

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.14
(906 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
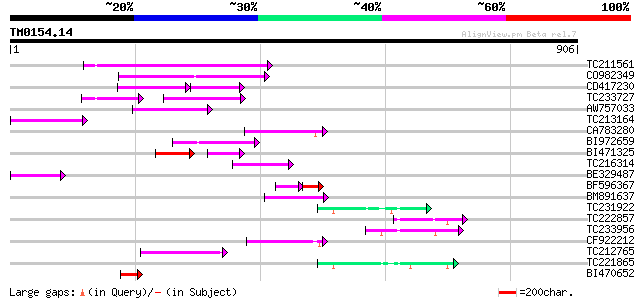
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 164 1e-40
CO982349 130 3e-30
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 89 7e-26
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 69 1e-22
AW757033 85 2e-16
TC213164 77 3e-14
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 73 6e-13
BI972659 71 2e-12
BI471325 52 1e-11
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 64 4e-10
BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase... 57 4e-08
BF596367 39 4e-08
BM891637 54 4e-07
TC231922 52 9e-07
TC222857 51 2e-06
TC233956 51 3e-06
CF922212 50 3e-06
TC212765 50 4e-06
TC221865 50 6e-06
BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase ... 47 5e-05
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 164 bits (416), Expect = 1e-40
Identities = 96/303 (31%), Positives = 149/303 (48%)
Frame = +1
Query: 118 NALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMT 177
+AL+A E K+ C + K+D KAYD V W FL ++ +M F W+ I
Sbjct: 7 SALIANEVIDEAKRSNKSC---LVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIEE 177
Query: 178 CVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIK 237
CV + S+++NG+P F P RGLRQGDPL+P+LF + E + L+ +AV +
Sbjct: 178 CVKSASISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRRAVEENLYKPYL 357
Query: 238 IARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVP 297
+ IS L +ADD++ F A + +IK IL S+E S IN KS V
Sbjct: 358 VGANGVPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKSCFGVF-GVT 534
Query: 298 DDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAG 357
D E L + YLG+P + +++ + + +KL WK R + G
Sbjct: 535 DQWKQEAANYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKCERKLSKWK*RHISFGG 714
Query: 358 REVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAK 417
R L+K+V +IP Y S F +P + +++ + F WGG ++ + + W+ LC K
Sbjct: 715 RVTLIKSVLTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGGGPDQKKISWIRWEKLCLPK 894
Query: 418 DNG 420
+ G
Sbjct: 895 ERG 903
>CO982349
Length = 795
Score = 130 bits (326), Expect = 3e-30
Identities = 74/240 (30%), Positives = 118/240 (48%)
Frame = +3
Query: 175 IMTCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLH 234
I C+ + SI++N +P F P RGLRQGDPL+P LF + E + L+ +AV +
Sbjct: 72 IRGCLHSASVSILVNESPINEFSPQRGLRQGDPLAPLLFNIVAEGLTGLMREAVDRKRFN 251
Query: 235 GIKIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSR 294
+ + +S L +ADD++ F AT++ IK IL +E ASG IN +S+
Sbjct: 252 SFLVGKNKEPVSILQYADDTIFFEEATMENMKVIKTILRCFELASGLKINFAESRFGAIW 431
Query: 295 NVPDDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLY 354
PD E + L + YLG+P +I + + + KL WK+R +
Sbjct: 432 K-PDHWCKEAAEFLNCSMLSMPFSYLGIPIEANPRCREI*DPIIRKCETKLARWKQRHIS 608
Query: 355 RAGREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLC 414
GR L+ A+ A+P Y S F P + ++ + +F WGG +R + + W+ +C
Sbjct: 609 LGGRVTLINAILTALPIYFFSFFRAPSKVVQKLVNIQRKFLWGGGSXQRRIAWVKWETVC 788
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 89.0 bits (219), Expect(2) = 7e-26
Identities = 45/118 (38%), Positives = 68/118 (57%)
Frame = -3
Query: 172 VSLIMTCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLAS 231
V L+ C+ST S++ NG + F P+ G+RQ DP++PYLF+LC E S LIN +
Sbjct: 686 VQLVWYCMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFVLCLERLSHLINLVMNKK 507
Query: 232 SLHGIKIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSK 289
I++ R +ISHL F DD ++F A + + I L + +SG +N+DK+K
Sbjct: 506 L*RSIQVNRGGLLISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKTK 333
Score = 47.8 bits (112), Expect(2) = 7e-26
Identities = 27/86 (31%), Positives = 44/86 (50%)
Frame = -2
Query: 289 KLSCSRNVPDDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGW 348
K S S+NV ++ L +++ + KYLG+P + F+F+ ++V K+ W
Sbjct: 333 KFSFSKNVNWWVEEDISSGLGLQSTDDLGKYLGVPIFHKNPTSHTFDFIIDKVNKRFSRW 154
Query: 349 KERSLYRAGREVLVKAVAHAIPSYVM 374
K L GR L K V A+PS++M
Sbjct: 153 KSNLLSMVGRLTLTK*VV*ALPSHIM 76
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 69.3 bits (168), Expect(2) = 1e-22
Identities = 39/99 (39%), Positives = 58/99 (58%)
Frame = -3
Query: 115 ITDNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSL 174
I D V E + + KK+ G N +ALK+D+ KA+D ++ FL SVL + G+ ++ +
Sbjct: 834 IKDCTCVTSEVINMLDKKVFGGN--LALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNW 661
Query: 175 IMTCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLF 213
I + + SI +NG P F RG+RQGDPLSP L+
Sbjct: 660 IRVILLSAKLSISVNGEPVGFFSCQRGVRQGDPLSPSLY 544
Score = 56.2 bits (134), Expect(2) = 1e-22
Identities = 38/132 (28%), Positives = 61/132 (45%)
Frame = -2
Query: 246 SHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDCFHELR 305
SH++ ADD +IF + T + ++ Y+ + GQVI+ KSK+ ++P +
Sbjct: 451 SHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIFIG-SIPGQRLRVIS 275
Query: 306 QLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREVLVKAV 365
LL + Y G+P GK + V +++ KL WK L GR LV +V
Sbjct: 274 DLLQYFHSDIPFNYFGVPIFKGKPTRRHLYCVADKIKCKLASWKGSILSIMGRIQLVNSV 95
Query: 366 AHAIPSYVMSCF 377
H + Y S +
Sbjct: 94 IHGMLLYSFSVY 59
>AW757033
Length = 441
Score = 84.7 bits (208), Expect = 2e-16
Identities = 47/128 (36%), Positives = 69/128 (53%)
Frame = -2
Query: 196 FQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIARTAPVISHLLFADDSV 255
F PHRGLRQGDPL+P LF + E + L+ +AV + + + +S L +ADD++
Sbjct: 431 FSPHRGLRQGDPLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKKPVSILQYADDTI 252
Query: 256 IFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDCFHELRQLLTVKAVES 315
F AT++ IK IL +E ASG IN KS+ + +VPD E + + +
Sbjct: 251 FFGEATMENVRVIKTILRGFELASGLKINFAKSRFG-AISVPD*WRKEAAEFMNCSLLSL 75
Query: 316 YDKYLGLP 323
YLG+P
Sbjct: 74 PFSYLGIP 51
>TC213164
Length = 446
Score = 77.4 bits (189), Expect = 3e-14
Identities = 44/124 (35%), Positives = 65/124 (51%)
Frame = +3
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V ++ K+PG DGL F ++FW I+ D+ F + + N L L
Sbjct: 42 EVRATMWDRGNAKSPGPDGLNFKFIKEFWDILKTDLLRFLDEFYANGVFLKRSNAFFLAL 221
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPKV P N+++ ISL I+KI+ K L+ R K +LP +I E QT F+ GR N +
Sbjct: 222 IPKVHDP*SLNEYKPISLIGCIYKIVAKLLSGRHKKVLPTIIDERQTVFMEGRHTLHNVV 401
Query: 121 VAFE 124
+A E
Sbjct: 402 IANE 413
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 72.8 bits (177), Expect = 6e-13
Identities = 37/139 (26%), Positives = 65/139 (46%), Gaps = 5/139 (3%)
Frame = +2
Query: 375 SCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNGRLGFRDFKMFNSAL 434
S F LP + ++E + RF WGG+ R + ++WK +C K G LG +D + FN+ L
Sbjct: 5 SFFSLP*KIIAKLESLQRRFLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTTL 184
Query: 435 VAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSIMRSGEI-----FAT 489
+ K W + + KV ++ Y + G + S W ++++ ++
Sbjct: 185 LGKWRWDLFYIQQEPWAKVLQSKYGGWRALEEGSSGSKDSAWWKDLIKTQQLQRNIPLKR 364
Query: 490 GGMWNIGNGKSVKAWSDKW 508
+W +G G +K W D W
Sbjct: 365 ETIWKVGGGDRIKFWEDLW 421
>BI972659
Length = 453
Score = 70.9 bits (172), Expect = 2e-12
Identities = 39/140 (27%), Positives = 72/140 (50%)
Frame = +1
Query: 260 ATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDCFHELRQLLTVKAVESYDKY 319
A+ + +K++L +E SG IN KS+ P+ H+ QLL + ++ Y
Sbjct: 10 ASWDNIIVLKSMLRGFEMVSGLRINYAKSQFGIVGFQPNWA-HDAAQLLNCRQLDIPFHY 186
Query: 320 LGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREVLVKAVAHAIPSYVMSCFLL 379
LG+P + S ++ + + KL W ++ L GR L+K+V +A+P Y++S F +
Sbjct: 187 LGMPIAVKASSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFFKI 366
Query: 380 PDGLCNQIEGMISRFYWGGD 399
P + +++ + F WGG+
Sbjct: 367 PQRIVDKLVSLQRTFMWGGN 426
>BI471325
Length = 421
Score = 52.0 bits (123), Expect(2) = 1e-11
Identities = 26/63 (41%), Positives = 40/63 (63%)
Frame = +2
Query: 233 LHGIKIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSC 292
+H +K+ R ++SHLL D +F R +EA +K IL ++E +SG V+NL KS++
Sbjct: 8 IHRVKVHRGITILSHLLSVDFYFLFCRTIDKEADVLKNILTTFEASSGLVVNLRKSEICF 187
Query: 293 SRN 295
SRN
Sbjct: 188 SRN 196
Score = 36.6 bits (83), Expect(2) = 1e-11
Identities = 18/61 (29%), Positives = 36/61 (58%), Gaps = 2/61 (3%)
Frame = +1
Query: 317 DKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGRE--VLVKAVAHAIPSYVM 374
D+YL +P++I K IFN++++ + ++ ++ RE +++K VA +I +Y M
Sbjct: 238 DEYLDMPSMIRTKKKAIFNYLRDEFGRIYSTLVQQESFKGQREKCLIIKYVAQSITTYCM 417
Query: 375 S 375
S
Sbjct: 418 S 420
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 63.5 bits (153), Expect = 4e-10
Identities = 30/97 (30%), Positives = 51/97 (51%)
Frame = +2
Query: 357 GREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKA 416
GR L+K+V A+P +S F +P + +++ + +F WGG + + W D+C
Sbjct: 8 GRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADICNP 187
Query: 417 KDNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKV 453
K +G LG +D FN+AL + W + SN L ++
Sbjct: 188 KIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWARL 298
>BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (15%)
Length = 376
Score = 57.0 bits (136), Expect = 4e-08
Identities = 31/87 (35%), Positives = 45/87 (51%)
Frame = -2
Query: 2 VEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVLI 61
++ A++Q K+PG DGL F + FW + D+ F + I N + + LI
Sbjct: 375 IKSAVWQCGSVKSPGPDGLNFKFIKHFWERLKPDIIRFLNEFHANGIFPKGGNASFIALI 196
Query: 62 PKVKKPMHANQFRLISLCNVIFKIITK 88
PKVK P N FR ISL ++KI+ K
Sbjct: 195 PKVKHPQALNDFRPISLIGCVYKIVAK 115
>BF596367
Length = 419
Score = 38.9 bits (89), Expect(2) = 4e-08
Identities = 15/34 (44%), Positives = 24/34 (70%)
Frame = +3
Query: 468 KRGYRPSYAWSSIMRSGEIFATGGMWNIGNGKSV 501
K+ Y P+Y+WSS+ S ++ G +W IG+G+SV
Sbjct: 189 KKAYMPNYSWSSV**SKDLAEHGAIWRIGDGRSV 290
Score = 37.7 bits (86), Expect(2) = 4e-08
Identities = 17/45 (37%), Positives = 27/45 (59%)
Frame = +1
Query: 425 RDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKR 469
R K F+ A++AK WR+ N ++L+ V A YFP ++ AK+
Sbjct: 58 RRIKAFHKAMLAKQMWRLIDNKDALVT*VLTAKYFPNSSLLDAKK 192
>BM891637
Length = 427
Score = 53.5 bits (127), Expect = 4e-07
Identities = 31/108 (28%), Positives = 52/108 (47%), Gaps = 6/108 (5%)
Frame = -1
Query: 408 LSWKDLCKAKDNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFP-RGTIRG 466
+SW+ C + D G LG +D K+ N+AL+ K W + P L ++ + Y RG +G
Sbjct: 403 ISWRQCCASGDVGGLGIKDIKILNNALLIKWKWLMFHQPHQLWNRILISKYKGWRGLDQG 224
Query: 467 AKRGYRPSYAWS---SIMRSGEIFATGGM--WNIGNGKSVKAWSDKWV 509
++ Y + W+ +I + + A W +G G + W D WV
Sbjct: 223 PQKYYFSPW-WADLRAINQHQSMIAASNQFCWKVGRGDQILFWEDSWV 83
>TC231922
Length = 803
Score = 52.4 bits (124), Expect = 9e-07
Identities = 52/209 (24%), Positives = 75/209 (35%), Gaps = 27/209 (12%)
Frame = -3
Query: 492 MWNIGNGKSVKAWSDKWVPGTDTL------VYSVGLAQHFGVSLVSDLMLPGL---LAWN 542
+W +G G +K W D W+ L +Y++ + +S + W
Sbjct: 708 VWRVGCGDKIKFWQDSWLSEGCNLQQKYNQLYTISRQ*NLTISKMGKFSQNARSWEFKWR 529
Query: 543 RELINF--ICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVVA 600
R L ++ I I + HD ++W S G YSTKS Y+ L +
Sbjct: 528 RRLFDYEYAMAVDFMNEISGISIQNQAHDS-MFWKADSSGVYSTKSAYRLL-------MP 373
Query: 601 SSSQVPSL------------PRA---DWRKLWKADALLPVRAYLHARGL-VTDPTCPRCL 644
S S PS PRA WR LP R L R + + D CP C
Sbjct: 372 SISPAPSRRIFQILWHLKIPPRAAVFSWRLFLDR---LPTRGNLSRRSIPIQDIMCPLCG 202
Query: 645 VALETVQHALLFCPMVQPIWFASSLGFRL 673
E H C M + +W+ S R+
Sbjct: 201 CQHEEAGHLFFHCKMTKGLWWESMRWIRV 115
>TC222857
Length = 657
Score = 51.2 bits (121), Expect = 2e-06
Identities = 34/128 (26%), Positives = 54/128 (41%), Gaps = 9/128 (7%)
Frame = -3
Query: 613 WRKLWKADALLPVRAYLHARGL-VTDPTCPRCLVALETVQHALLFCPMVQPIWFASSLGF 671
WR LW LP +A L AR + ++D TCP C ET H + C QPIW+ +
Sbjct: 571 WRLLWDR---LPTKANLRARQVQISDLTCPFCRRVEETASHMFIHCIKTQPIWWETMNWI 401
Query: 672 RLAHECRVHEFVSDFMQVADMSSLGA--------FLTLLYAIWMARNELCFKEKASTLEQ 723
+ FMQ + + G ++ + ++IW RN + F +
Sbjct: 400 NMQGPL-PWSITDHFMQFSSLKEAGIRSRRWQWWWMAVTWSIWQLRNSIVFSNATFDGNK 224
Query: 724 ILTHAASL 731
++ A+ L
Sbjct: 223 LVEDASFL 200
>TC233956
Length = 757
Score = 50.8 bits (120), Expect = 3e-06
Identities = 43/172 (25%), Positives = 70/172 (40%), Gaps = 16/172 (9%)
Frame = +2
Query: 569 DDVLYWPFSSDGHYSTKSGYQFLR-----HGAEAVVASSSQVPSLPRA---DWRKLWKAD 620
+D W S G STKS YQ ++ G ++ P+A WR LW
Sbjct: 56 NDTWVWRAESTGIISTKSAYQVIKSELDDEGQHLGFKKLWEIKVPPKALSFVWRLLWDR- 232
Query: 621 ALLPVRAYLHARGL-VTDPTCPRCLVALETVQHALLFCPMVQPIW--FASSLGFRLAHEC 677
LP + L R + V D CP C ET H C + P+W F S + C
Sbjct: 233 --LPTKDNLIKRQIQVEDDLCPFCHSQSETASHLFFTCGKIMPLWWEFLSWVKEDKVFHC 406
Query: 678 R-----VHEFVSDFMQVADMSSLGAFLTLLYAIWMARNELCFKEKASTLEQI 724
R + + S +V++ ++ + +IW RN++ F+ + + ++
Sbjct: 407 RPMDNFLQHYSSAASKVSNTRRTMWWIAVTNSIWRLRNDIIFQNQTVDITRL 562
>CF922212
Length = 445
Score = 50.4 bits (119), Expect = 3e-06
Identities = 35/141 (24%), Positives = 61/141 (42%), Gaps = 11/141 (7%)
Frame = -2
Query: 379 LPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNGRLGFRDFKMFNSALVAKN 438
+P+ + +++ + F WGG + ++ + ++W +C K+ LG RD + FN AL+ K
Sbjct: 441 VPNKVLDKLVRLQQWFLWGGGLGQKKIAWVNWDSVCLPKEKEGLGNRDLRKFNYALLGKR 262
Query: 439 WWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAW---SSIMRSGEIFATGGMW-- 493
W + + L +V + Y + +R S+ W S I S E G W
Sbjct: 261 RWNLFHHQGELGARVLDSKYKRWRNLDEERRVKSESFWWQEISFITHSTE----DGSWFE 94
Query: 494 ------NIGNGKSVKAWSDKW 508
+G V+ W D W
Sbjct: 93 KRTKTGRLGCRAKVRFWEDGW 31
>TC212765
Length = 637
Score = 50.1 bits (118), Expect = 4e-06
Identities = 41/141 (29%), Positives = 62/141 (43%), Gaps = 3/141 (2%)
Frame = +3
Query: 210 PYLFILCGEVFSALINKAVLASSLHGIKIARTAPVISHLLFADDSVIFARATVQEALSIK 269
PYLF+LC E + I+ I ++ ++ +L+ DD +F A A
Sbjct: 15 PYLFVLCMERLALCIHDLTSQ*IWRPILVSDDNRLVPYLMLVDDIFLFINAIGD*AYLAS 194
Query: 270 AILASYEQASGQVINLDKSKLSCSRNVPDDCFHELRQLLTVKAVESYDKYLGLPTIIGKS 329
L ++ Q G +NLDKS + CS + + L + S KYL L I G+S
Sbjct: 195 LTL*NFFQTFGLKLNLDKSSMYCS*RMAPVLRDLISNTLNIVHAPSLGKYLCLKFIHGRS 374
Query: 330 KTQIFNFVKERV---WKKLKG 347
K F V ER+ W + +G
Sbjct: 375 KIVDFLLV-ERIHNPWPRGRG 434
>TC221865
Length = 1045
Score = 49.7 bits (117), Expect = 6e-06
Identities = 57/260 (21%), Positives = 90/260 (33%), Gaps = 34/260 (13%)
Frame = -3
Query: 492 MWNIGNGKSVKAWSDKWVPGTDTLV------YSVGLAQHFGVSLV---SDLMLPGLLAWN 542
+W +G G + W DK V +TL+ Y + Q + V ++ W
Sbjct: 932 VWKVGCGDKFRFWEDKLVGEGETLMRKYPRMYQISCQQQQLIQQVGSHTETTWEWKFQWR 753
Query: 543 RELINFICCPPTARAIMAIPLPLVPH-DDVLYWPFSSDGHYSTKSGYQFLRHGAEAVVAS 601
R L + P+ H D W +GHYST+S Y ++H EA A+
Sbjct: 752 RPLFDNEVDTTIGFLEDISRFPIHRHLTDCWVWKSEPNGHYSTRSAYHLMQH--EAAKAN 579
Query: 602 SSQVPSLPRADWRKLWKADALLPVRAYLHARGLVTD--PT---------------CPRCL 644
+ + ++WK +P +A + A LV D PT C C
Sbjct: 578 TDPA-------FEEIWKLK--IPAKAAIFA*RLVKDRLPTKNNLRRRQVQLNGTLCLFCR 426
Query: 645 VALETVQHALLFCPMVQPIWFASSLGFRLAHECRV---HEFVSDFMQVADMSSLGA---- 697
E H C QP+W+ S + H F+ + +
Sbjct: 425 NYEEEASHLFFSCTKTQPVWWESLSWINTSGAFSANPRHHFLQHSFGLLGLKQYNR*KCW 246
Query: 698 FLTLLYAIWMARNELCFKEK 717
++ L + IW RN F +
Sbjct: 245 WVALTFTIWQHRNRTIFSNE 186
>BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase (EC
2.7.7.49) (clone AmLi2) - garden snapdragon (fragment),
partial (22%)
Length = 111
Score = 46.6 bits (109), Expect = 5e-05
Identities = 18/34 (52%), Positives = 25/34 (72%)
Frame = +1
Query: 178 CVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPY 211
C+S+ SI++NG+P E F+P RGLRQG P P+
Sbjct: 7 CLSSASISILVNGSPTEEFKPXRGLRQGGPFGPF 108
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.139 0.444
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,503,201
Number of Sequences: 63676
Number of extensions: 787797
Number of successful extensions: 4902
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 4814
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4881
length of query: 906
length of database: 12,639,632
effective HSP length: 106
effective length of query: 800
effective length of database: 5,889,976
effective search space: 4711980800
effective search space used: 4711980800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0154.14


