

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0153b.2
(402 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
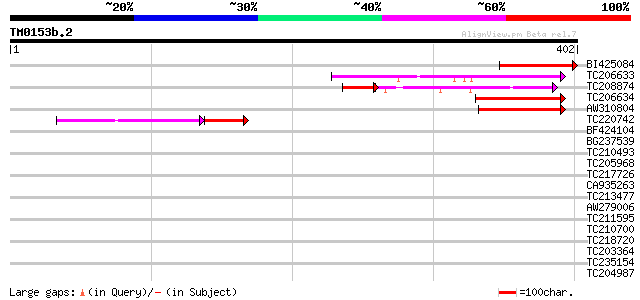
Score E
Sequences producing significant alignments: (bits) Value
BI425084 similar to GP|17381270|gb| AT3g24470/MXP5_4 {Arabidopsi... 113 1e-25
TC206633 113 2e-25
TC208874 80 6e-18
TC206634 77 2e-14
AW310804 66 3e-11
TC220742 52 1e-08
BF424104 37 0.021
BG237539 34 0.13
TC210493 33 0.23
TC205968 31 1.1
TC217726 UP|Q8W199 (Q8W199) Serine acetyltransferase, complete 31 1.1
CA935263 30 1.9
TC213477 30 1.9
AW279006 weakly similar to SP|P98204|ALA1 Phospholipid-transport... 30 1.9
TC211595 weakly similar to UP|Q9DE40 (Q9DE40) Zinc finger protei... 30 2.5
TC210700 30 2.5
TC218720 similar to GB|AAO42946.1|28416831|BT004700 At3g18880 {A... 29 4.3
TC203364 similar to UP|Q6SS00 (Q6SS00) YABBY-like transcription ... 29 4.3
TC235154 29 4.3
TC204987 similar to GB|AAC98030.1|4056457|F5O8 ESTs gb|234051 an... 28 5.7
>BI425084 similar to GP|17381270|gb| AT3g24470/MXP5_4 {Arabidopsis thaliana},
partial (12%)
Length = 425
Score = 113 bits (283), Expect = 1e-25
Identities = 48/55 (87%), Positives = 51/55 (92%)
Frame = +1
Query: 348 LLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAPIIWKNIHTEST 402
LL+GWNSHHSM+KWTIDVGWTS WV+IVNEWLAVCVYLWMLIAPIIWKN T ST
Sbjct: 1 LLIGWNSHHSMRKWTIDVGWTSTWVKIVNEWLAVCVYLWMLIAPIIWKNRQTGST 165
>TC206633
Length = 959
Score = 113 bits (282), Expect = 2e-25
Identities = 62/183 (33%), Positives = 92/183 (49%), Gaps = 17/183 (9%)
Frame = +3
Query: 229 VLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWCAIRSEPEGYEC--IRKSDSPNKTDW 286
+L + V+LHP VN +L ++ LY +LC+ A+ SEP YEC + K T
Sbjct: 3 ILAFVFAIVALHPAVNGSVLPASVISLYCTYLCYSALASEPRDYECNGLHKHSKAVSTGT 182
Query: 287 QNIISLVVGILALVIATFSTGIDSKCFQ---YRKGDKP-----AEEDD-------VPYGY 331
+ L +L++V + G + + KP A+ED+ V Y Y
Sbjct: 183 LTL-GLATTVLSVVYSAVRAGSSAAVLSPPSSPRAGKPLLPLDAKEDEEKEKAKPVTYSY 359
Query: 332 GFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAP 391
FFH +F+ +MY AMLL GW++ +DVGW S WVRI+ W +YLW L+AP
Sbjct: 360 AFFHLIFSLASMYSAMLLTGWSTSVGESGKLVDVGWPSVWVRIITSWATALLYLWSLVAP 539
Query: 392 IIW 394
I++
Sbjct: 540 IMF 548
>TC208874
Length = 769
Score = 79.7 bits (195), Expect(2) = 6e-18
Identities = 54/155 (34%), Positives = 74/155 (46%), Gaps = 26/155 (16%)
Frame = +1
Query: 260 LCWCAI----RSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATF---------ST 306
LC C + SEP YEC + NK+ + +LV+G+L V++ +T
Sbjct: 67 LCLCVLYTGLSSEPRDYEC----NGLNKSRAVSTSTLVLGMLTTVLSVLYSALRAG*STT 234
Query: 307 GIDSKCFQYRKGDKPAEED-------------DVPYGYGFFHFVFATGAMYFAMLLVGWN 353
+ + G KP E+ V Y Y FFH +FA +MY AMLL GW
Sbjct: 235 FLSPQSSPRMVGSKPLLEEAEEGKAKEEKEAQPVIYSYSFFHQIFALASMYSAMLLSGWT 414
Query: 354 SHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWML 388
S S IDVGWTS WVRI EW+ +Y+W++
Sbjct: 415 ST-SES*DLIDVGWTSVWVRIGTEWVTAGLYIWIV 516
Score = 28.9 bits (63), Expect(2) = 6e-18
Identities = 9/25 (36%), Positives = 17/25 (68%)
Frame = +2
Query: 237 VSLHPKVNAGILSPGLMGLYVVFLC 261
++LHP+VN +L ++ LY ++C
Sbjct: 8 IALHPQVNGSLLPSAVISLYCAYVC 82
>TC206634
Length = 410
Score = 76.6 bits (187), Expect = 2e-14
Identities = 31/64 (48%), Positives = 41/64 (63%)
Frame = +2
Query: 331 YGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIA 390
Y FFH +F+ +MY AMLL GW++ +DVGW S WVRI+ W +YLW LIA
Sbjct: 11 YAFFHLIFSLASMYSAMLLTGWSTSVGESGKLVDVGWPSVWVRIITSWATALLYLWSLIA 190
Query: 391 PIIW 394
PI++
Sbjct: 191 PIMF 202
>AW310804
Length = 199
Score = 65.9 bits (159), Expect = 3e-11
Identities = 27/62 (43%), Positives = 39/62 (62%)
Frame = -1
Query: 333 FFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAPI 392
F H +F+ G+MY AMLL GW + + + VGW S WVRI+ W+ +YLW +API
Sbjct: 199 FSHSIFSLGSMYCAMLLTGWFTSVAETGKLVHVGWLSVWVRIITFWVTALLYLWSFVAPI 20
Query: 393 IW 394
++
Sbjct: 19 MF 14
>TC220742
Length = 881
Score = 52.0 bits (123), Expect(2) = 1e-08
Identities = 32/106 (30%), Positives = 50/106 (46%), Gaps = 1/106 (0%)
Frame = +2
Query: 34 KMARYVYALIFLVCNLLAWASRD-ELPGRSVLTKLKGLKTCKDPKDCLGTNGVLRVSMGC 92
+ AR Y +F ++AW R+ P L + K ++ T+ VLRVS+G
Sbjct: 281 RSARIAYCGLFAFSLVVAWILREVAAPLMESLPWINHFKHTPS-REWFETDAVLRVSLGN 457
Query: 93 FLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLL 138
FLFF ++ + RD+ H G W +KI+ W + IF F +
Sbjct: 458 FLFFTILAVLMVGVKTQRDPRDSMHHGGWMMKIICWCLLVIFMFFV 595
Score = 25.0 bits (53), Expect(2) = 1e-08
Identities = 9/31 (29%), Positives = 19/31 (61%)
Frame = +1
Query: 139 PSELIDLYGEVAHFGAGVFLFIQLISIISFI 169
P+E+I Y ++ G+G+FL Q + ++ +
Sbjct: 595 PNEIISFYETISKVGSGMFLLGQGMLLVDLV 687
>BF424104
Length = 409
Score = 36.6 bits (83), Expect = 0.021
Identities = 13/32 (40%), Positives = 21/32 (65%)
Frame = +1
Query: 116 WHSGWWSIKIVLWVAVTIFPFLLPSELIDLYG 147
+H G W+ KIV+W+ + + F LP +I +YG
Sbjct: 262 FHHGGWTAKIVIWLLLVVLAFFLPDAVILVYG 357
>BG237539
Length = 199
Score = 33.9 bits (76), Expect = 0.13
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +2
Query: 238 SLHPKVNAGILSPGLMGLYVVFLCWCAIRSEPEGYE 273
+LHP N + + LY +LC+ A+ EP YE
Sbjct: 41 ALHPAENGSVAPASKISLYCTYLCYSALAREPRDYE 148
>TC210493
Length = 580
Score = 33.1 bits (74), Expect = 0.23
Identities = 24/90 (26%), Positives = 41/90 (44%), Gaps = 2/90 (2%)
Frame = -1
Query: 219 LNISFITFTLVLLQIMTSVSLH--PKVNAGILSPGLMGLYVVFLCWCAIRSEPEGYECIR 276
L++ F F ++L I S++ P++ P + +FL +C I +EP+ +
Sbjct: 301 LHVLFSVFCYLVLYISKCQSIYVDPRIYTYSAIP-----FAIFLIYCLICNEPQFRQGKM 137
Query: 277 KSDSPNKTDWQNIISLVVGILALVIATFST 306
K W ++S GI A V A F+T
Sbjct: 136 KLKEIMSIAWNTLVSSDQGIRAAVTANFAT 47
>TC205968
Length = 433
Score = 30.8 bits (68), Expect = 1.1
Identities = 17/59 (28%), Positives = 26/59 (43%)
Frame = +2
Query: 91 GCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEV 149
G LFF++ F R + + T H W +VLW+ + PF P L + +V
Sbjct: 182 GFVLFFVLEFVHG*RRRRGVKREGTIHGSLWK*VVVLWLLCKLAPFKTPEFLFHAFFQV 358
>TC217726 UP|Q8W199 (Q8W199) Serine acetyltransferase, complete
Length = 1186
Score = 30.8 bits (68), Expect = 1.1
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = +2
Query: 84 GVLRVSMGCFLFFMMMFWSTARASKL 109
G+LR+ MGC L ++ FW R K+
Sbjct: 719 GILRLGMGCLLVLVLPFWGILRLGKV 796
>CA935263
Length = 446
Score = 30.0 bits (66), Expect = 1.9
Identities = 16/38 (42%), Positives = 25/38 (65%), Gaps = 2/38 (5%)
Frame = -1
Query: 280 SPNKTDWQNIISLVVGILALVIA--TFSTGIDSKCFQY 315
SPN++ W++ + V+ I A V+A +FSTG SKC +
Sbjct: 197 SPNRSCWRSCLGHVISIAAAVMAQYSFSTG-SSKCILF 87
>TC213477
Length = 357
Score = 30.0 bits (66), Expect = 1.9
Identities = 17/56 (30%), Positives = 31/56 (55%), Gaps = 3/56 (5%)
Frame = +1
Query: 247 ILSPGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKT---DWQNIISLVVGILAL 299
++ L ++ VFLC +IRSE YE I + +KT +W ++++ + +L L
Sbjct: 52 MIEQNLDKVFKVFLCCFSIRSEVSSYENITLIEKQHKTLLKNWI*VLAITI*LLIL 219
>AW279006 weakly similar to SP|P98204|ALA1 Phospholipid-transporting ATPase 1
(EC 3.6.3.1) (Aminophospholipid flippase 1). [Mouse-ear
cress], partial (7%)
Length = 424
Score = 30.0 bits (66), Expect = 1.9
Identities = 20/82 (24%), Positives = 36/82 (43%), Gaps = 13/82 (15%)
Frame = +3
Query: 329 YGYGFFHFVFATGAMYFAMLLVGWNS------------HHSMKKWTIDVGWTSAWVRIVN 376
YG G H + +F M+ W S ++ W++ WT + V +VN
Sbjct: 81 YGAGHRHEAYNMQLFWFTMIDTLWQSLVLFYIPVFIYKDSTIDIWSMGSLWTISVVILVN 260
Query: 377 EWLAVCVYLWMLIAPI-IWKNI 397
LA+ + W L++ + +W +I
Sbjct: 261 VHLAMDINQWALVSHVAVWGSI 326
>TC211595 weakly similar to UP|Q9DE40 (Q9DE40) Zinc finger protein Zic2,
partial (5%)
Length = 607
Score = 29.6 bits (65), Expect = 2.5
Identities = 21/75 (28%), Positives = 35/75 (46%), Gaps = 6/75 (8%)
Frame = +3
Query: 130 AVTIFPFLLPSELIDLYGEVAHFGAG-----VFLFIQLISIISFITWLNDF-FASEKYAE 183
++ + P LPS + AHF G + +F IS+ S +TWLND +S +
Sbjct: 243 SMLLLPPSLPSSMQHNKIAFAHFA*GFCSSHILMFKTTISLFSLLTWLNDKPISSIPFPL 422
Query: 184 RCQIHVMLFATASYF 198
+ H+ L ++F
Sbjct: 423 KFPFHLSLSMMLTFF 467
>TC210700
Length = 860
Score = 29.6 bits (65), Expect = 2.5
Identities = 17/52 (32%), Positives = 29/52 (55%), Gaps = 2/52 (3%)
Frame = +3
Query: 162 LISIISFITWLNDFFASEKYAERCQI--HVMLFATASYFICMVGVILMYIWY 211
++SII + +L+ FF Y +CQ+ HV+ F +FI G +++ WY
Sbjct: 345 ILSIIPTLLFLSFFF----YLPKCQLR*HVLFFNI--FFINFKGTVILLTWY 482
>TC218720 similar to GB|AAO42946.1|28416831|BT004700 At3g18880 {Arabidopsis
thaliana;} , partial (79%)
Length = 804
Score = 28.9 bits (63), Expect = 4.3
Identities = 9/28 (32%), Positives = 18/28 (64%)
Frame = -3
Query: 337 VFATGAMYFAMLLVGWNSHHSMKKWTID 364
V+ T + + ++L W+SH+ M WT++
Sbjct: 547 VWTTLSQFMKLVLQSWSSHYQMLAWTVE 464
>TC203364 similar to UP|Q6SS00 (Q6SS00) YABBY-like transcription factor
GRAMINIFOLIA, partial (65%)
Length = 1094
Score = 28.9 bits (63), Expect = 4.3
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = -1
Query: 267 SEPEGYECIRKSDSPNKTDWQ 287
S PEGYE R+ PN+ +W+
Sbjct: 371 SPPEGYEMGRRKSDPNEIEWE 309
>TC235154
Length = 437
Score = 28.9 bits (63), Expect = 4.3
Identities = 20/67 (29%), Positives = 33/67 (48%), Gaps = 1/67 (1%)
Frame = +1
Query: 34 KMARYVYALIFLVCNLLAWASRD-ELPGRSVLTKLKGLKTCKDPKDCLGTNGVLRVSMGC 92
+ AR Y +F ++AW R+ P L + K ++ T+ VLRVS+G
Sbjct: 235 RSARIAYCGLFAFSLVVAWILREVAAPLMESLPWINHFKHTPS-REWFETDAVLRVSLGN 411
Query: 93 FLFFMMM 99
+LFF ++
Sbjct: 412 YLFFTIL 432
>TC204987 similar to GB|AAC98030.1|4056457|F5O8 ESTs gb|234051 and gb|F13722
come from this gene. {Arabidopsis thaliana;} , partial
(96%)
Length = 806
Score = 28.5 bits (62), Expect = 5.7
Identities = 13/39 (33%), Positives = 22/39 (56%)
Frame = -2
Query: 214 QPSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGL 252
+P C+ ++ TFT+ + ++ S VN G L+PGL
Sbjct: 202 RPFCITILALTTFTVKVCPLVPGRS*STLVNTGFLNPGL 86
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.140 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,515,948
Number of Sequences: 63676
Number of extensions: 418721
Number of successful extensions: 3203
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 3167
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3201
length of query: 402
length of database: 12,639,632
effective HSP length: 99
effective length of query: 303
effective length of database: 6,335,708
effective search space: 1919719524
effective search space used: 1919719524
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0153b.2


