

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
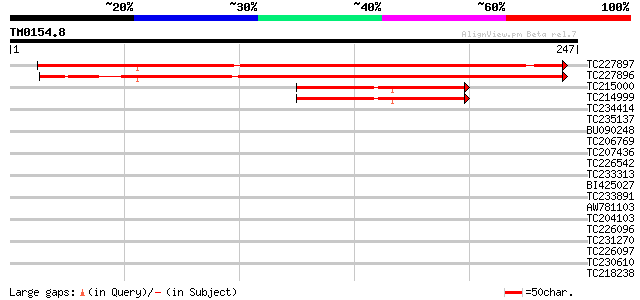
Query= TM0154.8
(247 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC227897 similar to GB|AAO63329.1|28950811|BT005265 At5g21920 {A... 327 2e-90
TC227896 similar to GB|AAO63329.1|28950811|BT005265 At5g21920 {A... 321 2e-88
TC215000 homologue to GB|AAP13371.1|30023676|BT006263 At3g07430 ... 78 3e-15
TC214999 78 3e-15
TC234414 similar to UP|Q6NN02 (Q6NN02) At4g14450, partial (24%) 35 0.041
TC235137 weakly similar to UP|Q7PV56 (Q7PV56) ENSANGP00000014031... 33 0.16
BU090248 similar to PIR|T03739|T03 nitrilase (EC 3.5.5.1) 4B - c... 31 0.59
TC206769 similar to UP|Q93ZG9 (Q93ZG9) AT4g25340/T30C3_20, parti... 30 1.0
TC207436 similar to GB|AAQ22666.1|33589800|BT010197 At3g63530 {A... 30 1.3
TC226542 weakly similar to UP|Q9ZWQ5 (Q9ZWQ5) UDP-glycose:flavon... 29 1.7
TC233313 29 1.7
BI425027 29 2.3
TC233891 similar to UP|Q94JR1 (Q94JR1) AT3g10030/T22K18_15, part... 28 3.0
AW781103 28 3.0
TC204103 similar to UP|Q75QC2 (Q75QC2) Glutamate-rich protein, p... 28 3.9
TC226096 similar to UP|ERF3_ARATH (O80339) Ethylene responsive e... 28 3.9
TC231270 28 3.9
TC226097 similar to UP|ERF3_ARATH (O80339) Ethylene responsive e... 28 3.9
TC230610 similar to UP|Q8RWR3 (Q8RWR3) At1g61040/T7P1_17, partia... 28 5.0
TC218238 weakly similar to UP|Q8H0F2 (Q8H0F2) Anthocyanin 3'-glu... 27 6.6
>TC227897 similar to GB|AAO63329.1|28950811|BT005265 At5g21920 {Arabidopsis
thaliana;} , partial (40%)
Length = 1008
Score = 327 bits (839), Expect = 2e-90
Identities = 170/233 (72%), Positives = 191/233 (81%), Gaps = 2/233 (0%)
Frame = +2
Query: 13 MATETDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPE--LHKSFTTATDNCFRFL 70
++TE D K S+CW N Q PFALP L P SN T+ LS P LH SFTTA DN FRF+
Sbjct: 113 LSTEIDCTKKVSNCWGNGQIPFALPFLTLPNSNFTTLLSSPPQLLHTSFTTAADNFFRFV 292
Query: 71 HSLASQNPFLNKILSLRSEFDSLCFQIRCSNYRSRRWLSSHNFAAVLPGDSVAGLVVGNG 130
HSLASQNPFLNK+LSL +EF +LC QIR R+ +SSHNFAAVLPG SVAGLVV NG
Sbjct: 293 HSLASQNPFLNKVLSLPAEFHTLCVQIR--KQRNVGLVSSHNFAAVLPGGSVAGLVVANG 466
Query: 131 IQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPIL 190
+ NFLNIYNTLL+VRLVLTWFPN+PP+IV PLSTICDPYLN+FRGLIPPLGGTLDLSPIL
Sbjct: 467 VLNFLNIYNTLLIVRLVLTWFPNTPPSIVSPLSTICDPYLNIFRGLIPPLGGTLDLSPIL 646
Query: 191 AFLVLNAFTSASAALPAELPVTQQSEQGVAATLQSTDITSSQKKWMKRLQGNR 243
AFLVLNAFTS +AALPAELPVT+QS+QG+AA L S T+SQKKWM+R QGNR
Sbjct: 647 AFLVLNAFTSTAAALPAELPVTEQSKQGLAAPLPS---TTSQKKWMRRFQGNR 796
>TC227896 similar to GB|AAO63329.1|28950811|BT005265 At5g21920 {Arabidopsis
thaliana;} , partial (44%)
Length = 1078
Score = 321 bits (822), Expect = 2e-88
Identities = 167/231 (72%), Positives = 188/231 (81%), Gaps = 1/231 (0%)
Frame = +2
Query: 14 ATETDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPE-LHKSFTTATDNCFRFLHS 72
+TE D KK+S+ CW N Q PFALP L SPP+ LH SFTTA DN FRFLHS
Sbjct: 101 STEIDTKKLSN-CWGNAQMPFALPTL---------LSSPPQHLHTSFTTAADNFFRFLHS 250
Query: 73 LASQNPFLNKILSLRSEFDSLCFQIRCSNYRSRRWLSSHNFAAVLPGDSVAGLVVGNGIQ 132
LAS N FLNKILSL SEF +LC QI R+ R +S HNFAAVLPGDSVAGLVV NG+
Sbjct: 251 LASHNSFLNKILSLPSEFHTLCVQI--GKQRNVRLVSGHNFAAVLPGDSVAGLVVANGVL 424
Query: 133 NFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
NFLNIYNTLL+VRLVLTWFPN+PP+IV PLSTICDPYLN+FRGLIPPLGGTLDLSPILAF
Sbjct: 425 NFLNIYNTLLIVRLVLTWFPNTPPSIVSPLSTICDPYLNIFRGLIPPLGGTLDLSPILAF 604
Query: 193 LVLNAFTSASAALPAELPVTQQSEQGVAATLQSTDITSSQKKWMKRLQGNR 243
LVLNAFTS ++ALPAELPVT+QS+QG+AA L ST++T+SQKKW+ R QGNR
Sbjct: 605 LVLNAFTSTASALPAELPVTEQSKQGLAAPLHSTNLTTSQKKWLTRFQGNR 757
>TC215000 homologue to GB|AAP13371.1|30023676|BT006263 At3g07430 {Arabidopsis
thaliana;} , partial (47%)
Length = 742
Score = 78.2 bits (191), Expect = 3e-15
Identities = 42/78 (53%), Positives = 55/78 (69%), Gaps = 3/78 (3%)
Frame = +2
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTI---CDPYLNLFRGLIPPLGG 182
VV G+ +L+IY+ +L+VR++L+WFPN P PLS I CDPYLNLFR +IPP+
Sbjct: 392 VVAAGLGKWLDIYSGVLMVRVLLSWFPNIPWERQ-PLSAIRDLCDPYLNLFRNIIPPVFD 568
Query: 183 TLDLSPILAFLVLNAFTS 200
TLD+SP+LAF VL S
Sbjct: 569 TLDVSPLLAFAVLGTLGS 622
>TC214999
Length = 1057
Score = 78.2 bits (191), Expect = 3e-15
Identities = 42/78 (53%), Positives = 55/78 (69%), Gaps = 3/78 (3%)
Frame = +1
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTI---CDPYLNLFRGLIPPLGG 182
VV G+ +L+IY+ +L+VR++L+WFPN P PLS I CDPYLNLFR +IPP+
Sbjct: 418 VVAAGLGKWLDIYSGVLMVRVLLSWFPNIPWERQ-PLSAIRDLCDPYLNLFRNIIPPVFD 594
Query: 183 TLDLSPILAFLVLNAFTS 200
TLD+SP+LAF VL S
Sbjct: 595 TLDVSPLLAFAVLGTLGS 648
>TC234414 similar to UP|Q6NN02 (Q6NN02) At4g14450, partial (24%)
Length = 424
Score = 34.7 bits (78), Expect = 0.041
Identities = 19/44 (43%), Positives = 23/44 (52%), Gaps = 2/44 (4%)
Frame = +2
Query: 19 PKKVSSHCWRNTQTPFAL--PILKPPISNLTSFLSPPELHKSFT 60
PKKV W++ PF P L PP N+ F SP +H SFT
Sbjct: 242 PKKVVFKKWQHPAAPFFYEQPSLVPPFVNV*GFQSPTLVHSSFT 373
>TC235137 weakly similar to UP|Q7PV56 (Q7PV56) ENSANGP00000014031 (Fragment),
partial (6%)
Length = 577
Score = 32.7 bits (73), Expect = 0.16
Identities = 21/62 (33%), Positives = 27/62 (42%)
Frame = -1
Query: 4 VELLPQLKAMATETDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFTTAT 63
+EL PQL+ + D + V H P L +L PP+ L PP LH AT
Sbjct: 313 IELQPQLQDVLRRLDEEPVRVHSGAARNQP--LQLLYPPLHTLLPPPPPPPLHLCHVLAT 140
Query: 64 DN 65
N
Sbjct: 139 HN 134
>BU090248 similar to PIR|T03739|T03 nitrilase (EC 3.5.5.1) 4B - common
tobacco, partial (29%)
Length = 415
Score = 30.8 bits (68), Expect = 0.59
Identities = 14/28 (50%), Positives = 18/28 (64%), Gaps = 2/28 (7%)
Frame = +3
Query: 27 WRNTQTP--FALPILKPPISNLTSFLSP 52
W T TP F LP+ KPP S+ T F++P
Sbjct: 105 WVPTSTPQRFELPLFKPPPSSTTHFIAP 188
>TC206769 similar to UP|Q93ZG9 (Q93ZG9) AT4g25340/T30C3_20, partial (10%)
Length = 1057
Score = 30.0 bits (66), Expect = 1.0
Identities = 26/66 (39%), Positives = 33/66 (49%), Gaps = 8/66 (12%)
Frame = -1
Query: 22 VSSHCWRNTQTPF-ALPILKPPISNL-TSFLSP------PELHKSFTTATDNCFRFLHSL 73
VS+ + T TPF A+ I KP S+L T +SP PEL SF+ CF F + L
Sbjct: 481 VSASAF*ETDTPFSAVGIGKPSSSSLSTRSVSPLATIGSPELLLSFSFN*ATCFFFFNFL 302
Query: 74 ASQNPF 79
PF
Sbjct: 301 VGSYPF 284
>TC207436 similar to GB|AAQ22666.1|33589800|BT010197 At3g63530 {Arabidopsis
thaliana;} , partial (39%)
Length = 1398
Score = 29.6 bits (65), Expect = 1.3
Identities = 14/47 (29%), Positives = 23/47 (48%)
Frame = +1
Query: 13 MATETDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSF 59
++T D K V +H WR+ P +PI++ +SP + SF
Sbjct: 328 LSTRKDSK*VGTHTWRSITIPLVIPIIQQEALLNILKVSPMNMSTSF 468
>TC226542 weakly similar to UP|Q9ZWQ5 (Q9ZWQ5) UDP-glycose:flavonoid
glycosyltransferase, partial (18%)
Length = 753
Score = 29.3 bits (64), Expect = 1.7
Identities = 18/55 (32%), Positives = 24/55 (42%), Gaps = 13/55 (23%)
Frame = -1
Query: 33 PFALPILKPPISNLTSFLSPPELHKS-------------FTTATDNCFRFLHSLA 74
P + + +S+L FL PP S FTT+ N F+FLHS A
Sbjct: 402 PTFIALFAATLSSLALFLIPPSSSSSLINEAIAFPISSRFTTSLPNSFQFLHSFA 238
>TC233313
Length = 681
Score = 29.3 bits (64), Expect = 1.7
Identities = 12/40 (30%), Positives = 20/40 (50%)
Frame = +2
Query: 34 FALPILKPPISNLTSFLSPPELHKSFTTATDNCFRFLHSL 73
F + + N T+ +PP+ + T+ CF+F HSL
Sbjct: 260 FTFGVFLSSLRNFTTLEAPPQCNSLLTSFHSLCFKFSHSL 379
>BI425027
Length = 400
Score = 28.9 bits (63), Expect = 2.3
Identities = 18/63 (28%), Positives = 33/63 (51%)
Frame = +2
Query: 74 ASQNPFLNKILSLRSEFDSLCFQIRCSNYRSRRWLSSHNFAAVLPGDSVAGLVVGNGIQN 133
A+QNPF++ + L+ E+ C+Q+R S S ++ L S LV+G + +
Sbjct: 26 AAQNPFISLLRPLKGEYPRECWQMRFSLVTDAFSDMSRMSSSFL---SQGELVIGQAVTS 196
Query: 134 FLN 136
+L+
Sbjct: 197 YLS 205
>TC233891 similar to UP|Q94JR1 (Q94JR1) AT3g10030/T22K18_15, partial (23%)
Length = 819
Score = 28.5 bits (62), Expect = 3.0
Identities = 17/50 (34%), Positives = 24/50 (48%)
Frame = +2
Query: 99 CSNYRSRRWLSSHNFAAVLPGDSVAGLVVGNGIQNFLNIYNTLLVVRLVL 148
CSN RRW+ + LP S+ +G+G + L L +V LVL
Sbjct: 83 CSNLL*RRWVFRLVYKLQLPCRSLLNHTIGSGPSDILRKEGLLYLVALVL 232
>AW781103
Length = 431
Score = 28.5 bits (62), Expect = 3.0
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = -2
Query: 8 PQLKAMATETDPKKVSSHCW 27
PQL A + DP +V SHCW
Sbjct: 220 PQLAARMHKGDPCRVQSHCW 161
>TC204103 similar to UP|Q75QC2 (Q75QC2) Glutamate-rich protein, partial (38%)
Length = 845
Score = 28.1 bits (61), Expect = 3.9
Identities = 24/63 (38%), Positives = 31/63 (49%), Gaps = 1/63 (1%)
Frame = -3
Query: 145 RLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDL-SPILAFLVLNAFTSASA 203
+LVL WFP + PL+++ P L L R L PL L L S +L L+ A S
Sbjct: 492 KLVLFWFPQRFQLVCSPLASV--PPLVLIRFL*FPLSLLLLLFSQVLGSLLPQASLPLSL 319
Query: 204 ALP 206
LP
Sbjct: 318 LLP 310
>TC226096 similar to UP|ERF3_ARATH (O80339) Ethylene responsive element
binding factor 3 (AtERF3), partial (28%)
Length = 672
Score = 28.1 bits (61), Expect = 3.9
Identities = 15/33 (45%), Positives = 19/33 (57%)
Frame = +1
Query: 35 ALPILKPPISNLTSFLSPPELHKSFTTATDNCF 67
ALP+L P + +T+ L PP S TTAT F
Sbjct: 298 ALPLLSPKTATVTATLPPP----SLTTATTTSF 384
>TC231270
Length = 764
Score = 28.1 bits (61), Expect = 3.9
Identities = 30/123 (24%), Positives = 51/123 (41%), Gaps = 2/123 (1%)
Frame = -1
Query: 23 SSHCWRNTQTPFALPILKPP--ISNLTSFLSPPELHKSFTTATDNCFRFLHSLASQNPFL 80
SSH + T T LP++ P I + F+S F+ + RF S F
Sbjct: 440 SSHFVQVTTTTIVLPLIPTPRIIPPILPFIS----FSMFSRSKIFFCRFGRSKMFFYMFG 273
Query: 81 NKILSLRSEFDSLCFQIRCSNYRSRRWLSSHNFAAVLPGDSVAGLVVGNGIQNFLNIYNT 140
++ L + +C QI C SR W+ V+P + L+V ++ N +N
Sbjct: 272 RSMILLPRFYSIICLQICCPALDSRIWIWCFGSIIVVPSGTRKLLLVQE--ESATNHHNL 99
Query: 141 LLV 143
+++
Sbjct: 98 VII 90
>TC226097 similar to UP|ERF3_ARATH (O80339) Ethylene responsive element
binding factor 3 (AtERF3), partial (36%)
Length = 466
Score = 28.1 bits (61), Expect = 3.9
Identities = 15/33 (45%), Positives = 19/33 (57%)
Frame = +3
Query: 35 ALPILKPPISNLTSFLSPPELHKSFTTATDNCF 67
ALP+L P + +T+ L PP S TTAT F
Sbjct: 327 ALPLLSPKTATVTATLPPP----SLTTATTTSF 413
>TC230610 similar to UP|Q8RWR3 (Q8RWR3) At1g61040/T7P1_17, partial (19%)
Length = 782
Score = 27.7 bits (60), Expect = 5.0
Identities = 13/30 (43%), Positives = 16/30 (53%)
Frame = +1
Query: 104 SRRWLSSHNFAAVLPGDSVAGLVVGNGIQN 133
SRRW S N+ A PG+ A GN + N
Sbjct: 61 SRRWTRSRNYYAAKPGEKAA---AGNNVVN 141
>TC218238 weakly similar to UP|Q8H0F2 (Q8H0F2) Anthocyanin
3'-glucosyltransferase, partial (26%)
Length = 936
Score = 27.3 bits (59), Expect = 6.6
Identities = 14/34 (41%), Positives = 19/34 (55%)
Frame = -3
Query: 41 PPISNLTSFLSPPELHKSFTTATDNCFRFLHSLA 74
PPI +F + FTT++ N F+FLHS A
Sbjct: 502 PPIKTPIAF----PISSLFTTSSPNSFQFLHSFA 413
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,147,839
Number of Sequences: 63676
Number of extensions: 216193
Number of successful extensions: 1366
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 1355
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1359
length of query: 247
length of database: 12,639,632
effective HSP length: 95
effective length of query: 152
effective length of database: 6,590,412
effective search space: 1001742624
effective search space used: 1001742624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0154.8


