

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0173b.3
(286 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
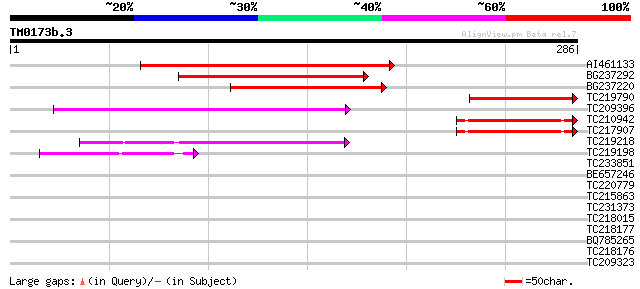
Score E
Sequences producing significant alignments: (bits) Value
AI461133 204 3e-53
BG237292 159 2e-39
BG237220 110 6e-25
TC219790 similar to GB|AAR24650.1|38603810|BT010872 At5g63160 {A... 92 2e-19
TC209396 weakly similar to GB|AAS77485.1|45825155|BT012360 At3g5... 67 1e-11
TC210942 58 6e-09
TC217907 similar to UP|Q9C5X9 (Q9C5X9) P300/CBP acetyltransferas... 57 1e-08
TC219218 weakly similar to UP|Q7F241 (Q7F241) Speckle-type POZ p... 46 2e-05
TC219198 weakly similar to UP|Q6ZAS6 (Q6ZAS6) POZ domain protein... 41 5e-04
TC233851 similar to UP|Q6NN60 (Q6NN60) RH61815p, partial (3%) 33 0.15
BE657246 similar to GP|6503306|gb| ESTs gb|AI993522 gb|T88335 a... 33 0.15
TC220779 similar to UP|Q6T285 (Q6T285) Predicted protein, partia... 33 0.15
TC215863 weakly similar to UP|Q70LK1 (Q70LK1) NADH5 protein, par... 32 0.25
TC231373 30 0.96
TC218015 similar to UP|Q9FN02 (Q9FN02) Ser/thr protein phosphata... 30 0.96
TC218177 similar to PIR|T52422|T52422 alternative oxidase-relate... 29 2.8
BQ785265 29 2.8
TC218176 similar to UP|Q9FEC9 (Q9FEC9) Plastid quinol oxidase (P... 29 2.8
TC209323 similar to GB|AAB64484.1|602374|SCE9871 Yel007wp {Sacch... 28 6.2
>AI461133
Length = 384
Score = 204 bits (520), Expect = 3e-53
Identities = 95/128 (74%), Positives = 111/128 (86%)
Frame = +1
Query: 67 AVTAFLTFLYSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQ 126
AVTAF+ FLYS RCTE+E+D+YGMHLLALSHVYMV LKQRC KGL+ R+ TE+VVD+LQ
Sbjct: 1 AVTAFVRFLYSSRCTEEEIDQYGMHLLALSHVYMVQQLKQRCIKGLTHRLTTESVVDVLQ 180
Query: 127 LARLCDAPDLHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEVLRFMDESETRREKSR 186
LARLCDAPDLHLRCM L +NFK VE TEGW+FL KHDPWLEL++LRF+DE ETR++KSR
Sbjct: 181 LARLCDAPDLHLRCMKLLAKNFKAVEATEGWKFLVKHDPWLELDILRFIDEHETRKKKSR 360
Query: 187 RSREVQGL 194
R R QG+
Sbjct: 361 RYRMEQGI 384
>BG237292
Length = 425
Score = 159 bits (401), Expect = 2e-39
Identities = 73/96 (76%), Positives = 82/96 (85%)
Frame = +1
Query: 86 DRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLT 145
D+YGMHLLALSHVY+VP LKQRC KGL+ R+ TENVVD+LQLARLCDAPDLHLRCM L
Sbjct: 1 DKYGMHLLALSHVYIVPQLKQRCIKGLTHRLTTENVVDVLQLARLCDAPDLHLRCMKLLA 180
Query: 146 RNFKTVEETEGWRFLTKHDPWLELEVLRFMDESETR 181
NFK VE TEGW+FL KHDP LEL++LRF+DE E R
Sbjct: 181 NNFKAVEATEGWKFLXKHDPCLELDILRFIDEHENR 288
>BG237220
Length = 450
Score = 110 bits (276), Expect = 6e-25
Identities = 50/79 (63%), Positives = 64/79 (80%)
Frame = +1
Query: 112 LSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEV 171
L+QR++TENVVD+LQL+RLCDAPDL+L+C+ L FK V+ETEGW+FL HDPWLEL+V
Sbjct: 1 LAQRLSTENVVDVLQLSRLCDAPDLYLKCVKLLRNRFKAVKETEGWKFLESHDPWLELDV 180
Query: 172 LRFMDESETRREKSRRSRE 190
LR M E E R+ + R+ RE
Sbjct: 181 LRLMGELEKRKRRVRKWRE 237
>TC219790 similar to GB|AAR24650.1|38603810|BT010872 At5g63160 {Arabidopsis
thaliana;} , partial (11%)
Length = 598
Score = 92.4 bits (228), Expect = 2e-19
Identities = 40/55 (72%), Positives = 45/55 (81%), Gaps = 1/55 (1%)
Frame = +3
Query: 233 FATCQGVQQLIRHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHAD-SCNVPLCR 286
FATCQG+Q LIRH+ATC K++ GGC+RCKRMWQLFRLHSY C D SC VP CR
Sbjct: 3 FATCQGLQVLIRHFATCEKKVRGGCVRCKRMWQLFRLHSYVCHDTDSSCKVPFCR 167
>TC209396 weakly similar to GB|AAS77485.1|45825155|BT012360 At3g56230
{Arabidopsis thaliana;} , partial (44%)
Length = 1116
Score = 66.6 bits (161), Expect = 1e-11
Identities = 34/150 (22%), Positives = 76/150 (50%)
Frame = +2
Query: 23 DGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTE 82
+G IPAH ++LA+ S + ++M++ + I I + + + + L FLYS
Sbjct: 551 NGPPIPAHKSVLAARSEIFKNMLECDECKAAPSNAITIPDLNHEELESLLEFLYSGTLNV 730
Query: 83 DEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMN 142
++++++ L + Y++PHL + C + L ++T N ++ L++A C +L +N
Sbjct: 731 EKLEKHVYALSQAADKYVIPHLLKHCERYLLSSLSTSNALETLEIADTCSNHNLKETTLN 910
Query: 143 HLTRNFKTVEETEGWRFLTKHDPWLELEVL 172
L +N + + + + P L ++++
Sbjct: 911 FLVKNIEHMVSSPKFEAFVHRSPHLTVQLV 1000
>TC210942
Length = 823
Score = 57.8 bits (138), Expect = 6e-09
Identities = 25/61 (40%), Positives = 38/61 (61%)
Frame = +2
Query: 226 KSGPCNKFATCQGVQQLIRHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSCNVPLC 285
+S C ++ C+ V+ L RH C R GGC+ CK+MW L +LH+ C+ ++ C+VP C
Sbjct: 59 RSAHC-QYPNCRKVKGLFRHGMHCKTRASGGCVLCKKMWYLLQLHARACKESE-CHVPRC 232
Query: 286 R 286
R
Sbjct: 233 R 235
>TC217907 similar to UP|Q9C5X9 (Q9C5X9) P300/CBP acetyltransferase-related
protein 2, partial (14%)
Length = 1911
Score = 57.0 bits (136), Expect = 1e-08
Identities = 25/61 (40%), Positives = 38/61 (61%)
Frame = +1
Query: 226 KSGPCNKFATCQGVQQLIRHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSCNVPLC 285
+S C ++ C+ V+ L RH C R GGC+ CK+MW L +LH+ C+ ++ C+VP C
Sbjct: 520 RSAHC-QYPNCRKVKGLFRHGMHCKIRASGGCVLCKKMWYLLQLHARACKESE-CHVPRC 693
Query: 286 R 286
R
Sbjct: 694 R 696
>TC219218 weakly similar to UP|Q7F241 (Q7F241) Speckle-type POZ protein-like
protein, partial (59%)
Length = 837
Score = 45.8 bits (107), Expect = 2e-05
Identities = 35/136 (25%), Positives = 60/136 (43%)
Frame = +1
Query: 36 SMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEMDRYGMHLLAL 95
S SPV +M+ R S I +I V D + AF+ +LY+ + D + +LL L
Sbjct: 220 SRSPVFRAMLKSDMTERRSGTI-KISDVSYDTLHAFVNYLYTAEASLD--NELACNLLVL 390
Query: 96 SHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLTRNFKTVEETE 155
Y V HLK C K L ++N + + A + L + + + ++ + E
Sbjct: 391 GEKYQVKHLKTYCEKYLIAKMNWDKAISNYAFAYQYNCKQLQSVSLAVILDHMDSLTQNE 570
Query: 156 GWRFLTKHDPWLELEV 171
+ L +P L +E+
Sbjct: 571 CYAELVDTNPRLVVEI 618
>TC219198 weakly similar to UP|Q6ZAS6 (Q6ZAS6) POZ domain protein
family-like, partial (53%)
Length = 1212
Score = 41.2 bits (95), Expect = 5e-04
Identities = 28/80 (35%), Positives = 43/80 (53%)
Frame = +3
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
D+ I T+DG+ + AH +L++ SPV +SM K + S I I + ++ TA L++L
Sbjct: 534 DLTIMTADGSTLRAHKAVLSASSPVFQSMYHLNLKEK*SS-TIHIEDMSLESCTALLSYL 710
Query: 76 YSRRCTEDEMDRYGMHLLAL 95
Y ED + H LAL
Sbjct: 711 YGTIKQED----FWKHRLAL 758
>TC233851 similar to UP|Q6NN60 (Q6NN60) RH61815p, partial (3%)
Length = 433
Score = 33.1 bits (74), Expect = 0.15
Identities = 15/39 (38%), Positives = 22/39 (55%)
Frame = +3
Query: 19 IHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERI 57
+ SD IPAH ++L S SPV ++M++ R S I
Sbjct: 315 VDDSDAVPIPAHKHLLVSRSPVFKAMLENDMAERRSGTI 431
>BE657246 similar to GP|6503306|gb| ESTs gb|AI993522 gb|T88335 and
gb|AA540932 come from this gene. {Arabidopsis thaliana},
partial (58%)
Length = 653
Score = 33.1 bits (74), Expect = 0.15
Identities = 18/62 (29%), Positives = 33/62 (53%)
Frame = -1
Query: 147 NFKTVEETEGWRFLTKHDPWLELEVLRFMDESETRREKSRRSREVQGLYAQLSEAMECLD 206
+F++ +GW + K + ++V+ M + E +R+ RE+ GL QL E CL+
Sbjct: 476 DFESANYPKGW-LIGKKRKLVNVDVVESMRRIAIQ-EMNRKDREIDGLNEQLEEDSRCLE 303
Query: 207 HI 208
H+
Sbjct: 302 HL 297
>TC220779 similar to UP|Q6T285 (Q6T285) Predicted protein, partial (71%)
Length = 769
Score = 33.1 bits (74), Expect = 0.15
Identities = 18/62 (29%), Positives = 33/62 (53%)
Frame = +3
Query: 147 NFKTVEETEGWRFLTKHDPWLELEVLRFMDESETRREKSRRSREVQGLYAQLSEAMECLD 206
+F++ +GW + K + ++V+ M + E +R+ RE+ GL QL E CL+
Sbjct: 402 DFESANYPKGW-LIGKKRKLVNVDVVESMRRIAIQ-EMNRKDREIDGLNEQLEEDSRCLE 575
Query: 207 HI 208
H+
Sbjct: 576 HL 581
>TC215863 weakly similar to UP|Q70LK1 (Q70LK1) NADH5 protein, partial (3%)
Length = 473
Score = 32.3 bits (72), Expect = 0.25
Identities = 26/99 (26%), Positives = 37/99 (37%), Gaps = 5/99 (5%)
Frame = -2
Query: 90 MHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARL-CDAPDLHLRCMNHLTRN- 147
MH LALS VYM R T + ++ + DA ++H C NH
Sbjct: 412 MHSLALSRVYMA-------------RKKTGFSLGAIETPKFTSDASNVHFHCPNHWNAAE 272
Query: 148 ---FKTVEETEGWRFLTKHDPWLELEVLRFMDESETRRE 183
F T W H P LE + R+ + T+ +
Sbjct: 271 PYLFGTDRRAHNWSLHRTHSPNLEKKRCRYRSNTATKNQ 155
>TC231373
Length = 741
Score = 30.4 bits (67), Expect = 0.96
Identities = 15/39 (38%), Positives = 22/39 (55%)
Frame = +3
Query: 156 GWRFLTKHDPWLELEVLRFMDESETRREKSRRSREVQGL 194
G ++LT W+EL V+ M+ S KSR SR ++ L
Sbjct: 207 GLKYLTSGAEWVELFVIEMMNASNMDDAKSRASRMLEAL 323
>TC218015 similar to UP|Q9FN02 (Q9FN02) Ser/thr protein phosphatase catalytic
subunit-like protein, partial (41%)
Length = 881
Score = 30.4 bits (67), Expect = 0.96
Identities = 16/56 (28%), Positives = 29/56 (51%)
Frame = +3
Query: 80 CTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPD 135
CTE+ + +Y + L+ SH K+ +G+ + ++VVD +L + APD
Sbjct: 237 CTEEFLKKYQLKLIIRSHEGPDAREKRDGLEGMDEGYTIDHVVDSGKLVTVFSAPD 404
>TC218177 similar to PIR|T52422|T52422 alternative oxidase-related protein
IMMUTANS [validated] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (70%)
Length = 1084
Score = 28.9 bits (63), Expect = 2.8
Identities = 22/79 (27%), Positives = 30/79 (37%), Gaps = 6/79 (7%)
Frame = +3
Query: 172 LRFMDESETRREKSRRSREVQGLYAQL-----SEAMECLD-HICTEGCTLVGPHHVEVDR 225
L DE +T R + R +++ LY EA C C L PH D
Sbjct: 837 LYLFDEFQTSRVPNSRRPKIENLYDVFVNIRDDEAEHCKTMKACQTHGNLRSPHSYAEDD 1016
Query: 226 KSGPCNKFATCQGVQQLIR 244
S C A C+G+ I+
Sbjct: 1017DSSVCALEADCEGIVDCIK 1073
>BQ785265
Length = 426
Score = 28.9 bits (63), Expect = 2.8
Identities = 19/66 (28%), Positives = 31/66 (46%), Gaps = 4/66 (6%)
Frame = +3
Query: 6 EQTTRVEPEPDIFIHTSDGTRIPAHSNILASMSPV----LESMIDRPRKHRSSERIIQIH 61
EQT +P+ D SDG P SN +S PV L + I +S+++ + H
Sbjct: 108 EQTKEQKPQVDNTSTQSDGKDEPVRSNGCSSQLPVYPLLLSTAIPEKLNENTSDKLRKGH 287
Query: 62 GVPGDA 67
+ G++
Sbjct: 288 AIVGNS 305
>TC218176 similar to UP|Q9FEC9 (Q9FEC9) Plastid quinol oxidase (Plastid
terminal oxidase), partial (34%)
Length = 701
Score = 28.9 bits (63), Expect = 2.8
Identities = 22/79 (27%), Positives = 30/79 (37%), Gaps = 6/79 (7%)
Frame = +3
Query: 172 LRFMDESETRREKSRRSREVQGLYAQL-----SEAMECLD-HICTEGCTLVGPHHVEVDR 225
L DE +T R + R +++ LY EA C C L PH D
Sbjct: 177 LYLFDEFQTSRVPNSRRPKIENLYDVFVNIRDDEAEHCKTMKACQTHGNLRSPHSYAEDD 356
Query: 226 KSGPCNKFATCQGVQQLIR 244
S C A C+G+ I+
Sbjct: 357 DSSVCALEADCEGIVDCIK 413
>TC209323 similar to GB|AAB64484.1|602374|SCE9871 Yel007wp {Saccharomyces
cerevisiae;} , partial (3%)
Length = 814
Score = 27.7 bits (60), Expect = 6.2
Identities = 20/65 (30%), Positives = 27/65 (40%), Gaps = 8/65 (12%)
Frame = -1
Query: 204 CLDHICTEGCTLVGPHHVEVDRKSGPCNKFATCQGV--------QQLIRHYATCTKRMGG 255
C+D + +G LVGPH V V P N+ + + L A C GG
Sbjct: 304 CIDRLDHDGAALVGPHRVRV----VPSNRLRHAKAYFRQ*TER*RWLRTVDAVCD---GG 146
Query: 256 GCLRC 260
GC+ C
Sbjct: 145 GCIGC 131
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.136 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,437,531
Number of Sequences: 63676
Number of extensions: 245403
Number of successful extensions: 1520
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 1513
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1520
length of query: 286
length of database: 12,639,632
effective HSP length: 96
effective length of query: 190
effective length of database: 6,526,736
effective search space: 1240079840
effective search space used: 1240079840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0173b.3


