

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0142.11
(270 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
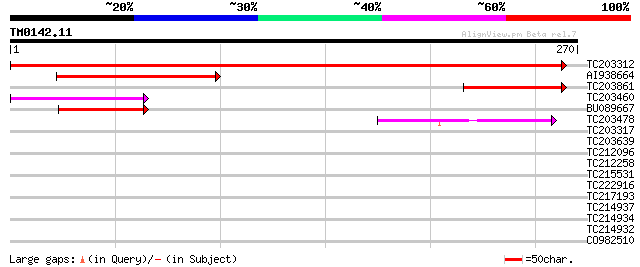
Score E
Sequences producing significant alignments: (bits) Value
TC203312 similar to UP|Q9FPH6 (Q9FPH6) AT3g15840, partial (77%) 446 e-126
AI938664 similar to GP|11762196|gb| AT3g15840 {Arabidopsis thali... 122 2e-28
TC203861 similar to UP|Q9ATM4 (Q9ATM4) Plasma membrane integral ... 95 3e-20
TC203460 similar to UP|Q8IQR2 (Q8IQR2) CG32182-PA, partial (4%) 54 6e-08
BU089667 52 4e-07
TC203478 similar to UP|Q9AXG0 (Q9AXG0) Ribulose-1,5-bisphosphate... 44 8e-05
TC203317 similar to UP|Q7X9A0 (Q7X9A0) Rubisco activase alpha fo... 39 0.003
TC203639 similar to UP|Q9AXG0 (Q9AXG0) Ribulose-1,5-bisphosphate... 38 0.004
TC212096 32 0.30
TC212258 UP|Q34597 (Q34597) Maturase-related protein (Fragment),... 28 3.3
TC215531 UP|HEM2_SOYBN (P43210) Delta-aminolevulinic acid dehydr... 27 7.5
TC222916 similar to UP|Q944S2 (Q944S2) At2g47790/F17A22.18 (Expr... 27 7.5
TC217193 similar to UP|Q7XJN2 (Q7XJN2) At2g40650 protein, partia... 27 9.7
TC214937 homologue to UP|Q43015 (Q43015) Alcohol dehydrogenase-1... 27 9.7
TC214934 homologue to UP|Q43016 (Q43016) Alcohol dehydrogenase-1... 27 9.7
TC214932 homologue to UP|Q8LJR2 (Q8LJR2) Alcohol dehydrogenase 1... 27 9.7
CO982510 27 9.7
>TC203312 similar to UP|Q9FPH6 (Q9FPH6) AT3g15840, partial (77%)
Length = 1165
Score = 446 bits (1146), Expect = e-126
Identities = 208/265 (78%), Positives = 233/265 (87%)
Frame = +2
Query: 1 MYDSSDLLFFSVPPSNPITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEI 60
++ SS VPP+ P T FLNVS PR YLVK+++VK ++ I+AA TT +EI
Sbjct: 107 IFASSTQPCLPVPPTIPNTLATPFLNVSSPRSYLVKKKHVKFSKKISAAAVATTTTTEEI 286
Query: 61 EEYKIPSWANFELGKASVYWKTMNGLPPTSGEKLKLFYNPTSTQLAPNEEFGIAFNGGFN 120
+EYK+PSWA FE+G+A+VYWKTMNGLPPTSGEKLKLFYNP +TQL PNEEFGIAFNGGFN
Sbjct: 287 QEYKLPSWAKFEIGRAAVYWKTMNGLPPTSGEKLKLFYNPAATQLVPNEEFGIAFNGGFN 466
Query: 121 QPIMCGGEPRAMLRKDRGKADAPIYSIQICVPKHALNLIFSFTNGVDWDGPYRLQFQVPK 180
QPIMCGGEPRAMLRKDRGKADAPIYSIQIC+PKHALNLIFSFTNGVDWDGPYRLQFQVPK
Sbjct: 467 QPIMCGGEPRAMLRKDRGKADAPIYSIQICIPKHALNLIFSFTNGVDWDGPYRLQFQVPK 646
Query: 181 VLQNKPIEFFNEGLAEELGKEGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEG 240
LQNKPI+FFN+GLAEEL KEGACEQAIFPD+NK+ITKCAM+GNL+ EGGDRCDLN V G
Sbjct: 647 ALQNKPIDFFNKGLAEELSKEGACEQAIFPDTNKIITKCAMIGNLSKEGGDRCDLNFVLG 826
Query: 241 CTDPSSHLYNPLANVDDGTCPLDLD 265
CTDPSSHLYNPLANVDDGTC ++LD
Sbjct: 827 CTDPSSHLYNPLANVDDGTCTIELD 901
>AI938664 similar to GP|11762196|gb| AT3g15840 {Arabidopsis thaliana},
partial (16%)
Length = 392
Score = 122 bits (305), Expect = 2e-28
Identities = 54/78 (69%), Positives = 67/78 (85%)
Frame = +1
Query: 23 TFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEIEEYKIPSWANFELGKASVYWKT 82
+FLNVS PR YLVK+++VK ++ I+AA TT +EI+EYK+PSWA FE+G+A+VYWKT
Sbjct: 154 SFLNVSSPRSYLVKKKHVKFSKKISAAAVATTTTTEEIQEYKLPSWAKFEIGRAAVYWKT 333
Query: 83 MNGLPPTSGEKLKLFYNP 100
MNGLPPTSGEKLKLFYNP
Sbjct: 334 MNGLPPTSGEKLKLFYNP 387
>TC203861 similar to UP|Q9ATM4 (Q9ATM4) Plasma membrane integral protein
ZmPIP2-7, partial (57%)
Length = 804
Score = 95.1 bits (235), Expect = 3e-20
Identities = 41/49 (83%), Positives = 45/49 (91%)
Frame = +1
Query: 217 TKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTCPLDLD 265
TKCAM+GNL+ EGGDRCDLN V GCTDPSSHLYNPLANVDDGTC ++LD
Sbjct: 1 TKCAMIGNLSKEGGDRCDLNFVLGCTDPSSHLYNPLANVDDGTCTIELD 147
>TC203460 similar to UP|Q8IQR2 (Q8IQR2) CG32182-PA, partial (4%)
Length = 458
Score = 54.3 bits (129), Expect = 6e-08
Identities = 29/66 (43%), Positives = 40/66 (59%)
Frame = +2
Query: 1 MYDSSDLLFFSVPPSNPITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEI 60
++ SS V P+ T FLNVS PR YLVK+++VK ++ I+AA TT +EI
Sbjct: 104 IFASSTQSCLPVSPTISNTLVSPFLNVSSPRSYLVKKKHVKFSKHISAAAVATTTTTEEI 283
Query: 61 EEYKIP 66
+EYK P
Sbjct: 284 QEYKFP 301
>BU089667
Length = 356
Score = 51.6 bits (122), Expect = 4e-07
Identities = 23/43 (53%), Positives = 32/43 (73%)
Frame = +2
Query: 24 FLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEIEEYKIP 66
FLNVS PR YLVK+++VK ++ I+AA TT +EI+EY+ P
Sbjct: 98 FLNVSSPRSYLVKKKHVKFSKKISAAAVATTTTTEEIQEYEFP 226
>TC203478 similar to UP|Q9AXG0 (Q9AXG0) Ribulose-1,5-bisphosphate
carboxylase/oxygenase activase 2 (Fragment), partial
(31%)
Length = 623
Score = 43.9 bits (102), Expect = 8e-05
Identities = 27/91 (29%), Positives = 45/91 (48%), Gaps = 6/91 (6%)
Frame = +2
Query: 176 FQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAMLGNLTVEG 229
F+ PK+ +K +E+ N + E+ + +A D+N K GN +G
Sbjct: 125 FEQPKMTLSKLLEYGNMLVQEQENVKRVQLADKYLNEAALGDANDDAIKT---GNFYGQG 295
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ + + EGCTDP++ Y+P A DDG+C
Sbjct: 296 AQQVHVPVPEGCTDPTAENYDPTARSDDGSC 388
>TC203317 similar to UP|Q7X9A0 (Q7X9A0) Rubisco activase alpha form precursor,
partial (94%)
Length = 1740
Score = 38.5 bits (88), Expect = 0.003
Identities = 25/91 (27%), Positives = 43/91 (46%), Gaps = 6/91 (6%)
Frame = +2
Query: 176 FQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAMLGNLTVEG 229
F PK+ +K +E+ N + E+ + +A D+N+ K G +
Sbjct: 1253 FDQPKMTLSKLLEYGNMLVQEQENVKRVQLADKYLNEAALGDANQDAIK---RGTFYGKA 1423
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ + + EGCTDP++ ++P A DDGTC
Sbjct: 1424 AQQVKIPVPEGCTDPNASNFDPTARSDDGTC 1516
>TC203639 similar to UP|Q9AXG0 (Q9AXG0) Ribulose-1,5-bisphosphate
carboxylase/oxygenase activase 2 (Fragment), partial
(30%)
Length = 665
Score = 38.1 bits (87), Expect = 0.004
Identities = 27/96 (28%), Positives = 43/96 (44%), Gaps = 11/96 (11%)
Frame = +2
Query: 176 FQVPKVLQNKPIEFFNE-----------GLAEELGKEGACEQAIFPDSNKVITKCAMLGN 224
F PK+ +K +E+ N LA++ KE A A N+ G
Sbjct: 122 FDQPKMTLSKLLEYGNMLVQEQENVKRVQLADKYLKEAALGDANQDSINR--------GT 277
Query: 225 LTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ + ++ + EGCTDP++ ++P A DDGTC
Sbjct: 278 FYGKAAQQVNIPVPEGCTDPNASNFDPTARSDDGTC 385
>TC212096
Length = 246
Score = 32.0 bits (71), Expect = 0.30
Identities = 13/27 (48%), Positives = 18/27 (66%)
Frame = +2
Query: 137 RGKADAPIYSIQICVPKHALNLIFSFT 163
R K + P+Y I CVPK+ L LI ++T
Sbjct: 122 RSKVEIPLYCITFCVPKYLLILISNYT 202
>TC212258 UP|Q34597 (Q34597) Maturase-related protein (Fragment), complete
Length = 2309
Score = 28.5 bits (62), Expect = 3.3
Identities = 18/51 (35%), Positives = 27/51 (52%), Gaps = 6/51 (11%)
Frame = +3
Query: 123 IMCGGEPRAMLRKDRGKAD--API----YSIQICVPKHALNLIFSFTNGVD 167
I GGE R ++R +RG A AP Y I+IC ++A +L+ V+
Sbjct: 843 IELGGEARLVIRSERGLARKLAPFKKNHYFIRICYVRYADDLLLGIVGAVE 995
>TC215531 UP|HEM2_SOYBN (P43210) Delta-aminolevulinic acid dehydratase,
chloroplast precursor (Porphobilinogen synthase)
(ALADH) , complete
Length = 1737
Score = 27.3 bits (59), Expect = 7.5
Identities = 13/46 (28%), Positives = 25/46 (54%), Gaps = 6/46 (13%)
Frame = -3
Query: 6 DLLFFSVPPSNPITSRHT------FLNVSLPRCYLVKERNVKVTRT 45
+L + S+PP P+ SRH+ F V+ R + ++ RN + ++
Sbjct: 601 NLFYKSMPPPKPVASRHSPNWSILFTFVNEKRIHEIRRRNARFLKS 464
>TC222916 similar to UP|Q944S2 (Q944S2) At2g47790/F17A22.18 (Expressed
protein), partial (5%)
Length = 419
Score = 27.3 bits (59), Expect = 7.5
Identities = 12/36 (33%), Positives = 17/36 (46%)
Frame = -1
Query: 72 ELGKASVYWKTMNGLPPTSGEKLKLFYNPTSTQLAP 107
++ + S++WK GL K FYN T L P
Sbjct: 419 KMTQVSIFWKKQKGL*NMLTYKYNFFYNKNPTHLNP 312
>TC217193 similar to UP|Q7XJN2 (Q7XJN2) At2g40650 protein, partial (88%)
Length = 1691
Score = 26.9 bits (58), Expect = 9.7
Identities = 15/34 (44%), Positives = 19/34 (55%)
Frame = +1
Query: 198 LGKEGACEQAIFPDSNKVITKCAMLGNLTVEGGD 231
LGKE E I D+ VI M+G +T+E GD
Sbjct: 868 LGKETGIEDTIVIDTGTVIMTETMIG-ITIESGD 966
>TC214937 homologue to UP|Q43015 (Q43015) Alcohol dehydrogenase-1CN ,
partial (41%)
Length = 803
Score = 26.9 bits (58), Expect = 9.7
Identities = 21/94 (22%), Positives = 37/94 (39%), Gaps = 7/94 (7%)
Frame = +1
Query: 97 FYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAMLRKDRGKADAPIYSIQICVPK--- 153
F NP E NGG ++ + C G +AM+ D ++ + VP
Sbjct: 52 FVNPKDHDKPVQEVIAAMTNGGVDRAVECTGSIQAMISAFECVHDGWGVAVLVGVPNKDD 231
Query: 154 ----HALNLIFSFTNGVDWDGPYRLQFQVPKVLQ 183
H +N + T + G Y+ + +P V++
Sbjct: 232 AFKTHPVNFLNERTLKGTFYGNYKPRTDLPSVVE 333
>TC214934 homologue to UP|Q43016 (Q43016) Alcohol dehydrogenase-1F ,
complete
Length = 1450
Score = 26.9 bits (58), Expect = 9.7
Identities = 21/94 (22%), Positives = 37/94 (39%), Gaps = 7/94 (7%)
Frame = +3
Query: 97 FYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAMLRKDRGKADAPIYSIQICVPK--- 153
F NP E NGG ++ + C G +AM+ D ++ + VP
Sbjct: 822 FVNPKDHDKPVQEVIAAMTNGGVDRAVECTGSIQAMISAFECVHDGWGVAVLVGVPNKDD 1001
Query: 154 ----HALNLIFSFTNGVDWDGPYRLQFQVPKVLQ 183
H +N + T + G Y+ + +P V++
Sbjct: 1002AFKTHPVNFLNERTLKGTFYGNYKPRTDLPSVVE 1103
>TC214932 homologue to UP|Q8LJR2 (Q8LJR2) Alcohol dehydrogenase 1 (Fragment),
complete
Length = 1456
Score = 26.9 bits (58), Expect = 9.7
Identities = 25/102 (24%), Positives = 40/102 (38%), Gaps = 10/102 (9%)
Frame = +3
Query: 97 FYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAMLRKDRGKADAPIYSIQICVPK--- 153
F NP E NGG ++ I C G +A + D ++ + VPK
Sbjct: 819 FVNPKDHNKPVQEVIAEMTNGGVDRAIECTGSIQASISAFECTHDGWGTAVLVSVPKKDA 998
Query: 154 ----HALNLIFSFTNGVDWDGPYRLQFQVPKVLQ---NKPIE 188
H + + T + G YR + +P V++ NK +E
Sbjct: 999 EFKTHPMKFMEGRTLKGTFYGHYRPRTDIPGVVEKYLNKELE 1124
>CO982510
Length = 488
Score = 26.9 bits (58), Expect = 9.7
Identities = 11/32 (34%), Positives = 15/32 (46%)
Frame = -1
Query: 108 NEEFGIAFNGGFNQPIMCGGEPRAMLRKDRGK 139
N FG+ G N C GE R + K+ G+
Sbjct: 317 NGSFGVEVGGSRNHHFACSGETRFHVEKEEGE 222
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,418,940
Number of Sequences: 63676
Number of extensions: 167000
Number of successful extensions: 749
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 747
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 749
length of query: 270
length of database: 12,639,632
effective HSP length: 96
effective length of query: 174
effective length of database: 6,526,736
effective search space: 1135652064
effective search space used: 1135652064
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0142.11


