

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0176a.2
(342 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
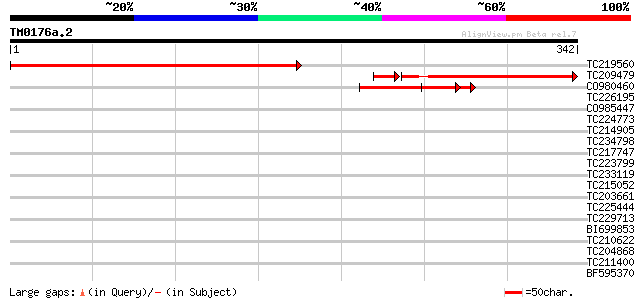
Score E
Sequences producing significant alignments: (bits) Value
TC219560 weakly similar to UP|Q94EI5 (Q94EI5) AT5g62130/mtg10_15... 326 1e-89
TC209479 weakly similar to UP|Q94EI5 (Q94EI5) AT5g62130/mtg10_15... 187 2e-50
CO980460 66 2e-11
TC226195 weakly similar to UP|Q95JD0 (Q95JD0) Basic proline-rich... 37 0.010
CO985447 32 0.54
TC224773 weakly similar to UP|Q9FF10 (Q9FF10) Similarity to hepa... 30 2.1
TC214905 similar to UP|Q8GT88 (Q8GT88) Fiber protein Fb1, partia... 30 2.1
TC234798 similar to UP|Q8GUL0 (Q8GUL0) GPAA1-like protein, parti... 29 2.7
TC217747 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxyg... 29 3.5
TC223799 weakly similar to UP|ML12_ARATH (O80961) MLO-like prote... 29 3.5
TC233119 weakly similar to GB|AAO39919.1|28372874|BT003691 At2g0... 29 3.5
TC215052 similar to UP|Q9XH69 (Q9XH69) Beta-amylase (Fragment), ... 28 4.6
TC203661 similar to GB|AAO63400.1|28950953|BT005336 At3g26700 {A... 28 4.6
TC225444 homologue to UP|RL24_CICAR (O65743) 60S ribosomal prote... 28 4.6
TC229713 28 6.0
BI699853 28 6.0
TC210622 homologue to UP|R10C_ARATH (P59231) 60S ribosomal prote... 28 6.0
TC204868 similar to UP|Q9LKY8 (Q9LKY8) Proline-rich protein, par... 28 6.0
TC211400 similar to UP|Q9FKJ0 (Q9FKJ0) Emb|CAB55405.1, partial (... 28 7.8
BF595370 28 7.8
>TC219560 weakly similar to UP|Q94EI5 (Q94EI5) AT5g62130/mtg10_150, partial
(43%)
Length = 646
Score = 326 bits (835), Expect = 1e-89
Identities = 150/177 (84%), Positives = 160/177 (89%), Gaps = 1/177 (0%)
Frame = +3
Query: 1 MIGGYVFAFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEH 60
M+ G+V AFLLVL SVEVIDASAGD DPRYR CI QCQETGCVAQ CFP+CKF SDGE
Sbjct: 114 MLDGWVCAFLLVLYCSVEVIDASAGDADPRYRVCITQCQETGCVAQRCFPNCKFSSDGEF 293
Query: 61 FDRPWYMQ-EPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQ 119
DRPWYMQ EPLYLQWKKWDCQ+DCRY+CMLDREKER+ NL PVKYHGKWPF RIYGMQ
Sbjct: 294 IDRPWYMQQEPLYLQWKKWDCQSDCRYYCMLDREKERESHNLGPVKYHGKWPFRRIYGMQ 473
Query: 120 EAASVAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSW 176
E ASVAFSALNLAMHFHGWVSFFIL++YKLPLKDGK+AYYEYAGLWH+YGLLSLNSW
Sbjct: 474 EPASVAFSALNLAMHFHGWVSFFILIHYKLPLKDGKKAYYEYAGLWHMYGLLSLNSW 644
>TC209479 weakly similar to UP|Q94EI5 (Q94EI5) AT5g62130/mtg10_150, partial
(13%)
Length = 735
Score = 187 bits (476), Expect(2) = 2e-50
Identities = 89/106 (83%), Positives = 94/106 (87%)
Frame = +2
Query: 237 VMYINFYKLDYGWNMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIY 296
VMYINFY LDY VCVVMAV QL +WAVWAG+S HPSRWKLWLVVIAGGLAMLLEIY
Sbjct: 62 VMYINFYLLDY-----VCVVMAVAQLSMWAVWAGVSNHPSRWKLWLVVIAGGLAMLLEIY 226
Query: 297 DFPPYEGLLDAHALWHATTIPLTYIWWSFIRDDAEFRTSIRVKKAK 342
DFPP++GL DAHALWHATTIPLTYIWWSFIRDDAEFRTS +KKAK
Sbjct: 227 DFPPHQGLFDAHALWHATTIPLTYIWWSFIRDDAEFRTSNLLKKAK 364
Score = 29.6 bits (65), Expect(2) = 2e-50
Identities = 14/16 (87%), Positives = 15/16 (93%)
Frame = +1
Query: 220 ATRVMVSAPLIAFVIT 235
ATRVMV+APLIAFV T
Sbjct: 10 ATRVMVAAPLIAFVTT 57
>CO980460
Length = 339
Score = 66.2 bits (160), Expect = 2e-11
Identities = 37/70 (52%), Positives = 45/70 (63%)
Frame = -2
Query: 212 RSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLDYGWNMIVCVVMAVVQLVIWAVWAGL 271
R+F++RDEATRVMV+APLIAFV THVMYINFY LDY V V + W +W
Sbjct: 338 RTFSIRDEATRVMVAAPLIAFVTTHVMYINFYLLDYA----VAVSLTYEGCKDW-IWLE* 174
Query: 272 SGHPSRWKLW 281
++ KLW
Sbjct: 173 LPSLTKLKLW 144
Score = 47.0 bits (110), Expect = 1e-05
Identities = 19/24 (79%), Positives = 22/24 (91%)
Frame = -1
Query: 249 WNMIVCVVMAVVQLVIWAVWAGLS 272
WNMIVCVVMA+ QL +WAVWAG+S
Sbjct: 75 WNMIVCVVMAMAQLSMWAVWAGVS 4
>TC226195 weakly similar to UP|Q95JD0 (Q95JD0) Basic proline-rich protein,
partial (17%)
Length = 654
Score = 37.4 bits (85), Expect = 0.010
Identities = 16/41 (39%), Positives = 23/41 (56%)
Frame = -2
Query: 253 VCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLL 293
+C+ A V LV+W W G S P R + W+ + G AM+L
Sbjct: 413 ICITSATVVLVVWRPWRGRSRWPRRRRSWMSIRWPGWAMVL 291
>CO985447
Length = 741
Score = 31.6 bits (70), Expect = 0.54
Identities = 16/80 (20%), Positives = 40/80 (50%), Gaps = 8/80 (10%)
Frame = -2
Query: 195 YSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLDYGWNM--- 251
++S + +GY L++A+ ++ T + + APL+ FV + ++ Y W++
Sbjct: 695 FASYSVRIGYFLLVALNTVLSLSSVLTTLFIEAPLLIFVSVFSCLVAWWIEIYHWSLHSE 516
Query: 252 -----IVCVVMAVVQLVIWA 266
+C+++ +V+W+
Sbjct: 515 DYARRRMCLLLIGFNVVLWS 456
>TC224773 weakly similar to UP|Q9FF10 (Q9FF10) Similarity to heparanase,
partial (56%)
Length = 1433
Score = 29.6 bits (65), Expect = 2.1
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = -1
Query: 65 WYMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYH 107
W +Q+ +QW K C + +C LD E L L P++YH
Sbjct: 1154 WGIQQ---MQWTKESCILEIARYCALDPTVEDLLRLLTPMEYH 1035
>TC214905 similar to UP|Q8GT88 (Q8GT88) Fiber protein Fb1, partial (74%)
Length = 796
Score = 29.6 bits (65), Expect = 2.1
Identities = 20/86 (23%), Positives = 37/86 (42%), Gaps = 3/86 (3%)
Frame = +1
Query: 130 NLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDVDL 189
N +F WV F+ + + L + E+A LW G W +F D D
Sbjct: 73 NFCSNFIYWVGVFLCVDFNCLLLVPSQDQREFAALWSCLG-------HWRGIFERYDKDR 231
Query: 190 T---ETLDYSSAVILLGYSLILAILR 212
+ + L+ A+ +GY++ ++L+
Sbjct: 232 SGKIDPLELKDALYGIGYAVPGSVLQ 309
>TC234798 similar to UP|Q8GUL0 (Q8GUL0) GPAA1-like protein, partial (12%)
Length = 652
Score = 29.3 bits (64), Expect = 2.7
Identities = 15/59 (25%), Positives = 27/59 (45%), Gaps = 11/59 (18%)
Frame = +3
Query: 228 PLIAFVITHVMYINFYKLDYG-----------WNMIVCVVMAVVQLVIWAVWAGLSGHP 275
P +AFV+ + +FY +++G WN + + V+ L WA+ + HP
Sbjct: 456 PPVAFVLLKGAFEDFYGMNFGDYWNWVESLWVWNSATYLYVGVIHLPCWALCIHILFHP 632
>TC217747 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxygenase-like
protein (AT5g05600/MOP10_14), partial (53%)
Length = 713
Score = 28.9 bits (63), Expect = 3.5
Identities = 17/45 (37%), Positives = 20/45 (43%), Gaps = 1/45 (2%)
Frame = -2
Query: 15 WSVEVIDASAG-DVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDG 58
W VE ++ S+G PR D Q CV SC HC P G
Sbjct: 484 WQVE*VNHSSGCRTRPRTHDLCLSSQSP-CVLHSCILHCSAPEFG 353
>TC223799 weakly similar to UP|ML12_ARATH (O80961) MLO-like protein 12
(AtMlo12) (AtMlo18), partial (25%)
Length = 449
Score = 28.9 bits (63), Expect = 3.5
Identities = 18/65 (27%), Positives = 30/65 (45%)
Frame = +2
Query: 78 WDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAFSALNLAMHFHG 137
W C D +H L + ++ L++ + H FA + G+ F+ L L + HG
Sbjct: 131 WHCGMDSSWHI*LQQMRQN-LISRTISREHLDEDFAAVVGITPTIWF-FAVLILLTNTHG 304
Query: 138 WVSFF 142
W S+F
Sbjct: 305 WYSYF 319
>TC233119 weakly similar to GB|AAO39919.1|28372874|BT003691 At2g04235
{Arabidopsis thaliana;} , partial (41%)
Length = 958
Score = 28.9 bits (63), Expect = 3.5
Identities = 18/64 (28%), Positives = 35/64 (54%), Gaps = 10/64 (15%)
Frame = +1
Query: 189 LTETLDYSSAVILLGYSLILAILRSFNVRDEATR----VMVSAPLIAFVIT------HVM 238
L + L ++ + +GYS I+ + R + +A+ +++ APLIAFV+ H++
Sbjct: 718 LVDELRTAAESVRVGYSRIIRLCRCISQVVQASTKSR*LVIMAPLIAFVVAMIVH*FHLV 897
Query: 239 YINF 242
Y+ F
Sbjct: 898 YLRF 909
>TC215052 similar to UP|Q9XH69 (Q9XH69) Beta-amylase (Fragment), partial (92%)
Length = 2020
Score = 28.5 bits (62), Expect = 4.6
Identities = 12/30 (40%), Positives = 15/30 (50%)
Frame = +3
Query: 295 IYDFPPYEGLLDAHALWHATTIPLTYIWWS 324
IYD PPY G + A W +T W+S
Sbjct: 1059 IYDQPPYNGFFNDGASWESTYGDFFLSWYS 1148
>TC203661 similar to GB|AAO63400.1|28950953|BT005336 At3g26700 {Arabidopsis
thaliana;} , partial (88%)
Length = 2451
Score = 28.5 bits (62), Expect = 4.6
Identities = 10/22 (45%), Positives = 15/22 (67%)
Frame = -1
Query: 86 YHCMLDREKERKLLNLVPVKYH 107
+HCMLD +LL++VP +H
Sbjct: 1818 FHCMLDNPDSHELLSIVPSVHH 1753
>TC225444 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 750
Score = 28.5 bits (62), Expect = 4.6
Identities = 23/83 (27%), Positives = 39/83 (46%), Gaps = 11/83 (13%)
Frame = -2
Query: 222 RVMVSAPLIAFVITHVMYINFYKLDYGWNMIVCVV---------MAVVQLV--IWAVWAG 270
+V AP + F+I H+ + N + N + +V ++V L +W +WA
Sbjct: 671 KVTSEAPNMGFLIKHINHANKISVQKFQNSSI*IVAKNKNKT*KLSVSHLPHQVWDLWAW 492
Query: 271 LSGHPSRWKLWLVVIAGGLAMLL 293
HPS++ L LV+ L +LL
Sbjct: 491 ---HPSKYFLSLVIFGT*LQLLL 432
>TC229713
Length = 1177
Score = 28.1 bits (61), Expect = 6.0
Identities = 16/54 (29%), Positives = 26/54 (47%)
Frame = +2
Query: 131 LAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHS 184
+A F W+SFF+LL Y L G+ ++ + + F+S +FHS
Sbjct: 764 IAPQFVIWISFFLLLLYPSLL------IMHVCGICLMFSCFLILAVFYSTIFHS 907
>BI699853
Length = 402
Score = 28.1 bits (61), Expect = 6.0
Identities = 11/19 (57%), Positives = 13/19 (67%)
Frame = -3
Query: 159 YEYAGLWHIYGLLSLNSWF 177
+ Y LW I LL+LNSWF
Sbjct: 85 FVYVYLWSIESLLTLNSWF 29
>TC210622 homologue to UP|R10C_ARATH (P59231) 60S ribosomal protein L10a-3,
partial (96%)
Length = 864
Score = 28.1 bits (61), Expect = 6.0
Identities = 10/31 (32%), Positives = 18/31 (57%)
Frame = +3
Query: 79 DCQNDCRYHCMLDREKERKLLNLVPVKYHGK 109
+C C++ C + EK K L+P +Y+G+
Sbjct: 618 ECTTQCQFPCFIIEEKLAKREVLIPEEYYGE 710
>TC204868 similar to UP|Q9LKY8 (Q9LKY8) Proline-rich protein, partial (91%)
Length = 865
Score = 28.1 bits (61), Expect = 6.0
Identities = 13/40 (32%), Positives = 20/40 (49%), Gaps = 4/40 (10%)
Frame = -3
Query: 249 WNMIVCVVMAVVQLVIWAVWAG----LSGHPSRWKLWLVV 284
W + V+ V++LV+W VWA P W +W+ V
Sbjct: 386 WVWVWVWVVLVLELVVWVVWASEEKVEDNQPWVWVVWVWV 267
>TC211400 similar to UP|Q9FKJ0 (Q9FKJ0) Emb|CAB55405.1, partial (52%)
Length = 767
Score = 27.7 bits (60), Expect = 7.8
Identities = 11/36 (30%), Positives = 21/36 (57%), Gaps = 4/36 (11%)
Frame = -2
Query: 247 YGWNMIVCVVMAVVQLVIW----AVWAGLSGHPSRW 278
+G ++VC++ A++ + V+ G+ GHPS W
Sbjct: 430 HGQGLLVCMIKALITRYLR*RNPPVYLGVKGHPSSW 323
>BF595370
Length = 415
Score = 27.7 bits (60), Expect = 7.8
Identities = 11/35 (31%), Positives = 19/35 (53%)
Frame = +2
Query: 78 WDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPF 112
W+C N +L + + LL L+ +K++ WPF
Sbjct: 170 WNCLNPFHQDQILSADHSQILLMLLHIKHNPLWPF 274
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.142 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,439,794
Number of Sequences: 63676
Number of extensions: 392102
Number of successful extensions: 2680
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 2651
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2676
length of query: 342
length of database: 12,639,632
effective HSP length: 98
effective length of query: 244
effective length of database: 6,399,384
effective search space: 1561449696
effective search space used: 1561449696
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0176a.2


