

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
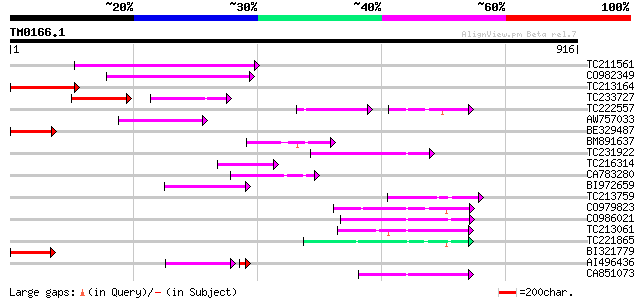
Query= TM0166.1
(916 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 206 5e-53
CO982349 157 3e-38
TC213164 108 8e-24
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 77 4e-22
TC222557 55 4e-18
AW757033 87 4e-17
BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase... 86 7e-17
BM891637 86 1e-16
TC231922 84 4e-16
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 78 2e-14
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 78 2e-14
BI972659 77 3e-14
TC213759 76 6e-14
CO979823 75 1e-13
CO986021 72 1e-12
TC213061 71 2e-12
TC221865 70 4e-12
BI321779 69 9e-12
AI496436 66 2e-11
CA851073 66 6e-11
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 206 bits (523), Expect = 5e-53
Identities = 113/301 (37%), Positives = 175/301 (57%), Gaps = 3/301 (0%)
Frame = +1
Query: 106 LIGNEVVDTMLSSGRGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTA 165
LI NEV+D S + + K+D+ KAYD++ W+FLM MM + F P+W WI CV +A
Sbjct: 13 LIANEVIDEAKRSNKSCLVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIEECVKSA 192
Query: 166 TLAVLVNGSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRE---RPCGVVLG 222
+++VLVNGSPT F ++GLRQGDPL+P LFN+ +GL+ + R A+ E +P +V
Sbjct: 193 SISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRR-AVEENLYKPY-LVGA 366
Query: 223 PGVCLNHLQFADDTLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIV 282
GV ++ LQ+ADDT+ F + +E + ++ +F S L++N+AKS V
Sbjct: 367 NGVPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKSCFGVFGVTDQWK 546
Query: 283 NQLAVELGVVVGTLPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVV 342
+ A L + P YLG +G + RR W +I K L+ +S GR+ +
Sbjct: 547 QEAANYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKCERKLSKWK*RHISFGGRVTL 726
Query: 343 LRSVASAIPIYHMAVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGG 402
++SV ++IPIY + ++ P VV + K+ R FLWG ++I+W+ WE++C KE GG
Sbjct: 727 IKSVLTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGGGPDQKKISWIRWEKLCLPKERGG 906
Query: 403 L 403
+
Sbjct: 907 I 909
>CO982349
Length = 795
Score = 157 bits (396), Expect = 3e-38
Identities = 85/240 (35%), Positives = 133/240 (55%), Gaps = 1/240 (0%)
Frame = +3
Query: 157 WIHSCVSTATLAVLVNGSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERP 216
WI C+ +A++++LVN SP F ++GLRQGDPL+PLLFN+ +GL+ + R+R
Sbjct: 69 WIRGCLHSASVSILVNESPINEFSPQRGLRQGDPLAPLLFNIVAEGLTGLMREAVDRKRF 248
Query: 217 CGVVLGPGV-CLNHLQFADDTLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGC 275
++G ++ LQ+ADDT+ F + M+ + ++ F SGLK+N+A+S
Sbjct: 249 NSFLVGKNKEPVSILQYADDTIFFEEATMENMKVIKTILRCFELASGLKINFAESRFGAI 428
Query: 276 NVDSAIVNQLAVELGVVVGTLPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVS 335
+ A L + ++P YLG + +PR R P+I K LA +S
Sbjct: 429 WKPDHWCKEAAEFLNCSMLSMPFSYLGIPIEANPRCREI*DPIIRKCETKLARWKQRHIS 608
Query: 336 LSGRLVVLRSVASAIPIYHMAVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVC 395
L GR+ ++ ++ +A+PIY + ++AP VV + + R FLWG RRIAWV WE VC
Sbjct: 609 LGGRVTLINAILTALPIYFFSFFRAPSKVVQKLVNIQRKFLWGGGSXQRRIAWVKWETVC 788
>TC213164
Length = 446
Score = 108 bits (271), Expect = 8e-24
Identities = 49/113 (43%), Positives = 77/113 (67%)
Frame = +3
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG NF F +EFW ++K ++++ +EFY G +K NA F+ALIPK P + +Y+
Sbjct: 84 PGPDGLNFKFIKEFWDILKTDLLRFLDEFYANGVFLKRSNAFFLALIPKVHDP*SLNEYK 263
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVD 113
ISLIG IYK++AK+L+ R + V+P ++ E Q F +GR + ++I NE+++
Sbjct: 264 PISLIGCIYKIVAKLLSGRHKKVLPTIIDERQTVFMEGRHTLHNVVIANEIME 422
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 77.0 bits (188), Expect(2) = 4e-22
Identities = 37/97 (38%), Positives = 61/97 (62%), Gaps = 1/97 (1%)
Frame = -3
Query: 101 IANCLLIGNEVVDTMLSSGRGGFL-LKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIH 159
I +C + +EV++ + GG L LK+D KA+D ++ DFL++++ G+ + WI
Sbjct: 834 IKDCTCVTSEVINMLDKKVFGGNLALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNWIR 655
Query: 160 SCVSTATLAVLVNGSPTEFFEIEKGLRQGDPLSPLLF 196
+ +A L++ VNG P FF ++G+RQGDPLSP L+
Sbjct: 654 VILLSAKLSISVNGEPVGFFSCQRGVRQGDPLSPSLY 544
Score = 47.0 bits (110), Expect(2) = 4e-22
Identities = 31/132 (23%), Positives = 62/132 (46%), Gaps = 1/132 (0%)
Frame = -2
Query: 228 NHLQFADDTLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAV 287
+H+ ADD ++FC + L + + G ++ KS++ ++ + ++
Sbjct: 451 SHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIFIGSIPGQRLRVISD 272
Query: 288 ELGVVVGTLPLPYLG-SVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSV 346
L +P Y G + G P RR + V +K++ LAS + +S+ GR+ ++ SV
Sbjct: 271 LLQYFHSDIPFNYFGVPIFKGKPTRRHLYC-VADKIKCKLASWKGSILSIMGRIQLVNSV 95
Query: 347 ASAIPIYHMAVY 358
+ +Y +VY
Sbjct: 94 IHGMLLYSFSVY 59
>TC222557
Length = 1002
Score = 55.5 bits (132), Expect(2) = 4e-18
Identities = 36/125 (28%), Positives = 59/125 (46%), Gaps = 2/125 (1%)
Frame = +3
Query: 464 DIVNVVLEDSVLGKTFRDQCFCVVGDGLRISFWRDPWVASKT-LAVLYPRFYAMASNQDI 522
D+ +V+ E +V G F+ VG G +I FW D W+ + L YPR Y ++ Q
Sbjct: 9 DLRSVIHEQAVHG-LFQSAIDWKVGCGDKIRFWEDCWLMEQEPLRAKYPRLYNISCQQQK 185
Query: 523 MVSEAGVFSNLGWEWNLGFRRRFFGWEEQVYDELIARCN-GVVPSREKDSLIWLGDTNGR 581
++ G S WEW L +RR F E Q + + G + D +W + NG+
Sbjct: 186 LILVMGFHSANVWEWKLEWRRHLFDNEVQAAASFLDDISWGHIDRWTLDCWVWKPEPNGQ 365
Query: 582 YTVKT 586
++ ++
Sbjct: 366 FSTRS 380
Score = 55.1 bits (131), Expect(2) = 4e-18
Identities = 36/144 (25%), Positives = 60/144 (41%), Gaps = 6/144 (4%)
Frame = +1
Query: 612 PQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQAEESVTHLILVCKFAWVLWS 671
P K F W+ + R+ NL R V ++ +C C + EE +HL C LW
Sbjct: 457 PLKATTFAWRLIKERIPTKGNLWRRRVQLN-NLMCPFCNRQEEEASHLFFNCPRILPLWW 633
Query: 672 KVLLREGVVWCVPASILSLLKEWDSLR-----KGSDL-VLWRLIPYALAWSVWMARNGLI 725
E + W + S + + ++ KG L W+ AL WS+W RN ++
Sbjct: 634 -----ESLAWTKTVGVFSPIPRQNYMQHTLISKGKHLQARWKCWWVALTWSIWRHRNRIV 798
Query: 726 FKGEEVSVETVWDIHITRMYWWIK 749
F + + + D I ++ W +
Sbjct: 799 FLNKAFNGSKLMDDAIFLVWSWFR 870
>AW757033
Length = 441
Score = 86.7 bits (213), Expect = 4e-17
Identities = 50/145 (34%), Positives = 76/145 (51%), Gaps = 1/145 (0%)
Frame = -2
Query: 176 TEFFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGP-GVCLNHLQFAD 234
T F +GLRQGDPL+PLLFN+ +GL+ + R +G ++ LQ+AD
Sbjct: 440 TREFSPHRGLRQGDPLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKKPVSILQYAD 261
Query: 235 DTLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVG 294
DT+ F + + + ++ F SGLK+N+AKS +V + A + +
Sbjct: 260 DTIFFGEATMENVRVIKTILRGFELASGLKINFAKSRFGAISVPD*WRKEAAEFMNCSLL 81
Query: 295 TLPLPYLGSVLGGDPRRRGFWAPVI 319
+LP YLG +G +PRRR W P+I
Sbjct: 80 SLPFSYLGIPIGANPRRRETWDPII 6
>BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (15%)
Length = 376
Score = 85.9 bits (211), Expect = 7e-17
Identities = 38/75 (50%), Positives = 52/75 (68%)
Frame = -2
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG NF F + FW +K +I++ EF+ G KG NA FIALIPK + P+ + D+R
Sbjct: 336 PGPDGLNFKFIKHFWERLKPDIIRFLNEFHANGIFPKGGNASFIALIPKVKHPQALNDFR 157
Query: 61 LISLIGSIYKLLAKV 75
ISLIG +YK++AK+
Sbjct: 156 PISLIGCVYKIVAKI 112
>BM891637
Length = 427
Score = 85.5 bits (210), Expect = 1e-16
Identities = 47/151 (31%), Positives = 80/151 (52%), Gaps = 7/151 (4%)
Frame = -1
Query: 383 GRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLLKWAWRFGQEPEALWRRVLCAKYGLN 442
G++IAW++W Q C ++GGLG++ +K N A+L+KW W +P LW R+L +KY
Sbjct: 421 GKKIAWISWRQCCASGDVGGLGIKDIKILNNALLIKWKWLMFHQPHQLWNRILISKY--- 251
Query: 443 SIPLVLHRVLEGL-RHPSRYMF-----DIVNVVLEDSVLGKTFRDQCFCVVGDGLRISFW 496
+ GL + P +Y F D+ + S++ + +Q VG G +I FW
Sbjct: 250 -------KGWRGLDQGPQKYYFSPWWADLRAINQHQSMIAAS--NQFCWKVGRGDQILFW 98
Query: 497 RDPWVASKT-LAVLYPRFYAMASNQDIMVSE 526
D WV T L +P Y ++S ++ ++++
Sbjct: 97 EDSWVDDGTPLKDQFPELYRLSSQRNFIMAD 5
>TC231922
Length = 803
Score = 83.6 bits (205), Expect = 4e-16
Identities = 53/202 (26%), Positives = 96/202 (47%), Gaps = 3/202 (1%)
Frame = -3
Query: 487 VGDGLRISFWRDPWVASK-TLAVLYPRFYAMASNQDIMVSEAGVFSNLGWEWNLGFRRRF 545
VG G +I FW+D W++ L Y + Y ++ ++ +S+ G FS W +RRR
Sbjct: 699 VGCGDKIKFWQDSWLSEGCNLQQKYNQLYTISRQ*NLTISKMGKFSQNARSWEFKWRRRL 520
Query: 546 FGWEEQVYDELIARCNGV-VPSREKDSLIWLGDTNGRYTVKTFCMEVELQLYGPPSWVVP 604
F +E + + + +G+ + ++ DS+ W D++G Y+ K+ + + PS +
Sbjct: 519 FDYEYAMAVDFMNEISGISIQNQAHDSMFWKADSSGVYSTKSAYRLLMPSISPAPSRRIF 340
Query: 605 SCVKKLA-PQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQAEESVTHLILVC 663
+ L P + +F W+ +R+ NL R + + + +C LCG E HL C
Sbjct: 339 QILWHLKIPPRAAVFSWRLFLDRLPTRGNLSRRSIPIQ-DIMCPLCGCQHEEAGHLFFHC 163
Query: 664 KFAWVLWSKVLLREGVVWCVPA 685
K LW + + V+ +PA
Sbjct: 162 KMTKGLWWESMRWIRVIGALPA 97
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 78.2 bits (191), Expect = 2e-14
Identities = 36/99 (36%), Positives = 54/99 (54%)
Frame = +2
Query: 336 LSGRLVVLRSVASAIPIYHMAVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVC 395
+ GR+ +++SV SA+PI ++ +K P +V + L R FLWG + RI WV W +C
Sbjct: 2 MGGRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADIC 181
Query: 396 KRKEIGGLGVEFLKFRNKAMLLKWAWRFGQEPEALWRRV 434
K GGLG++ L N A+ +W W LW R+
Sbjct: 182 NPKIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWARL 298
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 77.8 bits (190), Expect = 2e-14
Identities = 47/143 (32%), Positives = 68/143 (46%)
Frame = +2
Query: 358 YKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLL 417
+ P ++A +E L R FLWG R+IAWV W+ VC K GGLG++ L+ N +L
Sbjct: 11 FSLP*KIIAKLESLQRRFLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTTLLG 190
Query: 418 KWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVLGK 477
KW W + W +VL +KYG L G + + + I L+ ++
Sbjct: 191 KWRWDLFYIQQEPWAKVLQSKYGGWR---ALEEGSSGSKDSAWWKDLIKTQQLQRNI--- 352
Query: 478 TFRDQCFCVVGDGLRISFWRDPW 500
+ + VG G RI FW D W
Sbjct: 353 PLKRETIWKVGGGDRIKFWEDLW 421
>BI972659
Length = 453
Score = 77.0 bits (188), Expect = 3e-14
Identities = 43/140 (30%), Positives = 70/140 (49%)
Frame = +1
Query: 250 LFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGTLPLPYLGSVLGGDP 309
L ++ F SGL++NYAKS+ + A L +P YLG +
Sbjct: 34 LKSMLRGFEMVSGLRINYAKSQFGIVGFQPNWAHDAAQLLNCRQLDIPFHYLGMPIAVKA 213
Query: 310 RRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAVYKAPLGVVADIE 369
R W P+I K + L+ +S+ GR+ +++SV +A+PIY ++ +K P +V +
Sbjct: 214 SSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFFKIPQRIVDKLV 393
Query: 370 KLMRCFLWGRSGGGRRIAWV 389
L R F+WG + RI+WV
Sbjct: 394 SLQRTFMWGGNQHHNRISWV 453
>TC213759
Length = 681
Score = 76.3 bits (186), Expect = 6e-14
Identities = 47/156 (30%), Positives = 75/156 (47%), Gaps = 1/156 (0%)
Frame = +2
Query: 611 APQKVVLFLWQAAENRVAVMVNLHNRGVL-VDAEAVCVLCGQAEESVTHLILVCKFAWVL 669
AP KV++F WQ NR L R + VDA+ C +C + + THL C+F + +
Sbjct: 8 APLKVIVFGWQVILNRTQTREALF*RNIGGVDADLSCPMCHGDDVTTTHLFS-CRFGYSV 184
Query: 670 WSKVLLREGVVWCVPASILSLLKEWDSLRKGSDLVLWRLIPYALAWSVWMARNGLIFKGE 729
W + R G +P S + W G + W ++ W +W+ RN +IF+G+
Sbjct: 185 WQRCYQRLGFEQVLPGSSCAYF--WHHQGYGHWMTFWLVV----IWLIWLQRNTIIFRGK 346
Query: 730 EVSVETVWDIHITRMYWWIKAGKPHCSFSLCDFLQN 765
+ V+ V D + + + W+KA +FS D L N
Sbjct: 347 QACVQEVVDSVVFKTWSWVKAYYARDTFSFSD*LLN 454
>CO979823
Length = 853
Score = 75.1 bits (183), Expect = 1e-13
Identities = 59/236 (25%), Positives = 96/236 (40%), Gaps = 9/236 (3%)
Frame = -2
Query: 524 VSEAGVFSNLGWEWNLGFRRRFFGWEEQVYDELIARCNG--VVPSREKDSLIWLGDTNGR 581
+ + G +N GWE L +RR E + D + + P RE D +W + G
Sbjct: 843 IQQLGTITNSGWE*QLNWRRPXXDSEIAMADSFLGEITQQQIHPQRE-DKWLWKPEPGGH 667
Query: 582 YTVKTFCMEVELQLYGPPSWVVPSCVKKLA-PQKVVLFLWQAAENRVAVMVNLHNRGVLV 640
Y+ K+ + +L + + KL P K +F W+ +R+ NL R V+V
Sbjct: 666 YSTKSGYHVLWGELTEEIQDADFAEIWKLKIPTKAAVFAWRLVRDRLPTKSNLRRRQVMV 487
Query: 641 DAEAVCVLCGQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWDSLRKG 700
+ VC LC EE HL C LW E + W + + L+ G
Sbjct: 486 Q-DMVCPLCNNIEEGAAHLFFNCTKTLPLWW-----ESMSWVNLKTAMPQTPRQHFLQYG 325
Query: 701 SDLV------LWRLIPYALAWSVWMARNGLIFKGEEVSVETVWDIHITRMYWWIKA 750
+D+ W+ AL W++W RN ++F+ V + + ++ W KA
Sbjct: 324 TDIADGLKSKRWKCWWIALTWTIWQHRNKVVFQNATFHGNKVLEDALLLLWSWFKA 157
>CO986021
Length = 900
Score = 71.6 bits (174), Expect = 1e-12
Identities = 55/222 (24%), Positives = 96/222 (42%), Gaps = 6/222 (2%)
Frame = -1
Query: 535 WEWNLGFRRRFFGWEEQVYDELIARCNGV-VPSREKDSLIWLGDTNGRYTVKT-FCMEVE 592
W W+ +RR F E + + + + + + +DS++W D G Y+ K+ + + +
Sbjct: 891 WNWDFKWRRNLFDHENEQAIAFMDDISAISIHQQLQDSMLWKADPTGIYSTKSAYRLLLP 712
Query: 593 LQLYGPPSWVVPSCVKKLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQA 652
G S K P + LF W+ +R+++ NL R V + + +C LCG
Sbjct: 711 NNRPGQHSSNFKILWKLKIPPRAELFSWRLFRDRLSIRANLLRRHVALQ-DIMCPLCGNH 535
Query: 653 EESVTHLILVCKFAWVLWSKVL---LREGVVWCVPAS-ILSLLKEWDSLRKGSDLVLWRL 708
+E HL C+ LW + + G PA+ + + + R + LW +
Sbjct: 534 QEEAGHLFFHCRMTIGLWWESMNWTRTLGAFLDDPAAHFIQFSDGFGAQRNHNRRCLWWI 355
Query: 709 IPYALAWSVWMARNGLIFKGEEVSVETVWDIHITRMYWWIKA 750
AL ++W RN L+FKG V D + + W+KA
Sbjct: 354 ---ALTSTIWQHRNSLLFKGTPFQPPKVMDDALFHAWTWLKA 238
>TC213061
Length = 823
Score = 70.9 bits (172), Expect = 2e-12
Identities = 55/226 (24%), Positives = 92/226 (40%), Gaps = 6/226 (2%)
Frame = +2
Query: 530 FSNLGWEWNLGFRRRFFGWEEQVYDELIARCNG-VVPSREKDSLIWLGDTNGRYTVKTFC 588
F GWEW +RR F E ++ I G + S +D +W +++G Y ++
Sbjct: 5 FIETGWEWQFQWRRPLFDSEIEMLVAFIQEVEGHKIRSDIEDQWVWAAESSGSYLARSAY 184
Query: 589 MEVELQLYGPPSWVVPSCVKKL----APQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEA 644
+ G P K+L P KV +F W+ ++ + NL + V + E
Sbjct: 185 RVIR---EGIPEEEQDREFKELWKLKVPMKVTMFAWRLLKDILPTRDNLRRKRVELH-EY 352
Query: 645 VCVLCGQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWD-SLRKGSDL 703
VC LC +ES +HL C +W + L +V P + + +G
Sbjct: 353 VCPLCRSMDESASHLFFHCSKILPIWWESLSWVKLVGAFPHHPRHHFHQHSHEVYQGLQG 532
Query: 704 VLWRLIPYALAWSVWMARNGLIFKGEEVSVETVWDIHITRMYWWIK 749
W+ AL W++W RN +IF + V D + ++ W +
Sbjct: 533 NRWKWWWLALTWTIWKHRNDIIFSNATFNAHKVMDDAVFLIWTWFR 670
>TC221865
Length = 1045
Score = 70.1 bits (170), Expect = 4e-12
Identities = 64/289 (22%), Positives = 116/289 (39%), Gaps = 14/289 (4%)
Frame = -3
Query: 475 LGKTFRDQCFCVVGDGLRISFWRDPWVAS-KTLAVLYPRFYAMASNQDIMVSEAGVFSNL 533
L + +++ VG G + FW D V +TL YPR Y ++ Q ++ + G +
Sbjct: 959 LNRVLQNEIVWKVGCGDKFRFWEDKLVGEGETLMRKYPRMYQISCQQQQLIQQVGSHTET 780
Query: 534 GWEWNLGFRRRFFGWEEQV---YDELIARCNGVVPSREKDSLIWLGDTNGRYTVKTFCME 590
WEW +RR F E + E I+R + D +W + NG Y+ ++
Sbjct: 779 TWEWKFQWRRPLFDNEVDTTIGFLEDISRF--PIHRHLTDCWVWKSEPNGHYSTRSAYHL 606
Query: 591 VELQLYGPPSWVVPSCVKKL-APQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLC 649
++ + + + KL P K +F + ++R+ NL R V ++ +C+ C
Sbjct: 605 MQHEAAKANTDPAFEEIWKLKIPAKAAIFA*RLVKDRLPTKNNLRRRQVQLNG-TLCLFC 429
Query: 650 GQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWDSLRKGSDLV----- 704
EE +HL C +W E + W + S L+ L+
Sbjct: 428 RNYEEEASHLFFSCTKTQPVW-----WESLSWINTSGAFSANPRHHFLQHSFGLLGLKQY 264
Query: 705 ----LWRLIPYALAWSVWMARNGLIFKGEEVSVETVWDIHITRMYWWIK 749
W + AL +++W RN IF E + + + + ++ W +
Sbjct: 263 NR*KCWWV---ALTFTIWQHRNRTIFSNEPFNGSKMMEDAMFLVWSWFR 126
>BI321779
Length = 421
Score = 68.9 bits (167), Expect = 9e-12
Identities = 32/74 (43%), Positives = 47/74 (63%)
Frame = +2
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG++ + FW +IKEE++ F+EF++ L +G N FIALI K P + Y
Sbjct: 149 PGPDGYSLLLIKIFWEVIKEEVINFFKEFHKFDFLPRGTNRSFIALIAKCDNPLDIGQYC 328
Query: 61 LISLIGSIYKLLAK 74
ISL+G +YK++ K
Sbjct: 329 PISLVGCLYKIV*K 370
>AI496436
Length = 414
Score = 65.9 bits (159), Expect(2) = 2e-11
Identities = 41/114 (35%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Frame = +1
Query: 253 VMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVE-LGVVVGTLPLPYLGSVLGGDPRR 311
++ F SGLK+N+AKS+ G S I + A + L V ++P YLG +G + R
Sbjct: 4 ILRCFELSSGLKINFAKSQF-GTICKSDIWRKEAADYLNSSVLSMPFVYLGIPIGANSRH 180
Query: 312 RGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAVYKAPLGVV 365
W PV+ K LAS VS GR+ ++ SV +A+PIY ++ ++ P VV
Sbjct: 181 SDVWEPVVRKCERKLASWKQKYVSFGGRVTLINSVLTALPIYLLSFFRIPDKVV 342
Score = 21.9 bits (45), Expect(2) = 2e-11
Identities = 9/18 (50%), Positives = 11/18 (61%)
Frame = +2
Query: 371 LMRCFLWGRSGGGRRIAW 388
+ R FLWG R+IAW
Sbjct: 359 IQRRFLWGGDLE*RKIAW 412
>CA851073
Length = 633
Score = 66.2 bits (160), Expect = 6e-11
Identities = 48/188 (25%), Positives = 83/188 (43%), Gaps = 2/188 (1%)
Frame = +1
Query: 564 VPSREKDSLIWLGDTNGRYTVKT-FCMEVELQLYGPPSWVVPSCVKKLAPQKVVLFLWQA 622
+ S +D+++W D G Y+ K+ + + + + + + + K P + +F W+
Sbjct: 10 IQSHLQDTMVWKADPCGVYSTKSAYRILMTCNRHVSEANIFKTIWKLKIPPRAAVFSWRL 189
Query: 623 AENRVAVMVNLHNRGVLVDAEAVCVLCGQAEESVTHLILVCKFAWVLWSKVLLREGVVWC 682
++R+ NL R V + E C LCG +E HL CK LW + + +V
Sbjct: 190 IKDRLPTRHNLLRRNVSIQ-ENECPLCGYEQEEADHLFFNCKMTRGLWWESMRWNQIVGP 366
Query: 683 VPASILSLLKEW-DSLRKGSDLVLWRLIPYALAWSVWMARNGLIFKGEEVSVETVWDIHI 741
+ S S ++ D G + W AL S+W RN LIF+G + V D +
Sbjct: 367 LSVSPASHFVQFCDGFGAGRNHTRWCGWWIALTSSIWKHRNLLIFQGNQFEPSKVMDDAL 546
Query: 742 TRMYWWIK 749
+ W+K
Sbjct: 547 FLAWSWLK 570
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.142 0.462
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 49,970,973
Number of Sequences: 63676
Number of extensions: 838872
Number of successful extensions: 5325
Number of sequences better than 10.0: 133
Number of HSP's better than 10.0 without gapping: 5208
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5281
length of query: 916
length of database: 12,639,632
effective HSP length: 106
effective length of query: 810
effective length of database: 5,889,976
effective search space: 4770880560
effective search space used: 4770880560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0166.1


