

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0159.10
(353 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
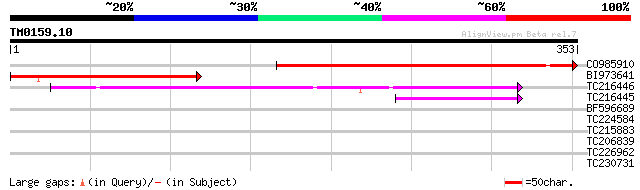
Score E
Sequences producing significant alignments: (bits) Value
CO985910 337 4e-93
BI973641 similar to GP|9758839|dbj UDP-galactose transporter rel... 198 3e-51
TC216446 weakly similar to UP|Q8NBK6 (Q8NBK6) Homo sapiens cDNA ... 129 2e-30
TC216445 weakly similar to UP|P78383 (P78383) UDP-galactose tran... 55 5e-08
BF596689 33 0.25
TC224584 similar to PIR|T49026|T49026 ubiquinol-cytochrome-c red... 33 0.25
TC215883 similar to PDB|1SE9_A.0|46015773|1SE9_A Chain A, Struct... 28 4.8
TC206839 similar to UP|Q8LG35 (Q8LG35) EF-hand Ca2+-binding prot... 28 6.3
TC226962 weakly similar to UP|Q9Y616 (Q9Y616) IL-1 receptor-asso... 28 6.3
TC230731 28 8.2
>CO985910
Length = 811
Score = 337 bits (865), Expect = 4e-93
Identities = 161/187 (86%), Positives = 176/187 (94%)
Frame = -1
Query: 167 AGEDISPYSRGRENTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCI 226
AG DISPY RGRENTVWGVLLM GYLGCDGFTSTFQDK+F+GY+MEIHNQIFYTTLCSCI
Sbjct: 811 AGTDISPYGRGRENTVWGVLLMLGYLGCDGFTSTFQDKMFKGYNMEIHNQIFYTTLCSCI 632
Query: 227 LSLTGLILQGHLIPAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMT 286
LSL GLI+QGHL+PA+EFVY H DCFFDIALLSTVATASQFFISYTIRTFGALTFATIMT
Sbjct: 631 LSLAGLIIQGHLLPAVEFVYIHKDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMT 452
Query: 287 TRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTSSDSIPLVQSGDS 346
TRQLVSI+LSCVWF+HPLSWEQWIGAVIVFG++YAKSF RKAP+KTTS S+ VQ+G+S
Sbjct: 451 TRQLVSILLSCVWFAHPLSWEQWIGAVIVFGAIYAKSFLRKAPEKTTS--SVEHVQNGNS 278
Query: 347 NNLKDNP 353
NNLK+NP
Sbjct: 277 NNLKENP 257
>BI973641 similar to GP|9758839|dbj UDP-galactose transporter related
protein-like {Arabidopsis thaliana}, partial (29%)
Length = 444
Score = 198 bits (504), Expect = 3e-51
Identities = 101/121 (83%), Positives = 109/121 (89%), Gaps = 2/121 (1%)
Frame = +3
Query: 1 MAESPSTSSSSSPPDS--RDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFK 58
MAE+ S S SS DS R+NKLWKG FAV+GIM+TL TYGVLQEKIMRVPYG K+YFK
Sbjct: 81 MAETTSASVSSPAGDSNWRENKLWKGTFAVAGIMLTLFTYGVLQEKIMRVPYGVNKDYFK 260
Query: 59 HSLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSF 118
+SLFLVFCNRITTSAVSAG+LLASKKALDPVAPIYKYCLVSVSNILTT+CQYEALKYVSF
Sbjct: 261 YSLFLVFCNRITTSAVSAGALLASKKALDPVAPIYKYCLVSVSNILTTSCQYEALKYVSF 440
Query: 119 P 119
P
Sbjct: 441 P 443
>TC216446 weakly similar to UP|Q8NBK6 (Q8NBK6) Homo sapiens cDNA PSEC0149
fis, clone PLACE1007338, partial (12%)
Length = 1397
Score = 129 bits (324), Expect = 2e-30
Identities = 95/300 (31%), Positives = 139/300 (45%), Gaps = 6/300 (2%)
Frame = +2
Query: 26 FAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHSLFLVFCNRITTSAVSAGSLLASKKA 85
F V+GI + GVLQE + + ++ F+H FL + S +
Sbjct: 116 FCVAGIWSAYIYQGVLQENVSTKRFNGER--FEHLAFLNLAQNVVCLIWSFIMIKMWASG 289
Query: 86 LDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTIIMQKRYQ 145
AP + Y ++N + EALKY+S+P Q LAK +KMIPVM+ GT++ RY
Sbjct: 290 NSGGAPWWSYWSAGITNTIGPAMGIEALKYISYPAQVLAKSSKMIPVMLMGTLVYGIRYT 469
Query: 146 GPDYMLAFLVTLGCSVF-ILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDGFTSTFQDK 204
P+Y+ FLV G S F +L + + IS + +G+ + L DGFT+ QD
Sbjct: 470 FPEYLCTFLVAGGVSTFALLKTSSKTISKLAHPNAPLGYGLCFLN--LAFDGFTNATQDS 643
Query: 205 LFRGYDMEIHNQI-----FYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIALLS 259
L Y I + T+ + I G A+ F +H + +DI L
Sbjct: 644 LKARYPKTSAWDIMLGMNLWGTIYNMIYMFGWPRASG--FEAVRFCQQHPEAAWDIFLYC 817
Query: 260 TVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSL 319
Q FI TI FG+L TI TTR+ VSI++S + +PLS +QW +VF L
Sbjct: 818 CCGAVGQNFIFLTISRFGSLANTTITTTRKFVSIVVSSLLSGNPLSTKQWGCVFMVFSGL 997
>TC216445 weakly similar to UP|P78383 (P78383) UDP-galactose transporter
related isozyme 1, partial (9%)
Length = 700
Score = 55.1 bits (131), Expect = 5e-08
Identities = 31/79 (39%), Positives = 42/79 (52%)
Frame = +2
Query: 241 AIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWF 300
A+ F H + +DI L Q FI TI FG+L TI TTR+ VSI++S +
Sbjct: 8 AVRFCQHHPEAAWDIFLYCCCGAVGQNFIFLTISRFGSLANTTITTTRKFVSIVVSSLLS 187
Query: 301 SHPLSWEQWIGAVIVFGSL 319
+PLS +QW +VF L
Sbjct: 188 GNPLSTKQWGCVSMVFSGL 244
>BF596689
Length = 292
Score = 32.7 bits (73), Expect = 0.25
Identities = 16/27 (59%), Positives = 18/27 (66%)
Frame = +2
Query: 1 MAESPSTSSSSSPPDSRDNKLWKGIFA 27
MAES + S+ S SRDNK WKG FA
Sbjct: 212 MAESCTNSAVSVDSISRDNKFWKGAFA 292
>TC224584 similar to PIR|T49026|T49026 ubiquinol-cytochrome-c reductase-like
protein - Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (86%)
Length = 595
Score = 32.7 bits (73), Expect = 0.25
Identities = 17/34 (50%), Positives = 21/34 (61%)
Frame = +1
Query: 2 AESPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTL 35
A +PSTS SSSP S ++ LW F SG +TL
Sbjct: 244 AATPSTSPSSSPAPSSESGLWITEFTRSGSTITL 345
>TC215883 similar to PDB|1SE9_A.0|46015773|1SE9_A Chain A, Structure Of
At3g01050, A Ubiquitin-Fold Protein From Arabidopsis
Thaliana. {Arabidopsis thaliana;} , partial (66%)
Length = 1110
Score = 28.5 bits (62), Expect = 4.8
Identities = 36/120 (30%), Positives = 52/120 (43%), Gaps = 7/120 (5%)
Frame = -1
Query: 68 RITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVS----FPVQTL 123
R TSA+ +G + L VA + SV+ TTTC + VS + T+
Sbjct: 690 RQMTSAIQSGCYKITHMHLFCVALFAAFFSFSVAGCCTTTCIVVTVSGVSHNGLWHSPTV 511
Query: 124 AKCAKMIPVMIWGTIIMQKRYQGPDYMLAF---LVTLGCSVFILYPAGEDISPYSRGREN 180
+K+ P +I T M GP + L+F TL +V IL AG+ + P S N
Sbjct: 510 LLFSKIFPALINFTSFM---VFGP-FSLSFGH*ARTLSFNVAILVAAGKLLGPISEPSVN 343
>TC206839 similar to UP|Q8LG35 (Q8LG35) EF-hand Ca2+-binding protein CCD1,
partial (74%)
Length = 552
Score = 28.1 bits (61), Expect = 6.3
Identities = 14/41 (34%), Positives = 23/41 (55%), Gaps = 4/41 (9%)
Frame = +2
Query: 267 FFISYTIRTFGALTFATIMTTRQLVSIMLSC----VWFSHP 303
F++SYT+ T AL+F TR+ + +L+C W+ P
Sbjct: 29 FYLSYTLTTLFALSFLHYGCTRKQLPRLLACDGQQAWWGWP 151
>TC226962 weakly similar to UP|Q9Y616 (Q9Y616) IL-1
receptor-associated-kinase-M (Interleukin-1
receptor-associated kinase 3), partial (8%)
Length = 1084
Score = 28.1 bits (61), Expect = 6.3
Identities = 24/99 (24%), Positives = 39/99 (39%), Gaps = 10/99 (10%)
Frame = -3
Query: 55 EYFKHSLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALK 114
E+ L C+R+ ++V G LAS ++++C SV+ + C Y
Sbjct: 617 EFLPAVLGASICSRVAKTSVIGGLALAS---------LFRHCRASVAAVKAPFCGYWPSN 465
Query: 115 YVSFPVQTLAKCAK----------MIPVMIWGTIIMQKR 143
VS + +L K IPVM+ T + R
Sbjct: 464 LVSIILNSLRLSLKYGFAQSTRFCSIPVMVLSTARLPDR 348
>TC230731
Length = 1006
Score = 27.7 bits (60), Expect = 8.2
Identities = 11/23 (47%), Positives = 14/23 (60%)
Frame = +3
Query: 2 AESPSTSSSSSPPDSRDNKLWKG 24
AE PST++ PPD+ K W G
Sbjct: 105 AEPPSTTADVPPPDAAAEKRWPG 173
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.137 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,911,068
Number of Sequences: 63676
Number of extensions: 266149
Number of successful extensions: 2155
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 2121
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2149
length of query: 353
length of database: 12,639,632
effective HSP length: 98
effective length of query: 255
effective length of database: 6,399,384
effective search space: 1631842920
effective search space used: 1631842920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0159.10


