

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
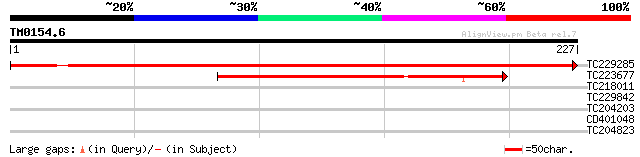
Query= TM0154.6
(227 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC229285 365 e-102
TC223677 129 1e-30
TC218011 30 0.90
TC229842 28 4.4
TC204203 homologue to UP|Q9SWS9 (Q9SWS9) Ribosomal protein S26, ... 27 5.8
CD401048 similar to GP|28416701|gb| At4g02100 {Arabidopsis thali... 27 9.9
TC204823 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, part... 27 9.9
>TC229285
Length = 1202
Score = 365 bits (938), Expect = e-102
Identities = 176/227 (77%), Positives = 199/227 (87%)
Frame = +3
Query: 1 MAKLYLVVILSFLHSASTGSAATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLA 60
M +L+V LSF+++A+T NTQIKVTNNPADKLV AINENRTA+KVS L DN GLA
Sbjct: 339 MTMWFLLVTLSFVYAAATA----NTQIKVTNNPADKLVVAINENRTAHKVSALTDNPGLA 506
Query: 61 CIALQYIKAYQGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYV 120
CIALQYIKAYQG+C AVGG D KKPPES+FAEVFAPNCGV+AS+LAPITGRFL CQTKYV
Sbjct: 507 CIALQYIKAYQGECDAVGGSDGKKPPESRFAEVFAPNCGVEASTLAPITGRFLACQTKYV 686
Query: 121 HAPEAFSEVLIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFEG 180
HAPEAFS++LI+NQ+SL+IL++ NHTQVGAAVTGTDGGSPYFWCVLFS GKPN TF FE
Sbjct: 687 HAPEAFSDILIRNQQSLDILYNNNHTQVGAAVTGTDGGSPYFWCVLFSSGKPNKTFTFES 866
Query: 181 GVAKLTKPGCFSGANDECSGAHDWSPLSVMWLFAASVLIALGFAFPL 227
GVAK+TKPGCFSGANDECSGA WSPL+ MW+FA S+LIA+GFA L
Sbjct: 867 GVAKITKPGCFSGANDECSGASYWSPLNEMWVFATSLLIAMGFALTL 1007
>TC223677
Length = 478
Score = 129 bits (324), Expect = 1e-30
Identities = 62/117 (52%), Positives = 78/117 (65%), Gaps = 1/117 (0%)
Frame = +3
Query: 84 KPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEAFSEVLIQNQRSLEILHSK 143
KPPE F EVFAPNCGV+ + ITG +GCQ KY+ AFSEVLI++++SL +L +K
Sbjct: 33 KPPEDDFTEVFAPNCGVELPTFGTITGHIVGCQRKYLEPSLAFSEVLIKDEKSLSLLKNK 212
Query: 144 NHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFEG-GVAKLTKPGCFSGANDECS 199
+HT+VG + G G P+FWCVLFS GK NSTF E G K GC+SG+ CS
Sbjct: 213 SHTEVGVGLVGLHKG-PFFWCVLFSNGKTNSTFVLENHGAGIQQKKGCYSGSTTPCS 380
>TC218011
Length = 1116
Score = 30.0 bits (66), Expect = 0.90
Identities = 23/73 (31%), Positives = 29/73 (39%), Gaps = 7/73 (9%)
Frame = -1
Query: 153 TGTDGGSPYFWCVLFSGGKPNSTFAFEGGVAKLTK-------PGCFSGANDECSGAHDWS 205
T DG SP F +TF F GG + P C S N +G+ +
Sbjct: 648 TSGDGDSPEL--AFFEECDELATFKFRGGKLRSDSIC*VPRAPVCESRGNGGSNGSELEA 475
Query: 206 PLSVMWLFAASVL 218
PLS WL S+L
Sbjct: 474 PLSTWWLLCKSLL 436
>TC229842
Length = 994
Score = 27.7 bits (60), Expect = 4.4
Identities = 18/57 (31%), Positives = 27/57 (46%), Gaps = 3/57 (5%)
Frame = +2
Query: 152 VTGTDGGSPYFWCVLF---SGGKPNSTFAFEGGVAKLTKPGCFSGANDECSGAHDWS 205
VTG++G P+ LF SGG A + + ++ G S ECS A+ W+
Sbjct: 365 VTGSNGYDPWSQGALFPSLSGGLKAQESATQNEMIYASQSGSASNHVHECSKANSWT 535
>TC204203 homologue to UP|Q9SWS9 (Q9SWS9) Ribosomal protein S26, partial
(27%)
Length = 449
Score = 27.3 bits (59), Expect = 5.8
Identities = 17/50 (34%), Positives = 25/50 (50%)
Frame = -3
Query: 159 SPYFWCVLFSGGKPNSTFAFEGGVAKLTKPGCFSGANDECSGAHDWSPLS 208
+P W L S G+ +S + G+ +LT+P C A DE SP+S
Sbjct: 159 APKTWNFLLSSGR-SSYSSRTWGLTRLTRPWCIIPAPDESLWRLTLSPIS 13
>CD401048 similar to GP|28416701|gb| At4g02100 {Arabidopsis thaliana},
partial (14%)
Length = 642
Score = 26.6 bits (57), Expect = 9.9
Identities = 12/28 (42%), Positives = 18/28 (63%)
Frame = +3
Query: 115 CQTKYVHAPEAFSEVLIQNQRSLEILHS 142
CQT+++ F +L+Q+QR E LHS
Sbjct: 87 CQTRFLSPSIYFLFLLLQSQRERERLHS 170
>TC204823 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (66%)
Length = 1596
Score = 26.6 bits (57), Expect = 9.9
Identities = 19/56 (33%), Positives = 22/56 (38%), Gaps = 1/56 (1%)
Frame = -1
Query: 162 FWCVLFSGGKPNSTFAFEGGVAKLTKPGCFS-GANDECSGAHDWSPLSVMWLFAAS 216
FW G P +FA GG AKL+ P G+ E P S W A S
Sbjct: 462 FWLSTS*GKFPTKSFASSGGSAKLSSPASSPLGSQLERPAWVSLRPCSSSWKVAVS 295
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,503,158
Number of Sequences: 63676
Number of extensions: 146581
Number of successful extensions: 594
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 592
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 593
length of query: 227
length of database: 12,639,632
effective HSP length: 94
effective length of query: 133
effective length of database: 6,654,088
effective search space: 884993704
effective search space used: 884993704
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0154.6


