

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.13
(433 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
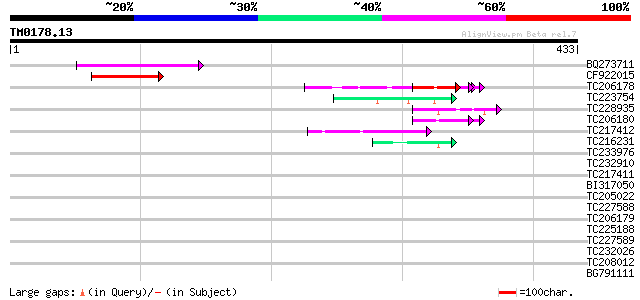
Score E
Sequences producing significant alignments: (bits) Value
BQ273711 92 6e-19
CF922015 60 2e-09
TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 47 2e-05
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 44 1e-04
TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana geno... 43 2e-04
TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 43 2e-04
TC217412 similar to UP|Q6L724 (Q6L724) ATP-dependent RNA helicas... 43 2e-04
TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (G... 42 7e-04
TC233976 41 0.001
TC232910 similar to UP|Q6I8N0 (Q6I8N0) Pol polypeptide, partial ... 39 0.005
TC217411 similar to UP|Q8L7S8 (Q8L7S8) At5g26740, partial (26%) 39 0.006
BI317050 38 0.010
TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, pa... 38 0.010
TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L1... 37 0.017
TC206179 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 37 0.023
TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like prot... 36 0.030
TC227589 36 0.030
TC232026 35 0.066
TC208012 similar to UP|Q761Z7 (Q761Z7) BRI1-KD interacting prote... 33 0.33
BG791111 32 0.43
>BQ273711
Length = 409
Score = 91.7 bits (226), Expect = 6e-19
Identities = 44/97 (45%), Positives = 57/97 (58%)
Frame = -2
Query: 52 EEREQAAAQTRGLNDFKRQDPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATY 111
+ER+ A+ RGL F++ PPK SG +DPE A LWL E EKIF + E KV YAT+
Sbjct: 291 QERDVEPAEYRGLMAFRKNHPPKFSGDYDPEGARLWLAETEKIFEAMGCLEEHKVLYATF 112
Query: 112 LLTGEAEYWWRGARAMMEADHQAITWECFRGAFLDKY 148
+L GEAE WW+ + A I W F+ FL+ Y
Sbjct: 111 MLQGEAENWWKFVKPSFVAPGGVIPWNAFKDKFLENY 1
>CF922015
Length = 172
Score = 59.7 bits (143), Expect = 2e-09
Identities = 26/55 (47%), Positives = 37/55 (67%)
Frame = -3
Query: 63 GLNDFKRQDPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEA 117
GL+ F+R +PP GG+DPE A+ WL+E+EKIF + KV +AT++L EA
Sbjct: 167 GLDRFQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADEA 3
>TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (69%)
Length = 1138
Score = 46.6 bits (109), Expect = 2e-05
Identities = 37/132 (28%), Positives = 56/132 (42%), Gaps = 1/132 (0%)
Frame = +2
Query: 226 EKTKGIEAIDNARGKFQSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGS 285
E+ K + ++R + +S+++ S M + R+D+ Y+ PY R +G + +
Sbjct: 185 ERKKKWKMSSDSRSRSRSRSR-------SPMDRKIRSDRFS-YRDAPYRRDSRRGFSRDN 340
Query: 286 FYPTGGNAIAPRPLAVSLDGVT-CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIP 344
N P A V C C GH AS C L CWNC +PGH+A C
Sbjct: 341 LCK---NCKRPGHYARECPNVAICHNCGLPGHIASECTTKSL-CWNCKEPGHMASSCPNE 508
Query: 345 KVEDAANVAGAR 356
+ AG R
Sbjct: 509 GICHTCGKAGHR 544
Score = 43.9 bits (102), Expect = 1e-04
Identities = 25/55 (45%), Positives = 28/55 (50%)
Frame = +2
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGG 362
C C + GH A CP P+ C CN GH+AR C PK ANV G R GG
Sbjct: 650 CNNCRKTGHLARDCPNDPI-CNLCNVSGHVARQC--PK----ANVLGDRSGGGGG 793
Score = 43.1 bits (100), Expect = 2e-04
Identities = 19/38 (50%), Positives = 23/38 (60%), Gaps = 1/38 (2%)
Frame = +2
Query: 308 CFKCNQKGHYASHCP-EGPLMCWNCNKPGHLARDCRIP 344
C+ C + GH AS CP EG +C C K GH AR+C P
Sbjct: 458 CWNCKEPGHMASSCPNEG--ICHTCGKAGHRARECSAP 565
Score = 43.1 bits (100), Expect = 2e-04
Identities = 20/47 (42%), Positives = 26/47 (54%)
Frame = +2
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVEDAANVAG 354
C C ++GH A+ C C NC K GHLARDC + + NV+G
Sbjct: 593 CNNCYKQGHIAAECTNEKA-CNNCRKTGHLARDCPNDPICNLCNVSG 730
Score = 38.5 bits (88), Expect = 0.006
Identities = 19/37 (51%), Positives = 21/37 (56%), Gaps = 1/37 (2%)
Frame = +2
Query: 306 VTCFKCNQKGHYASHCPEGPLM-CWNCNKPGHLARDC 341
V C C Q GH + C GPLM C NC GHLA +C
Sbjct: 836 VVCRNCQQLGHMSRDCM-GPLMICHNCGGRGHLAYEC 943
Score = 36.2 bits (82), Expect = 0.030
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 6/40 (15%)
Frame = +2
Query: 308 CFKCNQKGHYASHCPEGPL------MCWNCNKPGHLARDC 341
C C + GH A C P+ +C NC K GH+A +C
Sbjct: 515 CHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAEC 634
Score = 32.7 bits (73), Expect = 0.33
Identities = 17/62 (27%), Positives = 23/62 (36%), Gaps = 25/62 (40%)
Frame = +2
Query: 308 CFKCNQKGHYASHCPEGPLM-------------------------CWNCNKPGHLARDCR 342
C CN GH A CP+ ++ C NC + GH++RDC
Sbjct: 707 CNLCNVSGHVARQCPKANVLGDRSGGGGGGGGARGGGGGGYRDVVCRNCQQLGHMSRDCM 886
Query: 343 IP 344
P
Sbjct: 887 GP 892
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 43.9 bits (102), Expect = 1e-04
Identities = 35/122 (28%), Positives = 42/122 (33%), Gaps = 28/122 (22%)
Frame = +2
Query: 248 NQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQ------GTASGSFYPTGGNAIAPRPLAV 301
N G GG N G + + VR G G GS + GG R A
Sbjct: 98 NSGGGGGGSGGGCFNCGGFGHLARDCVRGGGGSVGIGGGGGGGSCFRCGGFGHMARDCAT 277
Query: 302 SL-------DGVTCFKCNQKGHYASHCPE---------------GPLMCWNCNKPGHLAR 339
G CF+C + GH A C G C+NC KPGH AR
Sbjct: 278 GKGNIGGGGSGGGCFRCGEVGHLARDCGMEGGRFGGGGGSGGGGGKSTCFNCGKPGHFAR 457
Query: 340 DC 341
+C
Sbjct: 458 EC 463
Score = 37.0 bits (84), Expect = 0.017
Identities = 17/47 (36%), Positives = 21/47 (44%), Gaps = 10/47 (21%)
Frame = +2
Query: 305 GVTCFKCNQKGHYASHCPEGPLM----------CWNCNKPGHLARDC 341
G C++C + GH A C C+NC GHLARDC
Sbjct: 35 GAACYQCGEFGHLARDCNRSSNSGGGGGGSGGGCFNCGGFGHLARDC 175
Score = 32.3 bits (72), Expect = 0.43
Identities = 18/53 (33%), Positives = 20/53 (36%), Gaps = 12/53 (22%)
Frame = +2
Query: 305 GVTCFKCNQKGHYASHCPEGPL------------MCWNCNKPGHLARDCRIPK 345
G CF C GH A C G C+ C GH+ARDC K
Sbjct: 125 GGGCFNCGGFGHLARDCVRGGGGSVGIGGGGGGGSCFRCGGFGHMARDCATGK 283
Score = 28.5 bits (62), Expect = 6.2
Identities = 14/39 (35%), Positives = 19/39 (47%)
Frame = +2
Query: 324 GPLMCWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGG 362
G C+ C + GHLARDC ++N G + GG
Sbjct: 32 GGAACYQCGEFGHLARDC-----NRSSNSGGGGGGSGGG 133
Score = 28.1 bits (61), Expect = 8.0
Identities = 9/17 (52%), Positives = 11/17 (63%)
Frame = +2
Query: 307 TCFKCNQKGHYASHCPE 323
TCF C + GH+A C E
Sbjct: 419 TCFNCGKPGHFARECVE 469
>TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K24M7, partial (36%)
Length = 1224
Score = 43.1 bits (100), Expect = 2e-04
Identities = 29/85 (34%), Positives = 42/85 (49%), Gaps = 17/85 (20%)
Frame = +1
Query: 308 CFKCNQKGHYASHCPEGPL-------MCWNCNKPGHLARDCRIPKVEDAANVAGARRPTA 360
CF CNQ+GH + +CP+ C C HLA+DC P + +VA A RP
Sbjct: 505 CFVCNQRGHLSKNCPQNTHGIYPKGGCCKICGGVTHLAKDC--PDKGKSGSVA-ANRPAD 675
Query: 361 G---------GRV-YFISGTEVDED 375
G G+V F+SG ++++D
Sbjct: 676 GWMRIDERPMGQVTKFVSGDDIEDD 750
Score = 38.9 bits (89), Expect = 0.005
Identities = 17/52 (32%), Positives = 23/52 (43%), Gaps = 6/52 (11%)
Frame = +1
Query: 308 CFKCNQKGHYASHCPE------GPLMCWNCNKPGHLARDCRIPKVEDAANVA 353
C +C ++GH A +CPE C+NC + GH C P E A
Sbjct: 346 CLRCRRRGHRAKNCPEVLDGAKDAKYCYNCGENGHALTQCLHPLQEGGTKFA 501
Score = 36.6 bits (83), Expect = 0.023
Identities = 20/54 (37%), Positives = 26/54 (48%), Gaps = 5/54 (9%)
Frame = +1
Query: 305 GVTCFKCNQKGHYASHCPEGP-----LMCWNCNKPGHLARDCRIPKVEDAANVA 353
G +CF C H A CPE +C C + GH A++C P+V D A A
Sbjct: 262 GESCFICKAMDHIAKLCPEKAEWEKNKICLRCRRRGHRAKNC--PEVLDGAKDA 417
>TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (42%)
Length = 655
Score = 43.1 bits (100), Expect = 2e-04
Identities = 20/47 (42%), Positives = 26/47 (54%)
Frame = +3
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVEDAANVAG 354
C C ++GH A+ C C NC K GHLARDC + + NV+G
Sbjct: 78 CNNCYKQGHIAAECTNEKA-CNNCRKTGHLARDCPNDPICNLCNVSG 215
Score = 41.6 bits (96), Expect = 7e-04
Identities = 24/55 (43%), Positives = 27/55 (48%)
Frame = +3
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGG 362
C C + GH A CP P+ C CN GH+AR C PK ANV G GG
Sbjct: 135 CNNCRKTGHLARDCPNDPI-CNLCNVSGHVARQC--PK----ANVLGDXXGGGGG 278
Score = 38.5 bits (88), Expect = 0.006
Identities = 19/37 (51%), Positives = 21/37 (56%), Gaps = 1/37 (2%)
Frame = +3
Query: 306 VTCFKCNQKGHYASHCPEGPLM-CWNCNKPGHLARDC 341
V C C Q GH + C GPLM C NC GHLA +C
Sbjct: 312 VVCRNCQQLGHMSRDCM-GPLMICHNCGGRGHLAYEC 419
Score = 33.9 bits (76), Expect = 0.15
Identities = 17/59 (28%), Positives = 23/59 (38%), Gaps = 22/59 (37%)
Frame = +3
Query: 308 CFKCNQKGHYASHCPEGPLM----------------------CWNCNKPGHLARDCRIP 344
C CN GH A CP+ ++ C NC + GH++RDC P
Sbjct: 192 CNLCNVSGHVARQCPKANVLGDXXGGGGGARGGGGGGYRDVVCRNCQQLGHMSRDCMGP 368
Score = 33.5 bits (75), Expect = 0.19
Identities = 14/37 (37%), Positives = 19/37 (50%), Gaps = 6/37 (16%)
Frame = +3
Query: 311 CNQKGHYASHCPEGPL------MCWNCNKPGHLARDC 341
C + GH A C P+ +C NC K GH+A +C
Sbjct: 9 CGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAEC 119
>TC217412 similar to UP|Q6L724 (Q6L724) ATP-dependent RNA helicase, partial
(3%)
Length = 868
Score = 43.1 bits (100), Expect = 2e-04
Identities = 29/95 (30%), Positives = 44/95 (45%)
Frame = +3
Query: 228 TKGIEAIDNARGKFQSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFY 287
++G A + RG F+S + + G +Q GR++ K + G +SG +
Sbjct: 78 SRGGSASRDRRG-FKSSRGLDVEDSGDDFGDQSSRRGGRNF-KTGNSWSRAAGKSSGDDW 251
Query: 288 PTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCP 322
GG + RP + G TCF C + GH AS CP
Sbjct: 252 LIGGGRRSSRPSSSDRFGGTCFNCGESGHRASDCP 356
>TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (GRP2)
(AT4g38680/F20M13_240), partial (42%)
Length = 891
Score = 41.6 bits (96), Expect = 7e-04
Identities = 24/77 (31%), Positives = 29/77 (37%), Gaps = 13/77 (16%)
Frame = +3
Query: 278 GQGTASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPL----------- 326
G G G Y GG G C+KC + GH A C +G
Sbjct: 171 GGGYGGGGGYGGGGGG----------GGGGCYKCGETGHIARDCSQGGGGGGRYGGGGGG 320
Query: 327 --MCWNCNKPGHLARDC 341
C+NC + GH ARDC
Sbjct: 321 GGSCYNCGESGHFARDC 371
Score = 30.8 bits (68), Expect = 1.2
Identities = 15/48 (31%), Positives = 19/48 (39%)
Frame = +3
Query: 275 RPQGQGTASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCP 322
R QG G Y GG G +C+ C + GH+A CP
Sbjct: 264 RDCSQGGGGGGRYGGGGGG-----------GGSCYNCGESGHFARDCP 374
>TC233976
Length = 763
Score = 40.8 bits (94), Expect = 0.001
Identities = 30/113 (26%), Positives = 45/113 (39%), Gaps = 19/113 (16%)
Frame = +1
Query: 254 SAMTNQGRNDKGR-HYQKKPYVRPQ-------GQGTASGSFYPTGGNAIAPRPL------ 299
+A + G+ K H + KPY P GQ T+ G G + + R
Sbjct: 52 NANASHGKEKKPMTHSRAKPYSAPLEYENHYGGQRTSGGHHLAGGSSQLVNRVSQPAGRG 231
Query: 300 -----AVSLDGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVE 347
A+ + C KC + GH A C + C+N GHL+ +C P+ E
Sbjct: 232 GSGAPAIVTTPLRCRKCGRLGHNAHECTYREVTCFNYQGKGHLSTNCPHPRKE 390
>TC232910 similar to UP|Q6I8N0 (Q6I8N0) Pol polypeptide, partial (3%)
Length = 690
Score = 38.9 bits (89), Expect = 0.005
Identities = 16/33 (48%), Positives = 21/33 (63%)
Frame = +3
Query: 326 LMCWNCNKPGHLARDCRIPKVEDAANVAGARRP 358
L C C KPGHL RDCR+ K ++ A +G+ P
Sbjct: 27 LSCSKCGKPGHLKRDCRVFKGKNKAGPSGSNDP 125
>TC217411 similar to UP|Q8L7S8 (Q8L7S8) At5g26740, partial (26%)
Length = 1275
Score = 38.5 bits (88), Expect = 0.006
Identities = 17/43 (39%), Positives = 23/43 (52%)
Frame = +3
Query: 280 GTASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCP 322
G +SG + GG+ + RP + G CF C + GH AS CP
Sbjct: 732 GRSSGDDWLIGGSRRSSRPSSSDRFGGACFNCGESGHRASDCP 860
>BI317050
Length = 425
Score = 37.7 bits (86), Expect = 0.010
Identities = 36/143 (25%), Positives = 65/143 (45%), Gaps = 3/143 (2%)
Frame = +3
Query: 79 FDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWWRGARAMMEADHQAITWE 138
FD A + ++ + F + RT + ++ ++ L GEA W++ + + Q +W
Sbjct: 27 FDDTNAPTLIFKISQFFDYHRTPEDERLQVTSFYLDGEALSWFQ----WLYRNDQLTSWS 194
Query: 139 CFRGAFLDKYFPNSARAAKEAQFLRLRQGGMTVAEYASKLESLAKHFRYFRGQIDEGYMC 198
F A L+ F S +L Q G +V + + E+LA + G + ++
Sbjct: 195 SFLQA-LEMRFAQSMYDDPRGTLFKLTQTG-SVNSFLAMFEALA---NWIVG-LPTPFLL 356
Query: 199 ERFIDGLSYELQR---AVQPLGL 218
FI GL +++R A+QPL L
Sbjct: 357 SCFISGLLPDIRREVLAMQPLSL 425
>TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (69%)
Length = 1623
Score = 37.7 bits (86), Expect = 0.010
Identities = 14/37 (37%), Positives = 20/37 (53%), Gaps = 2/37 (5%)
Frame = +2
Query: 308 CFKCNQKGHYASHCPEGPL--MCWNCNKPGHLARDCR 342
CF C GH+A C G C+ C + GH+ R+C+
Sbjct: 314 CFNCGLDGHWARDCKAGDWKNKCYRCGERGHIERNCK 424
Score = 30.0 bits (66), Expect = 2.1
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = +2
Query: 322 PEGPLMCWNCNKPGHLARDCR 342
P G C+NC GH ARDC+
Sbjct: 296 PPGSGRCFNCGLDGHWARDCK 358
>TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L16.11 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(10%)
Length = 1300
Score = 37.0 bits (84), Expect = 0.017
Identities = 15/45 (33%), Positives = 21/45 (46%), Gaps = 9/45 (20%)
Frame = +1
Query: 306 VTCFKCNQKGHYASHCPE---------GPLMCWNCNKPGHLARDC 341
++C+KC Q GH C P C+ C + GH AR+C
Sbjct: 412 ISCYKCGQLGHTGLACSRLRDEITSGATPSSCFKCGEEGHFAREC 546
Score = 33.5 bits (75), Expect = 0.19
Identities = 31/121 (25%), Positives = 44/121 (35%)
Frame = +1
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGGRVYFI 367
C+ C GH A C + C+ C K GH A+DC + ++A + G F
Sbjct: 127 CYVCGCLGHNARQCSKVQ-DCFICKKGGHRAKDCPEKHTSTSKSIAICLKCGNSGHDIFS 303
Query: 368 SGTEVDEDEGLIHGNREIPGNSLIVLFKLWCYTFCYRLSLCDVVKTRCDRITIRPDSIYS 427
+ +D+ L ++ CY C RL V T D T S Y
Sbjct: 304 CRNDYSQDD----------------LKEIQCYV-CKRLGHLCCVNT--DDATPGEISCYK 426
Query: 428 C 428
C
Sbjct: 427 C 429
Score = 33.1 bits (74), Expect = 0.25
Identities = 17/43 (39%), Positives = 23/43 (52%), Gaps = 2/43 (4%)
Frame = +1
Query: 308 CFKCNQKGHYASHCP--EGPLMCWNCNKPGHLARDCRIPKVED 348
CF C ++GH A +C + C+ C GH AR C KV+D
Sbjct: 61 CFNCGEEGHAAVNCSAVKRKKPCYVCGCLGHNARQC--SKVQD 183
Score = 29.3 bits (64), Expect = 3.6
Identities = 19/65 (29%), Positives = 25/65 (38%)
Frame = +1
Query: 307 TCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVEDAANVAGARRPTAGGRVYF 366
+CFKC ++GH+A C + N +D R K D A G R
Sbjct: 502 SCFKCGEEGHFARECTSS-IKSGKRNWESSHTKDKRSQKENDYMGNRSASNDMVGARRKK 678
Query: 367 ISGTE 371
S TE
Sbjct: 679 RSPTE 693
>TC206179 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (16%)
Length = 407
Score = 36.6 bits (83), Expect = 0.023
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = +3
Query: 311 CNQKGHYASHCPEGPLMCWNCNKPGHLARD 340
C + GHYA CP +C NC PGH+A +
Sbjct: 321 CKRPGHYARECP-NVAICHNCGLPGHIASE 407
>TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like protein, partial
(62%)
Length = 1131
Score = 36.2 bits (82), Expect = 0.030
Identities = 13/37 (35%), Positives = 20/37 (53%), Gaps = 2/37 (5%)
Frame = +1
Query: 308 CFKCNQKGHYASHCPEGPL--MCWNCNKPGHLARDCR 342
CF C GH+A C G C+ C + GH+ ++C+
Sbjct: 364 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 474
Score = 29.6 bits (65), Expect = 2.8
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = +1
Query: 322 PEGPLMCWNCNKPGHLARDCR 342
P G C+NC GH ARDC+
Sbjct: 346 PPGSGRCFNCGIDGHWARDCK 408
>TC227589
Length = 547
Score = 36.2 bits (82), Expect = 0.030
Identities = 15/48 (31%), Positives = 22/48 (45%), Gaps = 9/48 (18%)
Frame = +2
Query: 306 VTCFKCNQKGHYASHCPE---------GPLMCWNCNKPGHLARDCRIP 344
++C+KC Q GH C + P C C + GH A++C P
Sbjct: 398 ISCYKCGQLGHTGLACSKLPDEITSAATPSSCCKCGEAGHFAQECTSP 541
Score = 33.5 bits (75), Expect = 0.19
Identities = 14/34 (41%), Positives = 18/34 (52%)
Frame = +2
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDC 341
C+ C GH A C + C+ C K GH A+DC
Sbjct: 113 CYVCGGLGHNARQCTKAQ-DCFICKKGGHRAKDC 211
Score = 32.0 bits (71), Expect = 0.56
Identities = 14/36 (38%), Positives = 17/36 (46%), Gaps = 2/36 (5%)
Frame = +2
Query: 308 CFKCNQKGHYASHCPEGPLM--CWNCNKPGHLARDC 341
CF C + GH A +C C+ C GH AR C
Sbjct: 47 CFNCGEDGHAAVNCSAAKRKKPCYVCGGLGHNARQC 154
>TC232026
Length = 689
Score = 35.0 bits (79), Expect = 0.066
Identities = 13/32 (40%), Positives = 15/32 (46%)
Frame = +2
Query: 312 NQKGHYASHCPEGPLMCWNCNKPGHLARDCRI 343
NQ C CW C +PGHLA DC +
Sbjct: 2 NQHHWSKKTCSLSTYECWKCQRPGHLAEDCMV 97
>TC208012 similar to UP|Q761Z7 (Q761Z7) BRI1-KD interacting protein 117
(Fragment), partial (38%)
Length = 993
Score = 32.7 bits (73), Expect = 0.33
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = +1
Query: 323 EGPLMCWNCNKPGHLARDCR 342
+ P +C+ C K GHL+RDC+
Sbjct: 121 DAPKICYKCKKAGHLSRDCK 180
>BG791111
Length = 430
Score = 32.3 bits (72), Expect = 0.43
Identities = 9/16 (56%), Positives = 15/16 (93%)
Frame = +1
Query: 307 TCFKCNQKGHYASHCP 322
TC+KC+Q+GH++ +CP
Sbjct: 268 TCYKCDQRGHWSYYCP 315
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,913,854
Number of Sequences: 63676
Number of extensions: 266316
Number of successful extensions: 1645
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 1545
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1621
length of query: 433
length of database: 12,639,632
effective HSP length: 100
effective length of query: 333
effective length of database: 6,272,032
effective search space: 2088586656
effective search space used: 2088586656
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0178.13


