

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.5
(770 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
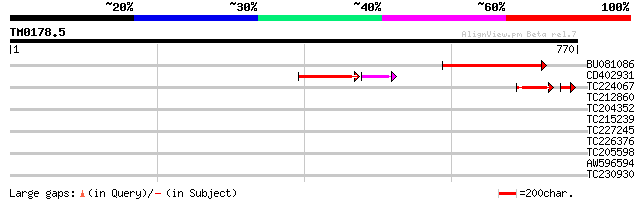
Score E
Sequences producing significant alignments: (bits) Value
BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza s... 148 1e-35
CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (ja... 101 1e-26
TC224067 39 6e-04
TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and... 38 0.019
TC204352 homologue to UP|Q9SP48 (Q9SP48) Homeodomain-leucine zip... 34 0.28
TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%) 33 0.62
TC227245 homologue to UP|Q941A9 (Q941A9) At1g26300/F28B23_4, par... 31 2.4
TC226376 similar to UP|VATC_ARATH (Q9SDS7) Vacuolar ATP synthase... 29 8.9
TC205598 similar to UP|BRU1_SOYBN (P35694) Brassinosteroid-regul... 29 8.9
AW596594 29 8.9
TC230930 similar to UP|Q94AA9 (Q94AA9) AT5g33290/F19N2_10, parti... 29 8.9
>BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 426
Score = 148 bits (373), Expect = 1e-35
Identities = 72/140 (51%), Positives = 94/140 (66%)
Frame = +2
Query: 589 DTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLCNDTR 648
D++ KSD T+EF+N S +PNH IKLKV IML+RN+DQ GLCN TR
Sbjct: 5 DSIEKSDTIESEGFSTITTEFINSLSTSRLPNHKIKLKVDSRIMLLRNLDQNEGLCNSTR 184
Query: 649 MIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTINK 708
+I+ +II A I++G G T +IPR++ +PS S PFK RR+FP+ + +AMTINK
Sbjct: 185 LIITMFADHIIEAKIMSGKGQGNTVYIPRLATSPSQSPWPFKLIRRKFPIIVSYAMTINK 364
Query: 709 SQGQSLSHVGLYLPRPVFTH 728
SQGQ L+ VGLYLP PVF+H
Sbjct: 365 SQGQLLASVGLYLPTPVFSH 424
>CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 617
Score = 101 bits (252), Expect(2) = 1e-26
Identities = 55/83 (66%), Positives = 64/83 (76%)
Frame = +2
Query: 393 LIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVISKGS 452
LIIWDETP++NK CFEALDR+L DI+ +Q+ PFGG VVLGGDFRQILPVI KGS
Sbjct: 227 LIIWDETPVMNKFCFEALDRTLQDIMASQNKDNATKPFGG-KVVLGGDFRQILPVIRKGS 403
Query: 453 RSEIVGSAINSSYLWKHCKVMKL 475
R +IVGSAIN+S KV+KL
Sbjct: 404 RQDIVGSAINASK-----KVLKL 457
Score = 37.4 bits (85), Expect(2) = 1e-26
Identities = 20/48 (41%), Positives = 26/48 (53%)
Frame = +1
Query: 478 NMILQNATSTSSPAEIKEFADWLLQVGDGTVKTIDEEETLIEIPPNLL 525
NM L + +I+EF DW+L + DG DE E I+IP NLL
Sbjct: 463 NMRLGTTNNNEDRDDIREFVDWILMIRDGNRDEEDEGE--IDIPKNLL 600
>TC224067
Length = 415
Score = 39.3 bits (90), Expect(2) = 6e-04
Identities = 27/52 (51%), Positives = 34/52 (64%), Gaps = 2/52 (3%)
Frame = -2
Query: 689 FKFQRRQFPVTLCFAMTINKSQGQSLSHV-GLYLPRPV-FTHGQLYVALSRV 738
FKFQ +C+ M INKS GQ+LS V G+ LPRP +HGQ YV L++V
Sbjct: 387 FKFQ--DVNSLVCYTMKINKSHGQTLSQVGGVLLPRPN**SHGQ-YVTLNQV 241
Score = 22.7 bits (47), Expect(2) = 6e-04
Identities = 9/20 (45%), Positives = 14/20 (70%)
Frame = -3
Query: 749 IDDEGVVSNTTRNVMYQEVF 768
ID+E +T RN++Y+E F
Sbjct: 233 IDEENQRVSTARNMIYKEFF 174
>TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and
recombination protein PIF1, mitochondrial precursor,
partial (8%)
Length = 596
Score = 37.7 bits (86), Expect = 0.019
Identities = 22/45 (48%), Positives = 30/45 (65%)
Frame = +2
Query: 702 FAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKM 746
+AM+I+K QG +L V L R F G +YVALSRV+S +GL +
Sbjct: 5 WAMSIHKCQGMTLERVHTDLSR-AFGCGMVYVALSRVRSLEGLHL 136
>TC204352 homologue to UP|Q9SP48 (Q9SP48) Homeodomain-leucine zipper protein
56, partial (72%)
Length = 645
Score = 33.9 bits (76), Expect = 0.28
Identities = 18/74 (24%), Positives = 37/74 (49%)
Frame = +1
Query: 526 IEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESVEEINNFMLAMIPGDETEY 585
++Q E LL+ + +L N+ + ++ + ++++ N + IPG +++
Sbjct: 313 LQQDNEALLKQIKELKSRLVQEENNNNTESDVSVKEEMIATLQDSNPLCESAIPGSDSKE 492
Query: 586 LSYDTLCKSDEDSG 599
LSY+ KSDE G
Sbjct: 493 LSYECFNKSDEVGG 534
>TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%)
Length = 340
Score = 32.7 bits (73), Expect = 0.62
Identities = 14/22 (63%), Positives = 18/22 (81%)
Frame = -3
Query: 692 QRRQFPVTLCFAMTINKSQGQS 713
QR FP+ +CFAMT NKS+GQ+
Sbjct: 338 QR**FPLIVCFAMTTNKSEGQT 273
>TC227245 homologue to UP|Q941A9 (Q941A9) At1g26300/F28B23_4, partial (14%)
Length = 787
Score = 30.8 bits (68), Expect = 2.4
Identities = 18/55 (32%), Positives = 29/55 (52%)
Frame = -1
Query: 88 INNMTS*EKRHFSWSIAIYSSWNGRKLLHACFTN*AKRL*QF*KHKNCKGCCLSH 142
I+ + + K HF +S++ S+W KLL F++ A L +H NC+ C H
Sbjct: 502 ISGIVASSKVHFDFSLSTKSNWGNPKLLQQ-FSSHASHL----QHLNCQSCQNHH 353
>TC226376 similar to UP|VATC_ARATH (Q9SDS7) Vacuolar ATP synthase subunit C
(V-ATPase C subunit) (Vacuolar proton pump C subunit) ,
partial (6%)
Length = 636
Score = 28.9 bits (63), Expect = 8.9
Identities = 14/41 (34%), Positives = 23/41 (55%)
Frame = -1
Query: 695 QFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVAL 735
QF +TLC A +K QS+SH+ +++ T+ + Y L
Sbjct: 552 QFIITLCRAALFSKGMSQSMSHLQVHIIADKKTYTKQYKLL 430
>TC205598 similar to UP|BRU1_SOYBN (P35694) Brassinosteroid-regulated protein
BRU1 precursor , partial (88%)
Length = 1015
Score = 28.9 bits (63), Expect = 8.9
Identities = 15/46 (32%), Positives = 27/46 (58%), Gaps = 1/46 (2%)
Frame = -1
Query: 450 KGSRSEIVGSAINSSYLWKHCKVMKLTV-NMILQNATSTSSPAEIK 494
K +RSE+ A+N + LW+ K++ + + + +L N + SS IK
Sbjct: 886 KKNRSEVSAPALNRNALWESLKIVAVVIDHEVLLNPSQFSSAIGIK 749
>AW596594
Length = 295
Score = 28.9 bits (63), Expect = 8.9
Identities = 14/41 (34%), Positives = 23/41 (55%)
Frame = -1
Query: 695 QFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVAL 735
QF +TLC A +K QS+SH+ +++ T+ + Y L
Sbjct: 286 QFIITLCRAALFSKGMSQSMSHLQVHIIADKKTYTKQYKLL 164
>TC230930 similar to UP|Q94AA9 (Q94AA9) AT5g33290/F19N2_10, partial (66%)
Length = 911
Score = 28.9 bits (63), Expect = 8.9
Identities = 22/67 (32%), Positives = 30/67 (43%)
Frame = -3
Query: 293 SKRLFIRMFWMLFCLIMVDSFFYMVLEELDKHLFGIHYLLLCVQGALSS*MSHLAELHRY 352
SK F+ + W CL LEE+ H+FG HYL S+ + L ELH
Sbjct: 579 SKTCFVPLVWYRLCL----------LEEI--HIFGHHYL------CPSNDSTRLFELHHV 454
Query: 353 CYLEVEL 359
Y + +L
Sbjct: 453 LYQQRKL 433
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.347 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,565,217
Number of Sequences: 63676
Number of extensions: 512935
Number of successful extensions: 4811
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 3546
Number of HSP's successfully gapped in prelim test: 135
Number of HSP's that attempted gapping in prelim test: 1230
Number of HSP's gapped (non-prelim): 3749
length of query: 770
length of database: 12,639,632
effective HSP length: 105
effective length of query: 665
effective length of database: 5,953,652
effective search space: 3959178580
effective search space used: 3959178580
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0178.5


