

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0162a.10
(265 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
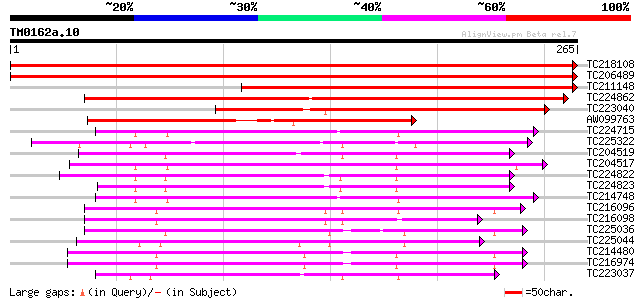
Score E
Sequences producing significant alignments: (bits) Value
TC218108 UP|NO26_SOYBN (P08995) Nodulin-26 (N-26), complete 371 e-103
TC206489 similar to UP|Q9XGG7 (Q9XGG7) Nodulin26-like major intr... 350 4e-97
TC211148 similar to UP|Q6QIL2 (Q6QIL2) NIP2, partial (58%) 216 8e-57
TC224862 similar to UP|Q6KAW0 (Q6KAW0) Nod26-like major intrinsi... 213 5e-56
TC223040 similar to UP|NI51_ARATH (Q9SV84) Probable aquaporin NI... 155 1e-38
AW099763 similar to PIR|JQ2285|JQ22 nodulin-26 - soybean, partia... 142 2e-34
TC224715 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%) 107 7e-24
TC225322 similar to UP|TI41_ARATH (O82316) Probable aquaporin TI... 105 2e-23
TC204519 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic... 104 4e-23
TC204517 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic... 103 1e-22
TC224822 similar to UP|TIP2_TOBAC (P24422) Probable aquaporin TI... 102 1e-22
TC224823 similar to UP|TIP1_TOBAC (P21653) Probable aquaporin TI... 102 2e-22
TC214748 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%) 101 4e-22
TC216096 homologue to UP|PI27_ARATH (P93004) Aquaporin PIP2.7 (P... 100 5e-22
TC216098 similar to UP|PI27_ARATH (P93004) Aquaporin PIP2.7 (Pla... 100 7e-22
TC225036 similar to UP|O65357 (O65357) Aquaporin 2, partial (92%) 100 7e-22
TC225044 homologue to UP|O22339 (O22339) Aquaporin-like transmem... 98 4e-21
TC214480 homologue to UP|Q39822 (Q39822) Pip1 protein, complete 97 1e-20
TC216974 homologue to UP|Q39822 (Q39822) Pip1 protein, complete 97 1e-20
TC223037 similar to UP|Q8H1Z5 (Q8H1Z5) Tonoplast intrinsic prote... 96 2e-20
>TC218108 UP|NO26_SOYBN (P08995) Nodulin-26 (N-26), complete
Length = 1122
Score = 371 bits (953), Expect = e-103
Identities = 178/265 (67%), Positives = 221/265 (83%)
Frame = +3
Query: 1 MANNSASFETHDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVV 60
MA+ SA E+ +VV++V K+ S+TI+ SD+ SV F+QKLVAE VGT+FLIF GCAS+VV
Sbjct: 93 MADYSAGTESQEVVVNVTKNTSETIQRSDSLVSVPFLQKLVAEAVGTYFLIFAGCASLVV 272
Query: 61 NKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIA 120
N+N N++T PGIA+VWGLVL VL+Y+VGHISG HFNPAVT AFA+T+RFP IQV Y+
Sbjct: 273 NENYYNMITFPGIAIVWGLVLTVLVYTVGHISGGHFNPAVTIAFASTRRFPLIQVPAYVV 452
Query: 121 SQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRA 180
+QLLG++LASG L++LF G HDQFSGT+P+GTNLQAFV EFI TF LMFVI VATDNRA
Sbjct: 453 AQLLGSILASGTLRLLFMGNHDQFSGTVPNGTNLQAFVFEFIMTFFLMFVICGVATDNRA 632
Query: 181 IGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAG 240
+GE+AGIAIGSTLLLN++I GP+TGASMNPAR+LGPA H +Y I +Y ++ + GA+AG
Sbjct: 633 VGELAGIAIGSTLLLNVIIGGPVTGASMNPARSLGPAFVHGEYEGIWIYLLAPVVGAIAG 812
Query: 241 AWVFNILRYTDKPLHEITKGSSILK 265
AWV+NI+RYTDKPL E TK +S LK
Sbjct: 813 AWVYNIVRYTDKPLSETTKSASFLK 887
>TC206489 similar to UP|Q9XGG7 (Q9XGG7) Nodulin26-like major intrinsic
protein, partial (86%)
Length = 1169
Score = 350 bits (898), Expect = 4e-97
Identities = 170/265 (64%), Positives = 214/265 (80%)
Frame = +1
Query: 1 MANNSASFETHDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVV 60
+A+NSA+ +H VVL+VN DA K + S V +QKLVAE VGT+FLIF GCAS+VV
Sbjct: 103 VADNSANNGSHQVVLNVNGDAPKKCDDSANQDCVPLLQKLVAEVVGTYFLIFAGCASVVV 282
Query: 61 NKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIA 120
N + D VVT PGI++VWGL +MVL+YSVGHISGAHFNPAVT A ATTKRFP QV Y+
Sbjct: 283 NLDKDKVVTQPGISIVWGLTVMVLVYSVGHISGAHFNPAVTIAHATTKRFPLKQVPAYVI 462
Query: 121 SQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRA 180
+Q++GA LASG L+++F+G +D F+GT+PSG++LQ+FV+EFI TF LMFVIS VATDNRA
Sbjct: 463 AQVVGATLASGTLRLIFNGKNDHFAGTLPSGSDLQSFVVEFIITFYLMFVISGVATDNRA 642
Query: 181 IGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAG 240
IGE+AG+A+GST+LLN++ +GPITGASMNPAR+LGPAI H +YR I +Y VS GAVAG
Sbjct: 643 IGELAGLAVGSTVLLNVMFAGPITGASMNPARSLGPAIVHHEYRGIWIYLVSPTLGAVAG 822
Query: 241 AWVFNILRYTDKPLHEITKGSSILK 265
W +N +RYT+KP+ EITK +S LK
Sbjct: 823 TWAYNFIRYTNKPVREITKSASFLK 897
>TC211148 similar to UP|Q6QIL2 (Q6QIL2) NIP2, partial (58%)
Length = 587
Score = 216 bits (550), Expect = 8e-57
Identities = 104/157 (66%), Positives = 128/157 (81%)
Frame = +2
Query: 109 RFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLM 168
RFP QV Y+ +Q++G+ LAS L++LFSG QFSGT+PSG+NLQAFVIEF+ TF LM
Sbjct: 11 RFPLKQVPVYVVAQVVGSTLASATLRLLFSGKETQFSGTLPSGSNLQAFVIEFLITFFLM 190
Query: 169 FVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVV 228
FVIS VATD+RAIGE+A IA+GST+LLN++ +GPITGASMNPAR +GPAI H++YR I +
Sbjct: 191 FVISGVATDDRAIGELAVIAVGSTVLLNVMFAGPITGASMNPARNIGPAILHNEYRGIWI 370
Query: 229 YFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
Y VS GAVAG WV+N +RYTDKPL EITK +S LK
Sbjct: 371 YIVSPTLGAVAGTWVYNTIRYTDKPLREITKSTSFLK 481
>TC224862 similar to UP|Q6KAW0 (Q6KAW0) Nod26-like major intrinsic protein,
partial (79%)
Length = 1192
Score = 213 bits (543), Expect = 5e-56
Identities = 102/226 (45%), Positives = 159/226 (70%)
Frame = +1
Query: 36 FVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAH 95
F +K+ AE +GTF L+F G S ++K ++++V+ G +L GL++ V+IYS+GHISGAH
Sbjct: 268 FPRKVFAEVIGTFLLVFVGSGSAGLSKIDESMVSKLGASLAGGLIVTVMIYSIGHISGAH 447
Query: 96 FNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQ 155
NPAV+ AF + PW Q+ Y+A+QL GA+ AS L+ L + D+ GT P+G+++Q
Sbjct: 448 MNPAVSLAFTAVRHLPWPQLPFYVAAQLTGAISASYTLRELLRPS-DEIGGTSPAGSHIQ 624
Query: 156 AFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTLG 215
A ++E ++T+ ++F+ AVATD+ A G+++G+A+GS++ + +++GPI+G SMNPARTLG
Sbjct: 625 ALIMEMVSTYTMVFISMAVATDSNATGQLSGVAVGSSVCIASIVAGPISGGSMNPARTLG 804
Query: 216 PAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGS 261
PAI S Y+ + VYFV I GAV AW +N++R T+ P I+ S
Sbjct: 805 PAIATSYYKGLWVYFVGPITGAVLAAWSYNVIRDTEHPGFPISLSS 942
>TC223040 similar to UP|NI51_ARATH (Q9SV84) Probable aquaporin NIP5.1
(NOD26-like intrinsic protein 5.1) (Nodulin-26-like
major intrinsic protein 6) (AtNLM6) (NLM6 protein)
(NodLikeMip6), partial (56%)
Length = 869
Score = 155 bits (393), Expect = 1e-38
Identities = 79/158 (50%), Positives = 111/158 (70%), Gaps = 2/158 (1%)
Frame = +1
Query: 97 NPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG--TIPSGTNL 154
NP++T AFA + FPW V YIA+Q+ ++ A LK ++ H SG T+P+ +
Sbjct: 10 NPSLTIAFAAFRHFPWTHVPAYIAAQVSASICACYALKGVY---HPFLSGGVTVPTVSVA 180
Query: 155 QAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTL 214
QAF EFI TF+L+FV++AVATD RA+GE+AGIA+G+T+LLNILISGP +G SMNP RTL
Sbjct: 181 QAFATEFIITFILLFVVTAVATDTRAVGELAGIAVGATVLLNILISGPTSGGSMNPVRTL 360
Query: 215 GPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDK 252
GPA+ Y+ I +Y V+ GA+AGA V+ +++ D+
Sbjct: 361 GPAVAAGNYKHIWIYLVAPTLGALAGAGVYTLVKLRDE 474
>AW099763 similar to PIR|JQ2285|JQ22 nodulin-26 - soybean, partial (34%)
Length = 472
Score = 142 bits (358), Expect = 2e-34
Identities = 78/165 (47%), Positives = 108/165 (65%), Gaps = 11/165 (6%)
Frame = +2
Query: 37 VQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHF 96
+ KLVAE VGT+FLIF GCAS+V N + D VVT PGI++VWGL +MVL+YS GHISGAHF
Sbjct: 8 LHKLVAEVVGTYFLIFAGCASVVENLDKDKVVTQPGISIVWGLTVMVLVYSFGHISGAHF 187
Query: 97 NPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASG-----------ILKMLFSGTHDQFS 145
PAVT A ++ P + ++ G L +L + G +D F+
Sbjct: 188 YPAVTIAHSS---------PP*VFTE-TGTSLCDNSGSSEPH*LVELLDLYSIGKNDSFA 337
Query: 146 GTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIG 190
GT+PS +++Q+FV+E+I T LM+VI+ VATD+ +IGE+A +A G
Sbjct: 338 GTLPSCSDVQSFVVEYIITIYLMYVIAGVATDHISIGELAWLAAG 472
>TC224715 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%)
Length = 1270
Score = 107 bits (266), Expect = 7e-24
Identities = 69/214 (32%), Positives = 108/214 (50%), Gaps = 7/214 (3%)
Frame = +1
Query: 41 VAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLPG----IALVWGLVLMVLIYSVGHISGAH 95
+AEF+ TF +F G S I NK DN P ++ L V + +ISG H
Sbjct: 571 LAEFISTFIFVFAGSGSGIAYNKLTDNGAATPAGLISASIAHAFALFVAVSVGANISGGH 750
Query: 96 FNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQ 155
NPAVTF +++ Y+ +QLLG+++AS +L + + T F + G
Sbjct: 751 VNPAVTFGAFVGGNITFLRGIVYVIAQLLGSIVASLLLAFVTASTVPAFGLSAGVGVG-N 927
Query: 156 AFVIEFITTFLLMFVISAVATDNRA--IGEMAGIAIGSTLLLNILISGPITGASMNPART 213
A V+E + TF L++ + A A D + +G +A IAIG + NIL+ G +GA+MNPA T
Sbjct: 928 ALVLEIVMTFGLVYTVYATAIDPKKGNLGIIAPIAIGFIVGANILLGGAFSGAAMNPAVT 1107
Query: 214 LGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNIL 247
GPA+ + +Y+ + G V+ ++
Sbjct: 1108FGPAVVSWTWTNHWIYWAGPLIGGGIAGLVYEVV 1209
Score = 37.0 bits (84), Expect = 0.009
Identities = 29/121 (23%), Positives = 55/121 (44%), Gaps = 11/121 (9%)
Frame = +1
Query: 154 LQAFVIEFITTFLLMFVISAVA------TDNRAIGE----MAGIAIGSTLLLNILISGPI 203
L+A + EFI+TF+ +F S TDN A A IA L + + + I
Sbjct: 559 LKAALAEFISTFIFVFAGSGSGIAYNKLTDNGAATPAGLISASIAHAFALFVAVSVGANI 738
Query: 204 TGASMNPARTLGPAI-FHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSS 262
+G +NPA T G + + + +VY ++ + G++ + + + + P ++ G
Sbjct: 739 SGGHVNPAVTFGAFVGGNITFLRGIVYVIAQLLGSIVASLLLAFVTASTVPAFGLSAGVG 918
Query: 263 I 263
+
Sbjct: 919 V 921
>TC225322 similar to UP|TI41_ARATH (O82316) Probable aquaporin TIP4.1
(Tonoplast intrinsic protein 4.1) (Epsilon-tonoplast
intrinsic protein) (Epsilon-TIP), partial (98%)
Length = 1378
Score = 105 bits (262), Expect = 2e-23
Identities = 80/242 (33%), Positives = 118/242 (48%), Gaps = 8/242 (3%)
Frame = +2
Query: 11 HDVVLSVNKDASKTIESSDTY--TSVFFVQKLVAEFVGTFFLIFTGC-ASIVVNK-NNDN 66
H ++L +++ A I T T +Q L+ EF+ TF +F G +S+VV+K D
Sbjct: 512 H*LILQLSRGAMAKIALGSTREATQPDCIQALIVEFIATFLFVFVGVGSSMVVDKLGGDA 691
Query: 67 VVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGA 126
+V L +A+ LV+ V+I S HISG H NPAVT + Y QL+ A
Sbjct: 692 LVGLFAVAVAHALVVAVMI-SAAHISGGHLNPAVTLGLLAGGHITIFRSLLYWIDQLVAA 868
Query: 127 VLASGILKMLFSGTHDQFSGTIPSGTNL-QAFVIEFITTFLLMFVISAVATDNRAIGEMA 185
AS +L L G T+ SG Q V E + TF L+F + A D + G +A
Sbjct: 869 AAASYLLYYLSGGQATPVH-TLASGVGYGQGVVWEIVLTFSLLFTVYATMVDPKK-GALA 1042
Query: 186 GIA---IGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAW 242
G+ +G + NIL G + ASMNPAR+ GPA+ + VY+V + G +
Sbjct: 1043GLGPTLVGFVVGANILAGGAYSAASMNPARSFGPALVTGNWTDHWVYWVGPLIGGGLAGF 1222
Query: 243 VF 244
++
Sbjct: 1223IY 1228
>TC204519 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic protein,
partial (97%)
Length = 989
Score = 104 bits (260), Expect = 4e-23
Identities = 69/212 (32%), Positives = 108/212 (50%), Gaps = 8/212 (3%)
Frame = +1
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLP-----GIALVWGLVLMVLIYS 87
S+ ++ +AEF+ T +F G S + + L +A+ G L V +
Sbjct: 100 SLASIKAYIAEFISTLLFVFAGVGSAIAYAKLTSDAALDPTGLVAVAICHGFALFVAVSV 279
Query: 88 VGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGT 147
+ISG H NPAVTF A + Y +QLLG+++AS +LK F +D +
Sbjct: 280 GANISGGHVNPAVTFGLALGGHITILTGLFYWIAQLLGSIVASLLLK--FVTGYDTPIHS 453
Query: 148 IPSGTNL-QAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISGPIT 204
+ +G + V E I TF L++ + A A D + ++G +A IAIG + NIL +GP +
Sbjct: 454 VAAGIGAGEGVVTEIIITFGLVYTVYATAADPKKGSLGTIAPIAIGFIVGANILAAGPFS 633
Query: 205 GASMNPARTLGPAIFHSKYRAIVVYFVSTIFG 236
G SMNPAR+ GPA+ + +Y+V + G
Sbjct: 634 GGSMNPARSFGPAVVSGDFHDNWIYWVGPLIG 729
>TC204517 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic protein,
complete
Length = 1016
Score = 103 bits (256), Expect = 1e-22
Identities = 72/231 (31%), Positives = 110/231 (47%), Gaps = 8/231 (3%)
Frame = +3
Query: 29 DTYTSVFFVQKLVAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLPG----IALVWGLVLMV 83
D S+ ++ +AEF T +F G S I K + P +A+ G L V
Sbjct: 96 DDSFSLTSIKAYIAEFHSTLLFVFAGVGSAIAYGKLTSDAALDPAGLLAVAICHGFALFV 275
Query: 84 LIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQ 143
+ +ISG H NPAVTF A + Y +QLLG+++A +L + G
Sbjct: 276 AVSVGANISGGHVNPAVTFGLALGGHITILTGFFYWIAQLLGSIVACFLLNYVTGGLPTP 455
Query: 144 FSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISG 201
++ V E I TF L++ + A A D + ++G +A IAIG + NIL +G
Sbjct: 456 IHSVASGVGAVEGVVTEIIITFGLVYTVYATAADPKKGSLGTIAPIAIGFIVGANILAAG 635
Query: 202 PITGASMNPARTLGPAIFHSKYRAIVVYFVSTIF-GAVAGAWVFNILRYTD 251
P +G SMNPAR+ GPA+ + +Y+V + G +AG N+ +D
Sbjct: 636 PFSGGSMNPARSFGPAVVSGDFHDNWIYWVGPLIGGGLAGLIYGNVFIRSD 788
>TC224822 similar to UP|TIP2_TOBAC (P24422) Probable aquaporin TIP-type
RB7-18C (Tonoplast intrinsic protein, root-specific
RB7-18C) (TobRB7) (RT-TIP), partial (96%)
Length = 1202
Score = 102 bits (255), Expect = 1e-22
Identities = 72/221 (32%), Positives = 111/221 (49%), Gaps = 8/221 (3%)
Frame = +1
Query: 24 TIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLP----GIALVWG 78
T+ + D V ++ +AEF T +F G S I N+ + P +A+
Sbjct: 112 TLGTFDDSFGVASLKAYLAEFHATLIFVFAGVGSAIAYNELTKDAALDPTGLVAVAVAHA 291
Query: 79 LVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFS 138
L V + +ISG H NPAVTF A I Y +QLLG+++A +L + +
Sbjct: 292 FALFVGVSVAANISGGHLNPAVTFGLAIGGNITLITGFLYWIAQLLGSIVACLLLNFITA 471
Query: 139 GTHDQFSGTIPSGTN-LQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLL 195
+ + +G N QA V E + TF L++ + A A D + ++G +A IAIG +
Sbjct: 472 KSIPSHAPA--TGVNDFQAVVFEIVITFGLVYTVYATAADPKKGSLGIIAPIAIGFVVGA 645
Query: 196 NILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFG 236
NIL +GP +G SMNPAR+ GPA+ + A +Y+V + G
Sbjct: 646 NILAAGPFSGGSMNPARSFGPAVVSGDFAANWIYWVGPLIG 768
>TC224823 similar to UP|TIP1_TOBAC (P21653) Probable aquaporin TIP-type
RB7-5A (Tonoplast intrinsic protein, root-specific
RB7-5A) (TobRB7) (RT-TIP), partial (96%)
Length = 1083
Score = 102 bits (254), Expect = 2e-22
Identities = 71/203 (34%), Positives = 104/203 (50%), Gaps = 8/203 (3%)
Frame = +2
Query: 42 AEFVGTFFLIFTGCAS-IVVNKNNDNVVTLP----GIALVWGLVLMVLIYSVGHISGAHF 96
AEF T +F G S I N+ + P +A+ L V + +ISG H
Sbjct: 101 AEFHATLIFVFAGVGSAIAYNELTKDAALDPTGLVAVAVAHAFALFVGVSVAANISGGHL 280
Query: 97 NPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTN-LQ 155
NPAVTF A I Y +QLLG+++A +L ++ + + S +G N LQ
Sbjct: 281 NPAVTFGLAIGGNITLITGFLYWIAQLLGSIVACLLLNLITAKSIPSHSPA--NGVNDLQ 454
Query: 156 AFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISGPITGASMNPART 213
A V E + TF L++ + A A D + ++G +A IAIG + NIL +GP +G SMNPAR+
Sbjct: 455 AVVFEIVITFGLVYTVYATAVDPKKGSLGIIAPIAIGFVVGANILAAGPFSGGSMNPARS 634
Query: 214 LGPAIFHSKYRAIVVYFVSTIFG 236
GPA+ A +Y+V + G
Sbjct: 635 FGPAVVSGDLAANWIYWVGPLIG 703
>TC214748 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%)
Length = 1254
Score = 101 bits (251), Expect = 4e-22
Identities = 66/214 (30%), Positives = 105/214 (48%), Gaps = 7/214 (3%)
Frame = +3
Query: 41 VAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLPG----IALVWGLVLMVLIYSVGHISGAH 95
+AEF+ T +F G S I NK DN P ++ L V + +ISG H
Sbjct: 198 LAEFISTLIFVFAGSGSGIAYNKLTDNGAATPAGLISASIAHAFALFVAVSVGANISGGH 377
Query: 96 FNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQ 155
NPAVTF ++ Y+ +QLLG+++AS +L + + F + G
Sbjct: 378 VNPAVTFGAFVGGNITLLRGIVYVIAQLLGSIVASLLLAFVTASPVPAFGLSAGVGVG-N 554
Query: 156 AFVIEFITTFLLMFVISAVATDNRA--IGEMAGIAIGSTLLLNILISGPITGASMNPART 213
A V+E + TF L++ + A A D + +G +A IAIG + NIL+ G +GA+MNPA T
Sbjct: 555 ALVLEIVMTFGLVYTVYATAVDPKKGNLGIIAPIAIGFIVGANILLGGAFSGAAMNPAVT 734
Query: 214 LGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNIL 247
GPA+ + +Y+ + G ++ ++
Sbjct: 735 FGPAVVSWTWTNHWIYWAGPLIGGGIAGLIYEVV 836
>TC216096 homologue to UP|PI27_ARATH (P93004) Aquaporin PIP2.7 (Plasma
membrane intrinsic protein 3) (Salt-stress induced major
intrinsis protein), partial (97%)
Length = 1043
Score = 100 bits (250), Expect = 5e-22
Identities = 65/221 (29%), Positives = 113/221 (50%), Gaps = 15/221 (6%)
Frame = +1
Query: 36 FVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV---VTLPGIALVWGLVLMVLIYSVGHIS 92
F + L+AEF+ T ++ A+++ +K V L GIA +G ++ VL+Y IS
Sbjct: 88 FYRALIAEFIATLLFLYVTVATVIGHKKQTGPCDGVGLLGIAWAFGGMIFVLVYCTAGIS 267
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG---TIP 149
G H NPAVTF ++ I+ Y+ +Q LGA+ G++K +++ G ++
Sbjct: 268 GGHINPAVTFGLFLARKVSLIRALFYMVAQCLGAICGVGLVKAFMKHSYNSLGGGANSVS 447
Query: 150 SGTNL-QAFVIEFITTFLLMFVISAVATDNRA-----IGEMAGIAIGSTLLLNILISGPI 203
+G N A E I TF+L++ + + R+ I +A + IG + + L + PI
Sbjct: 448 AGYNKGSALGAEIIGTFVLVYTVFSATDPKRSARDSHIPVLAPLPIGFAVFMVHLATIPI 627
Query: 204 TGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGA 241
TG +NPAR+ G A+ ++ + +++V GA+A A
Sbjct: 628 TGTGINPARSFGAAVIYNNGKVWDDHWIFWVGPFVGALAAA 750
>TC216098 similar to UP|PI27_ARATH (P93004) Aquaporin PIP2.7 (Plasma membrane
intrinsic protein 3) (Salt-stress induced major
intrinsis protein), partial (81%)
Length = 838
Score = 100 bits (249), Expect = 7e-22
Identities = 60/193 (31%), Positives = 104/193 (53%), Gaps = 7/193 (3%)
Frame = +2
Query: 36 FVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV---VTLPGIALVWGLVLMVLIYSVGHIS 92
F + L+AEF+ + ++ A+I+ +K V L GIA +G ++ VL+Y IS
Sbjct: 248 FYRALIAEFIASLLFLYVTVATIIGHKKQTGPCDGVGLLGIAWSFGGMIFVLVYCTAGIS 427
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG---TIP 149
G H NPAVTF ++ I+ Y+ +Q LGA+ G++K +++ G ++
Sbjct: 428 GGHINPAVTFGLFLARKVSLIRAVFYMVAQCLGAICGVGLVKAFMKHSYNSLGGGANSVS 607
Query: 150 SGTNL-QAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASM 208
+G N A E I TF+L++ + + R++ +A + IG + + L + PITG +
Sbjct: 608 AGYNKGSALGAEIIGTFVLVYTVFSATDPKRSV--LAPLPIGFAVFMVHLATIPITGTGI 781
Query: 209 NPARTLGPAIFHS 221
NPAR+LG A+ ++
Sbjct: 782 NPARSLGAAVIYN 820
>TC225036 similar to UP|O65357 (O65357) Aquaporin 2, partial (92%)
Length = 1610
Score = 100 bits (249), Expect = 7e-22
Identities = 67/230 (29%), Positives = 114/230 (49%), Gaps = 23/230 (10%)
Frame = +3
Query: 36 FVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLP-------GIALVWGLVLMVLIYSV 88
F + L+AEFV T ++ +++ + P GIA +G ++ VL+Y
Sbjct: 585 FFRALIAEFVATLLFLYVTILTVIGYNHQTATAAEPCSGVGVLGIAWAFGGMIFVLVYCT 764
Query: 89 GHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTI 148
ISG H NPAVTF ++ + Y+ +Q+LGA+ G++K L ++++ G +
Sbjct: 765 AGISGGHINPAVTFGLFLARKVSLTRAVGYMVAQVLGAISGVGLVKALQKSYYNRYKGGV 944
Query: 149 -------PSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGE------MAGIAIGSTLLL 195
GT L A E I TF+L++ + + ATD + + +A + IG + +
Sbjct: 945 NMLADGYSKGTGLGA---EIIGTFILVYTVFS-ATDPKRVARDSHVPVLAPLPIGFAVFM 1112
Query: 196 NILISGPITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGAW 242
L + PITG +NPAR+LGPA+ + +A +++V GA A+
Sbjct: 1113VHLATIPITGTGINPARSLGPAVIFNNEKAWDDQWIFWVGPFIGAALAAF 1262
>TC225044 homologue to UP|O22339 (O22339) Aquaporin-like transmembrane
channel protein, complete
Length = 1235
Score = 97.8 bits (242), Expect = 4e-21
Identities = 63/204 (30%), Positives = 106/204 (51%), Gaps = 13/204 (6%)
Frame = +2
Query: 32 TSVFFVQKLVAEFVGTFFLIFTGCASIV-VNKNNDNVVT--LPGIALVWGLVLMVLIYSV 88
TS F + +AEFV TF ++ +++ VN+++ T + GIA +G ++ L+Y
Sbjct: 176 TSWSFYRAGIAEFVATFLFLYITILTVMGVNRSSSKCATVGIQGIAWAFGGMIFALVYCT 355
Query: 89 GHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILK----MLFSGTHDQF 144
ISG H NPAVTF ++ + Y+ Q+LGA++ +G++K F G H+
Sbjct: 356 AGISGGHINPAVTFGLFLARKLSLTRALFYMVMQVLGAIVGAGVVKGFEGKTFYGQHNGG 535
Query: 145 SGTI-PSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGE-----MAGIAIGSTLLLNIL 198
+ + P T E + TF+L++ + + R+ + +A + IG + L L
Sbjct: 536 ANFVAPGYTKGDGLGAEIVGTFILVYTVFSATDAKRSARDSHVPILAPLPIGFAVFLVHL 715
Query: 199 ISGPITGASMNPARTLGPAIFHSK 222
+ PITG +NPAR+LG AI +K
Sbjct: 716 ATIPITGTGINPARSLGAAIIFNK 787
>TC214480 homologue to UP|Q39822 (Q39822) Pip1 protein, complete
Length = 1240
Score = 96.7 bits (239), Expect = 1e-20
Identities = 64/239 (26%), Positives = 117/239 (48%), Gaps = 24/239 (10%)
Frame = +1
Query: 28 SDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV---------VTLPGIALVWG 78
++ T F + L+AEF+ T ++ +++ K+ +V V + GIA +G
Sbjct: 178 AEELTQWSFYRALIAEFIATMLFLYITVLTVIGYKSQSDVKAGGDVCGGVGILGIAWAFG 357
Query: 79 LVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKML-- 136
++ +L+Y ISG H NPAVTF ++ I+ Y+ +Q LGA+ G++K
Sbjct: 358 GMIFILVYCTAGISGGHINPAVTFGLFLARKVSLIRAIMYMVAQCLGAICGVGLVKAFQK 537
Query: 137 -----FSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNR-----AIGEMAG 186
+ G ++ S +G L A E I TF+L++ + + R + +A
Sbjct: 538 AYYNRYGGGANELSEGYSTGVGLGA---EIIGTFVLVYTVFSATDPKRNARDSHVPVLAP 708
Query: 187 IAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGAW 242
+ IG + + L + P+TG +NPAR+LG A+ +++ +A +++V GA A+
Sbjct: 709 LPIGFAVFMVHLATIPVTGTGINPARSLGAAVMYNQQKAWDDHWIFWVGPFIGAAIAAF 885
>TC216974 homologue to UP|Q39822 (Q39822) Pip1 protein, complete
Length = 1254
Score = 96.7 bits (239), Expect = 1e-20
Identities = 64/239 (26%), Positives = 117/239 (48%), Gaps = 24/239 (10%)
Frame = +3
Query: 28 SDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV---------VTLPGIALVWG 78
++ T F + L+AEF+ T ++ +++ K+ +V V + GIA +G
Sbjct: 174 AEELTQWSFYRALIAEFIATMLFLYITVLTVIGYKSQSDVKAGGDVCGGVGILGIAWAFG 353
Query: 79 LVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKML-- 136
++ +L+Y ISG H NPAVTF ++ I+ Y+ +Q LGA+ G++K
Sbjct: 354 GMIFILVYCTAGISGGHINPAVTFGLFLARKVSLIRAIMYMVAQCLGAICGVGLVKAFQK 533
Query: 137 -----FSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNR-----AIGEMAG 186
+ G ++ S +G L A E I TF+L++ + + R + +A
Sbjct: 534 AYYNRYGGGANELSEGYSTGVGLGA---EIIGTFVLVYTVFSATDPKRNARDSHVPVLAP 704
Query: 187 IAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGAW 242
+ IG + + L + P+TG +NPAR+LG A+ +++ +A +++V GA A+
Sbjct: 705 LPIGFAVFMVHLATIPVTGTGINPARSLGAAVMYNQQKAWDDHWIFWVGPFIGAAIAAF 881
>TC223037 similar to UP|Q8H1Z5 (Q8H1Z5) Tonoplast intrinsic protein
bobTIP26-2, partial (76%)
Length = 723
Score = 95.9 bits (237), Expect = 2e-20
Identities = 66/196 (33%), Positives = 101/196 (50%), Gaps = 7/196 (3%)
Frame = +3
Query: 41 VAEFVGTFFLIFTGC-ASIVVNKNN-DNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNP 98
++EF+ T +F G +S+ VNK D L A+ L V + +ISG H NP
Sbjct: 138 LSEFISTLIFVFAGSGSSVAVNKLTVDKPSALVVAAVAHAFALFVAVSVSTNISGGHVNP 317
Query: 99 AVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNL-QAF 157
AVTF ++ + +Q+LG+V+A +LK + +G D + SG + A
Sbjct: 318 AVTFGAFVGGNLTLLRCVLFWIAQILGSVIACLLLKFI-TGGQDVPVFKLSSGVGVGNAV 494
Query: 158 VIEFITTFLLMFVISAVATDNRA----IGEMAGIAIGSTLLLNILISGPITGASMNPART 213
V+E + TF L++ + A D R+ +G MA I IG + N+L+ GP GASMNPA +
Sbjct: 495 VLEMVMTFGLVYTVYATTVDPRSRRGSLGVMAPIVIGFIVGANVLVGGPFDGASMNPAAS 674
Query: 214 LGPAIFHSKYRAIVVY 229
GPA+ ++ VY
Sbjct: 675 FGPAVVGWSWKNHWVY 722
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.138 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,884,643
Number of Sequences: 63676
Number of extensions: 118178
Number of successful extensions: 922
Number of sequences better than 10.0: 132
Number of HSP's better than 10.0 without gapping: 863
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 887
length of query: 265
length of database: 12,639,632
effective HSP length: 96
effective length of query: 169
effective length of database: 6,526,736
effective search space: 1103018384
effective search space used: 1103018384
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0162a.10


