Expect = 7e-09
Identities = 36/114 (31%), Positives = 54/114 (46%), Gaps = 7/114 (6%)
Frame = -3
Query: 1456 DRWSWKDNSDGISSVHDGYQWLREKKMAGVVHGDS------WLWI*KLHAPEKL*VFIWL 1509
D W+WK + G S Y+ L+E + GD+ LW KL P K VF W
Sbjct: 332 DSWTWKADPSGNYSTKTAYELLQE-----AIRGDNEDGIFVQLW--KLKIPPKAPVFTWR 174
Query: 1510 VLHDALPTNANRYRCKLAVSPG-CTRCSSPEEDTIHCLRDCPHSQELWGKLGAW 1562
+++D LPT N R ++ +S C C++ EED H +C + W + +W
Sbjct: 173 LINDRLPTKVNLRRRQVEISDSLCPLCNNSEEDAAHLFFNCNKTLPFWWESLSW 12
>BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (15%)
Length = 376
Score = 59.7 bits (143), Expect = 1e-08
Identities = 29/85 (34%), Positives = 46/85 (54%)
Frame = -2
Query: 895 ALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKV 954
A+ + S + GPDG + F K +WE + D+ + + + G+ + I LIPKV
Sbjct: 366 AVWQCGSVKSPGPDGLNFKFIKHFWERLKPDIIRFLNEFHANGIFPKGGNASFIALIPKV 187
Query: 955 DSPSTVKEFRPISLCNVVYKLITKV 979
P + +FRPISL VYK++ K+
Sbjct: 186 KHPQALNDFRPISLIGCVYKIVAKI 112
>TC212765
Length = 637
Score = 59.7 bits (143), Expect = 1e-08
Identities = 29/72 (40%), Positives = 39/72 (53%)
Frame = +2
Query: 1233 LSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQRG 1292
L+SWK LLN + CL V+ A P Y M+ WLP+SI I++ R IW K
Sbjct: 416 LASWKGKLLNQASKHCLT*LVLHAFPIYPMEASWLPQSIFQAIDKASRVTIWNKVGASLS 595
Query: 1293 WHRVNWKTLTQP 1304
WH V+ T ++P
Sbjct: 596 WHSVS*STASKP 631
>TC231841
Length = 791
Score = 59.3 bits (142), Expect = 1e-08
Identities = 55/203 (27%), Positives = 81/203 (39%), Gaps = 14/203 (6%)
Frame = +3
Query: 1456 DRWSWKDNSDGISSVHDGYQ--W--LREKKMAGVVHGDSWLWI*KLHAPEKL*VFIWLVL 1511
D+W WK + G + Y W + E++ GV LW KL P K+ +F W ++
Sbjct: 63 DQWVWKADPSGQYTAKSAYGVLWGEMFEEQQDGVFEE---LW--KLKLPSKITIFAWRLI 227
Query: 1512 HDALPTNANRYRCKLAV-SPGCTRCSSPEEDTIHCLRDCPHSQELWGKLGAWGWSTFRLP 1570
D LPT +N R ++ V P C C S EE H C +W + +W P
Sbjct: 228 RDRLPTRSNLRRKQIEVDDPRCPFCRSAEESAAHLFFHCSRIAPVWWESLSWVNLLGVFP 407
Query: 1571 D-----LKA*VKTQLQGSNNIR----FLAGLWGVWKWRNNVVLDPLPWTLDGAWQKLRHD 1621
+ + G R +LA W +WK RNN++ +G
Sbjct: 408 NHPRQHFLQHIYGVTAGMRASRWKWWWLALTWTIWKQRNNMIFS------NGT------- 548
Query: 1622 HDELLQFSSGDLLDEHHFLINLW 1644
F++ +LDE FLI W
Sbjct: 549 ------FNANKILDEAIFLIWTW 599
>TC233392
Length = 1145
Score = 58.2 bits (139), Expect = 3e-08
Identities = 46/164 (28%), Positives = 67/164 (40%), Gaps = 16/164 (9%)
Frame = -1
Query: 1456 DRWSWKDNSDGISSVHDGYQWLREKKMAGVVHGDSW------LWI*KLHAPEKL*VFIWL 1509
D W WK + + S Y+ L+ G + G+ LW KL P K +F W
Sbjct: 656 DTWVWKHEPNELYSTRSAYKLLQ-----GHLEGEDQDGALQDLW--KLKIPAKASIFAWR 498
Query: 1510 VLHDALPTNANRYRCKLAVSPG-CTRCSSPEEDTIHCLRDCPHSQELWGKLGAWGWSTFR 1568
++ D LPT +N +R ++ + C C EED H +C + LW + W S
Sbjct: 497 LIRDRLPTKSNLHRRQVVLEDSLCPFCRIREEDASHIFLECNKIRPLWWESQTWVRSLGV 318
Query: 1569 LPDLKA*VKTQ----LQGSNNIR-----FLAGLWGVWKWRNNVV 1603
P K Q GS ++A W +WK RN V+
Sbjct: 317 SPINKRQHFLQHVHGTPGSKRYNRWKTWWIALTWSLWKHRNQVI 186
>CF922212
Length = 445
Score = 57.8 bits (138), Expect = 4e-08
Identities = 35/115 (30%), Positives = 56/115 (48%)
Frame = -2
Query: 1267 LPRSINHKINQEMRRFIWLKNSGQRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKV 1326
+P + K+ + + F+W GQ+ VNW ++ PKE GL +D+ FN +LLGK
Sbjct: 441 VPNKVLDKLVRLQQWFLWGGGLGQKKIAWVNWDSVCLPKEKEGLGNRDLRKFNYALLGKR 262
Query: 1327 VWLLANNSSKLWVQVLAHKYLHGESIFSVHRRANASPIWQGILKARDQLQEGFQF 1381
W L ++ +L +VL KY ++ R + S WQ I ++G F
Sbjct: 261 RWNLFHHQGELGARVLDSKYKRWRNLDEERRVKSESFWWQEISFITHSTEDGSWF 97
>CO979823
Length = 853
Score = 57.8 bits (138), Expect = 4e-08
Identities = 45/161 (27%), Positives = 68/161 (41%), Gaps = 13/161 (8%)
Frame = -2
Query: 1456 DRWSWKDNSDGISSVHDGYQWLREKKMAGVVHGD-SWLWI*KLHAPEKL*VFIWLVLHDA 1514
D+W WK G S GY L + + D + +W KL P K VF W ++ D
Sbjct: 702 DKWLWKPEPGGHYSTKSGYHVLWGELTEEIQDADFAEIW--KLKIPTKAAVFAWRLVRDR 529
Query: 1515 LPTNANRYRCKLAVSP-GCTRCSSPEEDTIHCLRDCPHSQELWGKLGAWGWSTFRLPDLK 1573
LPT +N R ++ V C C++ EE H +C + LW + +W +P
Sbjct: 528 LPTKSNLRRRQVMVQDMVCPLCNNIEEGAAHLFFNCTKTLPLWWESMSWVNLKTAMPQTP 349
Query: 1574 A*VKTQLQGSNNIR-----------FLAGLWGVWKWRNNVV 1603
+ LQ +I ++A W +W+ RN VV
Sbjct: 348 R--QHFLQYGTDIADGLKSKRWKCWWIALTWTIWQHRNKVV 232
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like protein
{Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 57.8 bits (138), Expect = 4e-08
Identities = 24/45 (53%), Positives = 33/45 (73%)
Frame = -3
Query: 1061 VQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLC 1105
VQL+ C+S + +S+LWNG L F P G+R+ DP++PYLFVLC
Sbjct: 686 VQLVWYCMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFVLC 552
Score = 39.7 bits (91), Expect = 0.012
Identities = 17/33 (51%), Positives = 26/33 (78%)
Frame = -2
Query: 1231 KKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQ 1263
K+ S WK+NLL+MVG++ L K V+ A+P++ MQ
Sbjct: 171 KRFSRWKSNLLSMVGRLTLTK*VV*ALPSHIMQ 73
>BM891241
Length = 407
Score = 57.4 bits (137), Expect = 6e-08
Identities = 23/51 (45%), Positives = 34/51 (66%)
Frame = -2
Query: 1296 VNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKY 1346
V W+ + PK GGL IKD+ FN +LLGK +W LA++ +LW +++ KY
Sbjct: 373 VKWEVVCLPKTKGGLGIKDISKFNIALLGKWIWALASDQQQLWARIINSKY 221
>TC223713
Length = 420
Score = 57.0 bits (136), Expect = 7e-08
Identities = 32/104 (30%), Positives = 52/104 (49%), Gaps = 1/104 (0%)
Frame = +1
Query: 1460 WKDNSDGISSVHDGYQWLREKKMAGVVHGDSWLWI*KLHAPEKL*VFIWLVLHDALPTNA 1519
W+ + G + Y +RE ++ G+ ++ +*KL P K VF W +L D LPT A
Sbjct: 10 WEADQSGQYTAQTAYNLMREVEVGGI-QDRAFEEL*KLKVPIKFAVFAWRLLRDRLPTKA 186
Query: 1520 NRYRCKLAV-SPGCTRCSSPEEDTIHCLRDCPHSQELWGKLGAW 1562
N +R ++ V + C CS+ EE+ H C +W + +W
Sbjct: 187 NLHRRQIEVTNRSCPFCSNMEEEAGHLFFHCSKIIPIWWETSSW 318
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 56.6 bits (135), Expect = 1e-07
Identities = 36/109 (33%), Positives = 56/109 (51%)
Frame = -3
Query: 1005 IMDNAFLAQEIIHHMGASMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLI 1064
I D + E+I+ + + G LA KID+ KA+D++ FL L +FG+ + I
Sbjct: 834 IKDCTCVTSEVINMLDKKVFG-GNLALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNWI 658
Query: 1065 MRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLCSLHGAIVY 1113
+ + LS+ NG + F RG+R+GDPLSP L+ L +Y
Sbjct: 657 RVILLSAKLSISVNGEPVGFFSCQRGVRQGDPLSPSLYRQRCLEQGTIY 511
Score = 33.1 bits (74), Expect = 1.1
Identities = 14/36 (38%), Positives = 25/36 (68%)
Frame = -2
Query: 1230 KKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVF 1265
K KL+SWK ++L+++G++ L SVI + Y+ V+
Sbjct: 166 KCKLASWKGSILSIMGRIQLVNSVIHGMLLYSFSVY 59
>AW620606
Length = 420
Score = 50.4 bits (119), Expect(2) = 2e-07
Identities = 29/66 (43%), Positives = 39/66 (58%), Gaps = 2/66 (3%)
Frame = +1
Query: 1509 LVLHDALPTNANRYRCKLAVSPGC--TRCSSPEEDTIHCLRDCPHSQELWGKLGAWGWST 1566
L+L++ALPTN RYR LA S + CS ++D IHCL DCP ++E+ LG
Sbjct: 7 LILNNALPTNFLRYRPHLASSSSSAFSPCS*NDQDIIHCLWDCPKAREV*SCLGFLSPKA 186
Query: 1567 FRLPDL 1572
F +P L
Sbjct: 187 FLIPTL 204
Score = 25.0 bits (53), Expect(2) = 2e-07
Identities = 10/25 (40%), Positives = 13/25 (52%)
Frame = +2
Query: 1577 KTQLQGSNNIRFLAGLWGVWKWRNN 1601
K L I ++ L +WKWRNN
Sbjct: 215 KA*LLAPRGILYVVAL*WIWKWRNN 289
>BI972659
Length = 453
Score = 55.5 bits (132), Expect = 2e-07
Identities = 27/70 (38%), Positives = 41/70 (58%)
Frame = +1
Query: 1229 FKKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNS 1288
FK KLS W L+M G+V L KSV++A+P Y + F +P+ I K+ R F+W
Sbjct: 250 FKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFFKIPQRIVDKLVSLQRTFMW---G 420
Query: 1289 GQRGWHRVNW 1298
G + +R++W
Sbjct: 421 GNQHHNRISW 450
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.333 0.144 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 97,279,652
Number of Sequences: 63676
Number of extensions: 1628529
Number of successful extensions: 9586
Number of sequences better than 10.0: 122
Number of HSP's better than 10.0 without gapping: 9277
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 9557
length of query: 1813
length of database: 12,639,632
effective HSP length: 111
effective length of query: 1702
effective length of database: 5,571,596
effective search space: 9482856392
effective search space used: 9482856392
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0178.7


