

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0153b.6
(156 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
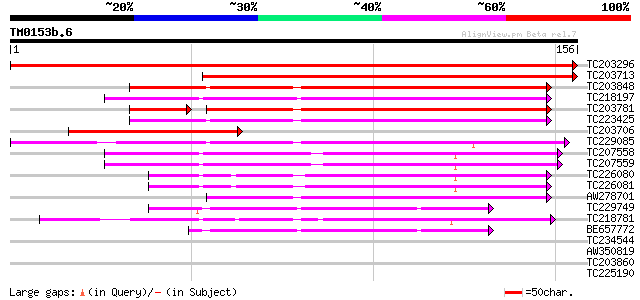
Score E
Sequences producing significant alignments: (bits) Value
TC203296 similar to GB|AAP21358.1|30102880|BT006550 At5g19590 {A... 275 6e-75
TC203713 similar to GB|AAP21358.1|30102880|BT006550 At5g19590 {A... 189 4e-49
TC203848 weakly similar to GB|AAC19282.1|3193298|T14P8 coded for... 94 3e-20
TC218197 weakly similar to GB|AAC19269.1|3193285|T14P8 coded for... 86 9e-18
TC203781 weakly similar to GB|AAC19282.1|3193298|T14P8 coded for... 76 5e-17
TC223425 weakly similar to GB|AAC19282.1|3193298|T14P8 coded for... 79 7e-16
TC203706 weakly similar to UP|Q6T520 (Q6T520) Zn-finger transcri... 79 9e-16
TC229085 73 5e-14
TC207558 similar to UP|Q9FFE5 (Q9FFE5) Gb|AAF02163.1, partial (43%) 69 7e-13
TC207559 similar to UP|Q9FFE5 (Q9FFE5) Gb|AAF02163.1, partial (44%) 63 5e-11
TC226080 similar to UP|Q7XA63 (Q7XA63) At5g19860, partial (62%) 62 1e-10
TC226081 similar to UP|Q7XA63 (Q7XA63) At5g19860, partial (62%) 60 6e-10
AW278701 55 1e-08
TC229749 similar to UP|Q70VA9 (Q70VA9) Cp protein, complete 48 2e-06
TC218781 weakly similar to UP|Q9FIV1 (Q9FIV1) Gb|AAF02153.1, par... 45 1e-05
BE657772 42 1e-04
TC234544 weakly similar to UP|Q9FIV1 (Q9FIV1) Gb|AAF02153.1, par... 35 0.011
AW350819 30 0.62
TC203860 similar to UP|Q8L5P7 (Q8L5P7) LHY protein, partial (48%) 30 0.62
TC225190 weakly similar to UP|Q8VZG3 (Q8VZG3) AT3g48690/T8P19_20... 28 1.8
>TC203296 similar to GB|AAP21358.1|30102880|BT006550 At5g19590 {Arabidopsis
thaliana;} , partial (64%)
Length = 813
Score = 275 bits (703), Expect = 6e-75
Identities = 132/156 (84%), Positives = 143/156 (91%)
Frame = +1
Query: 1 MNPKILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVN 60
MNPKIL +FAL LNL S SFQNPNPK PKST S AH+EL N+G P GLLP TTVLG+AVN
Sbjct: 55 MNPKILFVFALLLNLVSNSFQNPNPKTPKSTSSQAHVELINHGLPAGLLPATTVLGYAVN 234
Query: 61 QTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGI 120
+TSGDF+V+LGGACKITLPPDNYVATYS+TITGKIVKG+IAEL GIRVRAFF+WWSITGI
Sbjct: 235 RTSGDFTVKLGGACKITLPPDNYVATYSDTITGKIVKGKIAELEGIRVRAFFKWWSITGI 414
Query: 121 RSSGDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS 156
RSSGD+IVFEVGMVTAKYPSKNFDDSPACEGQ SSS
Sbjct: 415 RSSGDDIVFEVGMVTAKYPSKNFDDSPACEGQHSSS 522
>TC203713 similar to GB|AAP21358.1|30102880|BT006550 At5g19590 {Arabidopsis
thaliana;} , partial (64%)
Length = 645
Score = 189 bits (481), Expect = 4e-49
Identities = 90/103 (87%), Positives = 99/103 (95%)
Frame = +3
Query: 54 VLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQ 113
V G+AVN+TSG+F+V+L GACKITLPPDNYVATYS+TITGKIVKG+IAEL GIRVRAFF+
Sbjct: 18 VSGYAVNRTSGEFTVKLSGACKITLPPDNYVATYSDTITGKIVKGKIAELEGIRVRAFFK 197
Query: 114 WWSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS 156
WWSITGIRSSGD+IVFEVGMVTAKYPSKNFDDSPACEGQ SSS
Sbjct: 198 WWSITGIRSSGDDIVFEVGMVTAKYPSKNFDDSPACEGQHSSS 326
>TC203848 weakly similar to GB|AAC19282.1|3193298|T14P8 coded for by A.
thaliana cDNA R65118; coded for by A. thaliana cDNA
T43996 {Arabidopsis thaliana;} , partial (79%)
Length = 1128
Score = 93.6 bits (231), Expect = 3e-20
Identities = 47/116 (40%), Positives = 71/116 (60%)
Frame = +3
Query: 34 SAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITG 93
SA+ L YGFPVGLLP + G+++N+ SG+F+V GAC + ++Y Y +TITG
Sbjct: 462 SAYDVLMEYGFPVGLLPKGAI-GYSLNRDSGEFAVYFEGACSFDI--ESYALKYKSTITG 632
Query: 94 KIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
I KGR+ L G+ V+ W +I + G++I F VG+ +A + +NF +SP C
Sbjct: 633 VISKGRLYNLKGVTVKILLLWLNIVEVSRQGNDIYFSVGIASADFGVENFLESPQC 800
>TC218197 weakly similar to GB|AAC19269.1|3193285|T14P8 coded for by A.
thaliana cDNA N65001; coded for by A. thaliana cDNA
R64991 {Arabidopsis thaliana;} , partial (74%)
Length = 1021
Score = 85.5 bits (210), Expect = 9e-18
Identities = 45/123 (36%), Positives = 71/123 (57%)
Frame = +3
Query: 27 PPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVAT 86
P + +A+ L ++ FP G+LP V G+ ++ +SG FS L G+C +L +Y +
Sbjct: 45 PAITATPTAYEMLESFHFPAGILPKG-VTGYELDPSSGKFSADLNGSCSFSLE-GSYQLS 218
Query: 87 YSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGMVTAKYPSKNFDDS 146
Y TITG + +GR+ EL GI V+ F W +I + G+++ F VG+ +A +P NF S
Sbjct: 219 YQKTITGYVSEGRLTELRGISVKILFFWLNILDVVRVGEDLDFSVGVASASFPLDNFFVS 398
Query: 147 PAC 149
P C
Sbjct: 399 PQC 407
>TC203781 weakly similar to GB|AAC19282.1|3193298|T14P8 coded for by A.
thaliana cDNA R65118; coded for by A. thaliana cDNA
T43996 {Arabidopsis thaliana;} , partial (63%)
Length = 752
Score = 76.3 bits (186), Expect(2) = 5e-17
Identities = 35/95 (36%), Positives = 58/95 (60%)
Frame = +1
Query: 55 LGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQW 114
+G+++N+ SG+F+V GAC + ++Y Y +TITG I KGR+ L G+ V+ W
Sbjct: 145 IGYSLNRDSGEFAVYFQGACSFDI--ESYTLKYKSTITGVISKGRLYNLKGVTVKILLLW 318
Query: 115 WSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
+I + G++I F VG+ +A + +NF +SP C
Sbjct: 319 LNIVEVSRQGNDIYFSVGIASADFGVENFLESPQC 423
Score = 27.3 bits (59), Expect(2) = 5e-17
Identities = 12/17 (70%), Positives = 13/17 (75%)
Frame = +3
Query: 34 SAHIELTNYGFPVGLLP 50
SA+ L YGFPVGLLP
Sbjct: 84 SAYDVLMEYGFPVGLLP 134
>TC223425 weakly similar to GB|AAC19282.1|3193298|T14P8 coded for by A.
thaliana cDNA R65118; coded for by A. thaliana cDNA
T43996 {Arabidopsis thaliana;} , partial (70%)
Length = 597
Score = 79.3 bits (194), Expect = 7e-16
Identities = 41/116 (35%), Positives = 65/116 (55%)
Frame = +3
Query: 34 SAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITG 93
SA+ L Y PVGLLP G+ +N+ +G F+ L G C ++ ++Y Y +TI G
Sbjct: 249 SAYEVLEKYDLPVGLLPKGAT-GYELNEKNGHFTAYLNGTCYFSI--ESYELKYKSTIKG 419
Query: 94 KIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
I KG++++L GI V+ W I + GD++ F VG+ +A + +F +SP C
Sbjct: 420 VISKGKLSKLKGISVKIEVLWLKIVEVTRDGDDLQFSVGVASAGFSVDSFSESPQC 587
>TC203706 weakly similar to UP|Q6T520 (Q6T520) Zn-finger transcription factor
1, partial (5%)
Length = 586
Score = 79.0 bits (193), Expect = 9e-16
Identities = 36/48 (75%), Positives = 40/48 (83%)
Frame = +1
Query: 17 SKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSG 64
S SFQNPNPK PKST S AH+EL N+G P GLLP TTVLG+AVN+TSG
Sbjct: 442 SNSFQNPNPKTPKSTSSQAHVELINHGLPAGLLPATTVLGYAVNRTSG 585
>TC229085
Length = 732
Score = 73.2 bits (178), Expect = 5e-14
Identities = 46/156 (29%), Positives = 77/156 (48%), Gaps = 2/156 (1%)
Frame = +2
Query: 1 MNPKILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVN 60
M P+ LILF+ + L + P H L YGFP GLLP+ V + ++
Sbjct: 35 MFPQTLILFSALIILFHGDLVLST-----TAPDDIHDVLPQYGFPKGLLPNNAV-SYTLS 196
Query: 61 QTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGI 120
G F+V+L C + D+ + Y + I+G + G ++ ++GI+ + F W +TGI
Sbjct: 197 PDDGFFTVQLDAPCYVHW--DDQLVYYHSQISGTLTYGSVSHVSGIQAQKLFLWLPVTGI 370
Query: 121 RSSGDN--IVFEVGMVTAKYPSKNFDDSPACEGQRS 154
+ D+ + F VG ++ P+ +F D P C + S
Sbjct: 371 KVHQDSGMLEFFVGALSQTLPASDFQDVPGCSRRGS 478
>TC207558 similar to UP|Q9FFE5 (Q9FFE5) Gb|AAF02163.1, partial (43%)
Length = 923
Score = 69.3 bits (168), Expect = 7e-13
Identities = 41/129 (31%), Positives = 67/129 (51%), Gaps = 3/129 (2%)
Frame = +3
Query: 27 PPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVAT 86
P ++ ++ + EL G PVGLLP + +++N TSG+F V + AC + +
Sbjct: 141 PARANSTTIYEELRAQGLPVGLLPKG-IAKYSMNATSGEFEVWMKEACNAKFENEVH--- 308
Query: 87 YSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIR---SSGDNIVFEVGMVTAKYPSKNF 143
Y + I G + GRI EL+G+ + F W+ + GIR + I F+VG+ ++ F
Sbjct: 309 YDSNIKGVLGYGRIGELSGVSAQELFLWFPVKGIRVDVPTSGLIHFDVGVADKQFSLSLF 488
Query: 144 DDSPACEGQ 152
+D P C Q
Sbjct: 489 EDPPDCNPQ 515
>TC207559 similar to UP|Q9FFE5 (Q9FFE5) Gb|AAF02163.1, partial (44%)
Length = 860
Score = 63.2 bits (152), Expect = 5e-11
Identities = 38/129 (29%), Positives = 65/129 (49%), Gaps = 3/129 (2%)
Frame = +3
Query: 27 PPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVAT 86
P + ++ + EL G PVGLLP + +++N +SG+F V + C + +
Sbjct: 36 PATANSTTIYEELRAQGLPVGLLPKG-IAKYSLNASSGEFEVWMKEPCNAKFENEVH--- 203
Query: 87 YSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIR---SSGDNIVFEVGMVTAKYPSKNF 143
Y + I G + GRI +L+G+ + F W+ + GIR + I F+VG+ ++ F
Sbjct: 204 YDSNIKGVLGYGRIGKLSGVSAQELFLWFPVKGIRVDVPTSGLIHFDVGVADKQFSLSLF 383
Query: 144 DDSPACEGQ 152
+D P C Q
Sbjct: 384 EDPPDCNPQ 410
>TC226080 similar to UP|Q7XA63 (Q7XA63) At5g19860, partial (62%)
Length = 877
Score = 62.0 bits (149), Expect = 1e-10
Identities = 40/114 (35%), Positives = 59/114 (51%), Gaps = 3/114 (2%)
Frame = +1
Query: 39 LTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKG 98
L YG P GLLPDT V + +++ G F V L C I +Y+ Y + I+GK+ G
Sbjct: 181 LPKYGLPSGLLPDT-VTDYKLDE-DGQFVVVLPKPCYIQF---DYLVYYESKISGKLSYG 345
Query: 99 RIAELNGIRVRAFFQWWSITGIR---SSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
I L GI+V+ F W+++ IR D+I F+VG++ K F +C
Sbjct: 346 SITNLKGIQVQRLFIWFNVDEIRVDLPPSDSIYFQVGIINKKLDVDQFKTVRSC 507
>TC226081 similar to UP|Q7XA63 (Q7XA63) At5g19860, partial (62%)
Length = 919
Score = 59.7 bits (143), Expect = 6e-10
Identities = 38/114 (33%), Positives = 58/114 (50%), Gaps = 3/114 (2%)
Frame = +2
Query: 39 LTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKG 98
L YG P GLLPDT V + +++ G F V L C I +Y+ Y I+GK+ G
Sbjct: 227 LPKYGLPSGLLPDT-VTDYTLDE-DGQFVVVLAKPCYIQF---DYLVYYETKISGKLSYG 391
Query: 99 RIAELNGIRVRAFFQWWSITGIR---SSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
I L GI+V+ F W+++ I+ ++I F+VG++ K F +C
Sbjct: 392 SITNLKGIQVQRLFIWFNVDEIKVDLPPSNSIYFQVGIINKKLDVHQFKTVRSC 553
>AW278701
Length = 331
Score = 55.1 bits (131), Expect = 1e-08
Identities = 30/95 (31%), Positives = 49/95 (51%)
Frame = +2
Query: 55 LGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQW 114
LG++ + +G+F+ L GAC + D Y Y +TIT I KGR+ L + + W
Sbjct: 20 LGYSYTKDTGEFAR*LQGACSFDI--DTYTLKY*STITDVISKGRLYNLKCVTRKILLLW 193
Query: 115 WSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
+I + +G++I VG+ Y +N +SP C
Sbjct: 194 LNIVEVSRNGNDIYLSVGINCDDYSVQNLLESPQC 298
>TC229749 similar to UP|Q70VA9 (Q70VA9) Cp protein, complete
Length = 890
Score = 47.8 bits (112), Expect = 2e-06
Identities = 27/96 (28%), Positives = 49/96 (50%), Gaps = 1/96 (1%)
Frame = +1
Query: 39 LTNYGFPVGLLP-DTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVK 97
L Y P+G+ P D T + N+ +G V + C++ D+ V + T+TG + K
Sbjct: 316 LKEYDLPIGIFPRDAT--NYEFNEETGKLVVYIPQVCEVGYK-DSSVLRFFTTVTGYMEK 486
Query: 98 GRIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGM 133
G++A++ G++ + W ++T I S G + GM
Sbjct: 487 GKLADIEGMKTKVLI-WVNVTTILSEGSKLYVTAGM 591
>TC218781 weakly similar to UP|Q9FIV1 (Q9FIV1) Gb|AAF02153.1, partial (82%)
Length = 788
Score = 45.4 bits (106), Expect = 1e-05
Identities = 38/145 (26%), Positives = 64/145 (43%), Gaps = 3/145 (2%)
Frame = +1
Query: 9 FALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSV 68
F+L L+L S +P+P S H L + G P GLLP+ V ++ ++T G V
Sbjct: 91 FSLSLSLLQLSLSSPSP--------SIHDLLRSKGLPPGLLPEE-VKSYSFSET-GHLEV 240
Query: 69 RLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGI---RSSGD 125
L C +N V + +T + G + + G++ F W + I S
Sbjct: 241 FLDAPCLTKY--ENRVL-FEQVVTANLTYGSLIGVEGLQQEELFVWLPVKDIIVNDPSSG 411
Query: 126 NIVFEVGMVTAKYPSKNFDDSPACE 150
I+F++G+ + F+D P C+
Sbjct: 412 LILFDIGLAHKQLSLSLFEDPPHCK 486
>BE657772
Length = 421
Score = 42.0 bits (97), Expect = 1e-04
Identities = 23/84 (27%), Positives = 42/84 (49%)
Frame = -1
Query: 50 PDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVR 109
PD T + N+ +G V + C++ D+ V + T+TG + KG++A++ G++ +
Sbjct: 412 PDAT--NYEFNEETGKLVVYIPQVCEVGYK-DSSVLRFFTTVTGYLEKGKLADIEGMKTK 242
Query: 110 AFFQWWSITGIRSSGDNIVFEVGM 133
W +T I S G + GM
Sbjct: 241 VLI-WVKVTTILSEGSKLYVTAGM 173
>TC234544 weakly similar to UP|Q9FIV1 (Q9FIV1) Gb|AAF02153.1, partial (79%)
Length = 468
Score = 35.4 bits (80), Expect = 0.011
Identities = 27/107 (25%), Positives = 47/107 (43%), Gaps = 3/107 (2%)
Frame = +3
Query: 30 STPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSN 89
S+P++ H L + G P GLLP+ V + ++T G V L C L +
Sbjct: 132 SSPTTIHDLLRSKGLPPGLLPE-EVKSYTFSET-GHLEVFLDAPC---LTKYENRVLFEQ 296
Query: 90 TITGKIVKGRIAELNGIRVRAFFQWWSITGI---RSSGDNIVFEVGM 133
+T + G + + G++ F W + I S I+F++G+
Sbjct: 297 VVTANLTYGSLIGVEGLQQEELFVWLPVKDIIVNDPSSGLILFDIGL 437
>AW350819
Length = 562
Score = 29.6 bits (65), Expect = 0.62
Identities = 18/43 (41%), Positives = 25/43 (57%), Gaps = 2/43 (4%)
Frame = +1
Query: 3 PKILILFALFLNLTSKSFQNPN--PKPPKSTPSSAHIELTNYG 43
P I+ILFA+ SF +P+ P PKS P S+ + +TN G
Sbjct: 442 PTIVILFAI-------SFSSPDNIPSSPKSLPCSSFL*ITNCG 549
>TC203860 similar to UP|Q8L5P7 (Q8L5P7) LHY protein, partial (48%)
Length = 1950
Score = 29.6 bits (65), Expect = 0.62
Identities = 14/32 (43%), Positives = 19/32 (58%)
Frame = -3
Query: 13 LNLTSKSFQNPNPKPPKSTPSSAHIELTNYGF 44
LN + F P P+P S+PSS+ + LTN F
Sbjct: 1471 LNPVRRVFSFPGPRPMVSSPSSSLLSLTNSRF 1376
>TC225190 weakly similar to UP|Q8VZG3 (Q8VZG3) AT3g48690/T8P19_200, partial
(50%)
Length = 1298
Score = 28.1 bits (61), Expect = 1.8
Identities = 16/65 (24%), Positives = 29/65 (44%)
Frame = +1
Query: 12 FLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLG 71
F+NLTS SF P+ PPK+ + I P+ P ++++ + + + L
Sbjct: 82 FVNLTSNSFLLPS-LPPKAHNNPLQIPSQKLAKPLNSTPPSSIMASTTTEDDSEVTYDLS 258
Query: 72 GACKI 76
K+
Sbjct: 259 PVLKV 273
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,094,607
Number of Sequences: 63676
Number of extensions: 98655
Number of successful extensions: 985
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 957
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 969
length of query: 156
length of database: 12,639,632
effective HSP length: 89
effective length of query: 67
effective length of database: 6,972,468
effective search space: 467155356
effective search space used: 467155356
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0153b.6


