

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.7
(571 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
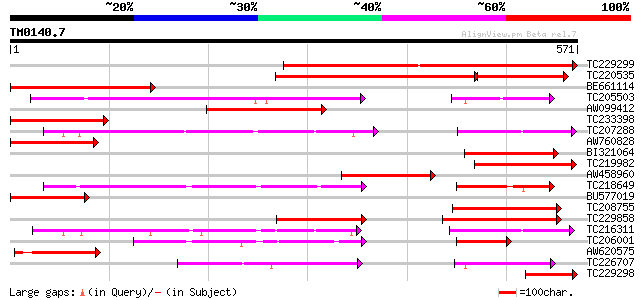
Score E
Sequences producing significant alignments: (bits) Value
TC229299 similar to UP|GT10_HUMAN (O95528) Solute carrier family... 526 e-149
TC220535 similar to UP|HXT5_YEAST (P38695) Probable glucose tran... 348 e-131
BE661114 265 3e-71
TC205503 UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, complete 191 8e-49
AW099412 191 1e-48
TC233398 187 8e-48
TC207288 UP|Q7XA51 (Q7XA51) Monosaccharide transporter, complete 176 3e-44
AW760828 166 2e-41
BI321064 165 4e-41
TC219982 162 5e-40
AW458960 153 2e-37
TC218649 similar to UP|Q39416 (Q39416) Integral membrane protein... 148 6e-36
BU577019 145 5e-35
TC208755 similar to UP|HXT4_YEAST (P32467) Low-affinity glucose ... 143 2e-34
TC229858 weakly similar to UP|Q8VHD6 (Q8VHD6) Glucose transporte... 143 2e-34
TC216311 UP|Q7XA52 (Q7XA52) Monosaccharide transporter, complete 129 5e-30
TC206001 similar to UP|Q8VZI5 (Q8VZI5) AT5g18840/F17K4_90, parti... 110 2e-24
AW620575 102 5e-22
TC226707 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter... 101 8e-22
TC229298 weakly similar to UP|Q8MKK4 (Q8MKK4) AT19440p (CG8234-P... 99 7e-21
>TC229299 similar to UP|GT10_HUMAN (O95528) Solute carrier family 2,
facilitated glucose transporter, member 10 (Glucose
transporter type 10), partial (6%)
Length = 1060
Score = 526 bits (1354), Expect = e-149
Identities = 254/298 (85%), Positives = 278/298 (93%), Gaps = 2/298 (0%)
Frame = +2
Query: 276 GMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKT 335
G+GL FQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLI SGLNAFGSILSIYFIDKT
Sbjct: 2 GVGLLIFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLIISGLNAFGSILSIYFIDKT 181
Query: 336 GRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAIETSHYNN--TCPDFRAAVNPGRW 393
GRKKLALISLCGVV SLALLT AFRESE+H+P VSAI++S +NN TCPD++ A+N W
Sbjct: 182 GRKKLALISLCGVVFSLALLTAAFRESEIHSPMVSAIQSSQFNNNNTCPDYKTALNSAEW 361
Query: 394 TCMTCLKASPSCGFCAASPNKLLPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALV 453
TCMTCLKASPSCG+CAA +K LPGACLIS+D TK MCG+DHRAWYTRGCPSK+GWAAL+
Sbjct: 362 TCMTCLKASPSCGYCAAD-DKFLPGACLISNDGTKKMCGDDHRAWYTRGCPSKYGWAALI 538
Query: 454 GLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIG 513
GLALYII FSPGMGTVPWV+NSEIYPLRYRGVCGGIASTTVWISNLIVS+SFLSLT+ +G
Sbjct: 539 GLALYIIFFSPGMGTVPWVVNSEIYPLRYRGVCGGIASTTVWISNLIVSESFLSLTKALG 718
Query: 514 TAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQKRDSGSEKH 571
TAWTFMMFGI+A+VAIFFVI+FVPETKGVPMEEVEKMLEQRS+QFKFW+KRDSGSEKH
Sbjct: 719 TAWTFMMFGIVAIVAIFFVIIFVPETKGVPMEEVEKMLEQRSVQFKFWEKRDSGSEKH 892
>TC220535 similar to UP|HXT5_YEAST (P38695) Probable glucose transporter
HXT5, partial (3%)
Length = 1007
Score = 348 bits (892), Expect(2) = e-131
Identities = 163/206 (79%), Positives = 180/206 (87%)
Frame = +1
Query: 268 TVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSIL 327
TVRRGLYAGMGLQ FQQFVGINTVMYYSPTIVQLAGFASNR ALLLSL+T+GLNAFGSIL
Sbjct: 1 TVRRGLYAGMGLQIFQQFVGINTVMYYSPTIVQLAGFASNRVALLLSLVTAGLNAFGSIL 180
Query: 328 SIYFIDKTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAIETSHYNNTCPDFRAA 387
SIYFIDKTGR+KL L SLCGVV+SL +LTVAF E+ H+P VS IETSH+NNTCPD+ A
Sbjct: 181 SIYFIDKTGRRKLLLFSLCGVVVSLVVLTVAFHETTTHSPMVSTIETSHFNNTCPDYSTA 360
Query: 388 VNPGRWTCMTCLKASPSCGFCAASPNKLLPGACLISDDTTKDMCGNDHRAWYTRGCPSKF 447
NPG W CM CLKASP CGFCA+ NKLLPGACLIS+DTT++ C + R WYTRGCPS++
Sbjct: 361 FNPGEWDCMKCLKASPECGFCASRANKLLPGACLISNDTTENQCQKEDRLWYTRGCPSQY 540
Query: 448 GWAALVGLALYIITFSPGMGTVPWVI 473
GW ALVGLALYII FSPGMGTVPWV+
Sbjct: 541 GWLALVGLALYIIFFSPGMGTVPWVV 618
Score = 140 bits (354), Expect(2) = e-131
Identities = 66/91 (72%), Positives = 80/91 (87%)
Frame = +2
Query: 472 VINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFF 531
++ SEIYPLRYRG+CGG+AST+ W+SNLIV+QSFLSLTQ IGT+ TFM+F + V AI F
Sbjct: 614 LLYSEIYPLRYRGICGGMASTSNWVSNLIVAQSFLSLTQAIGTSSTFMIFIFITVAAIVF 793
Query: 532 VIVFVPETKGVPMEEVEKMLEQRSLQFKFWQ 562
VI+FVPETKG+P+EEVE MLE+RSL FKFWQ
Sbjct: 794 VIIFVPETKGLPIEEVENMLERRSLNFKFWQ 886
>BE661114
Length = 634
Score = 265 bits (678), Expect = 3e-71
Identities = 133/147 (90%), Positives = 140/147 (94%)
Frame = +3
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYI+D+FKA
Sbjct: 177 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIKDEFKA 356
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 120
VDRKTWLQEAIVSTAIAGAIIGA+VGGWINDRFGR+ I+IADTLF IGSVIMAAA +PA
Sbjct: 357 VDRKTWLQEAIVSTAIAGAIIGASVGGWINDRFGRKKGIVIADTLFFIGSVIMAAASSPA 536
Query: 121 ILLFGRVFVGFGVGMASMASPLYISEA 147
IL+ GRVFVG GV MASMASPLYISEA
Sbjct: 537 ILIVGRVFVGIGVXMASMASPLYISEA 617
>TC205503 UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, complete
Length = 1928
Score = 191 bits (485), Expect = 8e-49
Identities = 113/351 (32%), Positives = 185/351 (52%), Gaps = 14/351 (3%)
Frame = +1
Query: 22 KNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAII 81
+N Y A A + +L GYD GV+SGA +YI+ D K D + + I++ ++I
Sbjct: 244 RNKYAFACAMLASMTSILLGYDIGVMSGAAIYIKRDLKVSDEQIEILLGIINLY---SLI 414
Query: 82 GAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASP 141
G+ + G +D GRR I + +FL+GS +M P+ + L+ GR G G+G A M +P
Sbjct: 415 GSCLAGRTSDWIGRRYTIGLGGAIFLVGSTLMGFYPHYSFLMCGRFVAGIGIGYALMIAP 594
Query: 142 LYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFT--NAPGTWRWMLGVAAAPAIIQ 199
+Y +E SP RG L S I GG L Y+ N F+ WR MLGV A P+++
Sbjct: 595 VYTAEVSPASSRGFLTSFPEVFINGGILLGYISNYGFSKLTLKVGWRMMLGVGAIPSVVL 774
Query: 200 IVLMLSLPESPRWLYRKGREEESKSILKKIY-APEDVDAEIEALKESV-------ESEIE 251
+L++PESPRWL +GR E++ +L K + E+ + +K++ + ++
Sbjct: 775 TEGVLAMPESPRWLVMRGRLGEARKVLNKTSDSKEEAQLRLAEIKQAAGIPESCNDDVVQ 954
Query: 252 ESKTSK----ISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASN 307
+K S + L T +R + A +G+ FFQQ G++ V+ YSP I + AG ++
Sbjct: 955 VNKQSNGEGVWKELFLYPTPAIRHIVIAALGIHFFQQASGVDAVVLYSPRIFEKAGITND 1134
Query: 308 RTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVA 358
LL ++ + + + + +D+ GR+ L L S+ G+VLSL L ++
Sbjct: 1135THKLLATVAVGFVKTVFILAATFTLDRVGRRPLLLSSVGGMVLSLLTLAIS 1287
Score = 68.9 bits (167), Expect = 6e-12
Identities = 40/108 (37%), Positives = 60/108 (55%), Gaps = 5/108 (4%)
Frame = +1
Query: 446 KFGWAALVGLAL---YIITFSPGMGTVPWVINSEIYPLRYR--GVCGGIASTTVWISNLI 500
K WA +A+ Y+ TFS G G + WV +SEI+PLR R G G+A ++ +
Sbjct: 1315 KLMWAVGSSIAMVLAYVATFSIGAGPITWVYSSEIFPLRLRAQGAAAGVAVNRT--TSAV 1488
Query: 501 VSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVE 548
VS +FLSLT+ I F ++ +A V F +PET+G +E++E
Sbjct: 1489 VSMTFLSLTRAITIGGAFFLYCGIATVGWIFFYTVLPETRGKTLEDME 1632
>AW099412
Length = 373
Score = 191 bits (484), Expect = 1e-48
Identities = 97/122 (79%), Positives = 111/122 (90%), Gaps = 1/122 (0%)
Frame = +1
Query: 199 QIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTS-K 257
QIVLM+ LPESPRWL+RKGREEE K+IL+KIY P++V+AEI LKESVE EI+E++ S K
Sbjct: 7 QIVLMMMLPESPRWLFRKGREEEGKAILRKIYPPQEVEAEINTLKESVEIEIKEAEASDK 186
Query: 258 ISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLIT 317
+S++ +LKT TVRRGLYAGMGLQ FQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLIT
Sbjct: 187 VSIVKMLKTKTVRRGLYAGMGLQIFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLIT 366
Query: 318 SG 319
SG
Sbjct: 367 SG 372
>TC233398
Length = 442
Score = 187 bits (476), Expect = 8e-48
Identities = 91/99 (91%), Positives = 95/99 (95%)
Frame = +3
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
MEGGVPEAD+SAFRECLSLSWKNPYVLRLAFSAGIGG LFGYDTGVISGALLYIRDDFK
Sbjct: 144 MEGGVPEADISAFRECLSLSWKNPYVLRLAFSAGIGGFLFGYDTGVISGALLYIRDDFKE 323
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAI 99
VDRKTWLQEAIVS A+AGAIIGA+VGGWINDRFGR+ AI
Sbjct: 324 VDRKTWLQEAIVSMALAGAIIGASVGGWINDRFGRKKAI 440
>TC207288 UP|Q7XA51 (Q7XA51) Monosaccharide transporter, complete
Length = 1813
Score = 176 bits (445), Expect = 3e-44
Identities = 116/360 (32%), Positives = 190/360 (52%), Gaps = 23/360 (6%)
Frame = +1
Query: 35 IGGLLFGYDTGVISGALL---YIRDDFKAVDRKTWLQ--------------EAIVSTAIA 77
+GG LFGYD GV G ++++ F V R+ + S+
Sbjct: 130 LGGSLFGYDLGVSGGVPSMDDFLKEFFPKVYRRKQMHLHETDYCKYDDQVLTLFTSSLYF 309
Query: 78 GAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMAS 137
A++ ++ + GR+ +II+ FL G+++ AAA N A+L+ GRV +G G+G +
Sbjct: 310 SALVMTFFASFLTRKKGRKASIIVGALSFLAGAILNAAAKNIAMLIIGRVLLGGGIGFGN 489
Query: 138 MASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNA--PGTWRWMLGVAAAP 195
A PLY+SE +P + RGA+ L F G ++ L+N FT P WR LG+A P
Sbjct: 490 QAVPLYLSEMAPAKNRGAVNQLFQFTTCAGILIANLVNY-FTEKIHPYGWRISLGLAGLP 666
Query: 196 AIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKT 255
A +V + E+P L +GR +++K +L++I E+V+AE E LKE+ EE++
Sbjct: 667 AFAMLVGGICCAETPNSLVEQGRLDKAKQVLQRIRGTENVEAEFEDLKEA----SEEAQA 834
Query: 256 SKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSL 315
K TLLK + + +G+ FQQ G N++++Y+P I Q GF +N +L S
Sbjct: 835 VKSPFRTLLKRKYRPQLIIGALGIPAFQQLTGNNSILFYAPVIFQSLGFGAN-ASLFSSF 1011
Query: 316 ITSGLNAFGSILSIYFIDKTGRKKLALIS----LCGVVLSLALLTVAFRESELHAPTVSA 371
IT+G +++S++ +DK GR+K L + +C ++++ A+L V F + VSA
Sbjct: 1012ITNGALLVATVISMFLVDKYGRRKFFLEAGFEMICCMIITGAVLAVNFGHGKEIGKGVSA 1191
Score = 56.6 bits (135), Expect = 3e-08
Identities = 33/119 (27%), Positives = 57/119 (47%)
Frame = +1
Query: 452 LVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQT 511
+V + L+++ + G + W++ SE++PL R I I +V+Q FL
Sbjct: 1198 VVVIFLFVLAYGRSWGPLGWLVPSELFPLEIRSSAQSIVVCVNMIFTALVAQLFLMSLCH 1377
Query: 512 IGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQKRDSGSEK 570
+ F++F L + FFV +PETK VP+EE+ + E +F +D + K
Sbjct: 1378 LKFG-IFLLFASLIIFMSFFVFFLLPETKKVPIEEIYLLFENHWFWRRFVTDQDPETSK 1551
>AW760828
Length = 423
Score = 166 bits (421), Expect = 2e-41
Identities = 81/89 (91%), Positives = 84/89 (94%)
Frame = +1
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
MEGG E D SAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGA+LYIRDDFKA
Sbjct: 157 MEGGGVEVDASAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGAILYIRDDFKA 336
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWI 89
VDRKTWLQEAIVS A+AGAI+GAAVGGWI
Sbjct: 337 VDRKTWLQEAIVSMALAGAIVGAAVGGWI 423
>BI321064
Length = 284
Score = 165 bits (418), Expect = 4e-41
Identities = 76/94 (80%), Positives = 87/94 (91%)
Frame = +3
Query: 459 IITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTF 518
II FSPGM TVPWV+NSEIYPLRYRGVCGGIASTT W+SNLIVSQSFL+LT IGTAWTF
Sbjct: 3 IIFFSPGMDTVPWVVNSEIYPLRYRGVCGGIASTTCWVSNLIVSQSFLTLTVAIGTAWTF 182
Query: 519 MMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLE 552
M+FG +A++ IFFV++FVPETKGVP+EEVE+MLE
Sbjct: 183 MLFGFVALIGIFFVLIFVPETKGVPVEEVEQMLE 284
>TC219982
Length = 590
Score = 162 bits (409), Expect = 5e-40
Identities = 73/102 (71%), Positives = 88/102 (85%)
Frame = +2
Query: 469 VPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVA 528
VPWV+NSEIYPLRYRG+CGG+AST+ W+SNLIV+QSFLSLTQ IGT+WTFM+F + + A
Sbjct: 17 VPWVVNSEIYPLRYRGICGGMASTSNWVSNLIVAQSFLSLTQAIGTSWTFMIFIFITIAA 196
Query: 529 IFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQKRDSGSEK 570
I FVI+FVPETKG+PMEEVEKMLE R L FKFWQ+ +E+
Sbjct: 197 IIFVIIFVPETKGLPMEEVEKMLEGRDLNFKFWQRSSRHAEE 322
>AW458960
Length = 285
Score = 153 bits (386), Expect = 2e-37
Identities = 69/94 (73%), Positives = 77/94 (81%)
Frame = +2
Query: 335 TGRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAIETSHYNNTCPDFRAAVNPGRWT 394
TGRKKL L SLCGVV SL +LTV F +S H+P VSA+ETSH+NNTCPD+ +AVNPG W
Sbjct: 2 TGRKKLVLFSLCGVVFSLVVLTVVFHQSTTHSPMVSALETSHFNNTCPDYHSAVNPGGWD 181
Query: 395 CMTCLKASPSCGFCAASPNKLLPGACLISDDTTK 428
CM CLKASP CGFCA+ NKLLPGACLIS DTTK
Sbjct: 182 CMKCLKASPGCGFCASGANKLLPGACLISGDTTK 283
>TC218649 similar to UP|Q39416 (Q39416) Integral membrane protein, partial
(93%)
Length = 1695
Score = 148 bits (374), Expect = 6e-36
Identities = 101/326 (30%), Positives = 164/326 (49%), Gaps = 1/326 (0%)
Frame = +2
Query: 35 IGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFG 94
+G + FG+ G S I +D + L S + GA++GA G I + G
Sbjct: 200 LGPIQFGFTAGYTSPTQSAIINDLGLSVSEFSL---FGSLSNVGAMVGAIASGQIAEYIG 370
Query: 95 RRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYISEASPTRVRG 154
R+ +++IA +IG + ++ A + + L GR+ GFGVG+ S P+YI+E SP +RG
Sbjct: 371 RKGSLMIASIPNIIGWLAISFAKDSSFLYMGRLLEGFGVGIISYTVPVYIAEISPPNLRG 550
Query: 155 ALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLY 214
LVS+N +T G L+YL+ + WR + + P I I + +PESPRWL
Sbjct: 551 GLVSVNQLSVTIGIMLAYLLGIFV-----EWRILAIIGILPCTILIPALFFIPESPRWLA 715
Query: 215 RKGREEESKSILKKIYA-PEDVDAEIEALKESVESEIEESKTSKISMMTLLKTTTVRRGL 273
+ G EE ++ L+ + D+ E+ +K +V S + T LK L
Sbjct: 716 KMGMTEEFETSLQVLRGFDTDISVEVNEIKRAVAS----TNTRITVRFADLKQRRYWLPL 883
Query: 274 YAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFID 333
G+GL QQ GIN V++YS TI + AG +S+ A + + + L+++ D
Sbjct: 884 MIGIGLLILQQLSGINGVLFYSSTIFRNAGISSSDAA---TFGVGAVQVLATSLTLWLAD 1054
Query: 334 KTGRKKLALISLCGVVLSLALLTVAF 359
K+GR+ L ++S G+ SL ++ + F
Sbjct: 1055KSGRRLLLIVSATGMSFSLLVVAITF 1132
Score = 76.3 bits (186), Expect = 4e-14
Identities = 36/102 (35%), Positives = 62/102 (60%), Gaps = 4/102 (3%)
Frame = +2
Query: 451 ALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQ 510
+LVG+ +I FS GMG +PW+I SEI P+ +G+ G +A+ W+ + +V +LT
Sbjct: 1190 SLVGVVAMVIAFSLGMGAMPWIIMSEILPINIKGLAGSVATLANWLFSWLV-----TLTA 1354
Query: 511 TIGTAW----TFMMFGILAVVAIFFVIVFVPETKGVPMEEVE 548
+ W TF ++ ++ + + FV ++VPETKG +EE++
Sbjct: 1355 NMLLDWSSGGTFTIYAVVCALTVVFVTIWVPETKGKTIEEIQ 1480
>BU577019
Length = 409
Score = 145 bits (366), Expect = 5e-35
Identities = 72/80 (90%), Positives = 74/80 (92%)
Frame = +3
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
MEGG E D+SAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFK
Sbjct: 168 MEGGGVEVDVSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKE 347
Query: 61 VDRKTWLQEAIVSTAIAGAI 80
VD KTWLQEAIVS A+AGAI
Sbjct: 348 VDSKTWLQEAIVSMALAGAI 407
>TC208755 similar to UP|HXT4_YEAST (P32467) Low-affinity glucose transporter
HXT4 (Low-affinity glucose transporter LGT1), partial
(3%)
Length = 701
Score = 143 bits (361), Expect = 2e-34
Identities = 62/109 (56%), Positives = 86/109 (78%)
Frame = +3
Query: 447 FGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFL 506
+GW A++GLALYI FSPGMG VPW +NSE+YP YRG+CGG+++T W+SNLIV QSFL
Sbjct: 18 YGWLAILGLALYIAFFSPGMGPVPWTVNSEVYPEEYRGICGGMSATVNWVSNLIVVQSFL 197
Query: 507 SLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRS 555
S+ +GT TF++ I+AV+A FV+V+VPETKG+ +EVE + ++R+
Sbjct: 198 SVAAAVGTGPTFLIIAIIAVLAFMFVVVYVPETKGLTFDEVELLWKERA 344
>TC229858 weakly similar to UP|Q8VHD6 (Q8VHD6) Glucose transporter 10 (Mus
musculus 13 days embryo heart cDNA, RIKEN full-length
enriched library, clone:D330001H08 product:solute
carrier family 2 (facilitated glucose transporter),
member 10, full insert sequence), partial (4%)
Length = 715
Score = 143 bits (360), Expect = 2e-34
Identities = 66/121 (54%), Positives = 92/121 (75%), Gaps = 2/121 (1%)
Frame = +3
Query: 437 AWYTRGCPSK--FGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTV 494
A+Y + S +GW A+VGLALYI FSPGMG VPW ++SEIYP YRG+CGG+++T
Sbjct: 273 AFYKQSSTSNELYGWLAVVGLALYIGFFSPGMGPVPWTLSSEIYPEEYRGICGGMSATVC 452
Query: 495 WISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQR 554
W+SNLIVS++FLS+ + IG TF++ G++AVVA FV+V+VPETKG+ +EVE + +R
Sbjct: 453 WVSNLIVSETFLSIAEGIGIGSTFLIIGVIAVVAFVFVLVYVPETKGLTFDEVEVIWRER 632
Query: 555 S 555
+
Sbjct: 633 A 635
Score = 114 bits (286), Expect = 9e-26
Identities = 59/91 (64%), Positives = 68/91 (73%)
Frame = +3
Query: 269 VRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILS 328
+R G GL FQQF GINTVMYYSPTIVQ+AGF +N ALLLSLI +G+NA G+IL
Sbjct: 6 IRLAFLVGAGLLAFQQFTGINTVMYYSPTIVQMAGFHANELALLLSLIVAGMNAAGTILG 185
Query: 329 IYFIDKTGRKKLALISLCGVVLSLALLTVAF 359
IY ID GRKKLAL SL GV++SL +L AF
Sbjct: 186 IYLIDHAGRKKLALSSLGGVIVSLVILAFAF 278
>TC216311 UP|Q7XA52 (Q7XA52) Monosaccharide transporter, complete
Length = 1925
Score = 129 bits (323), Expect = 5e-30
Identities = 105/361 (29%), Positives = 174/361 (48%), Gaps = 30/361 (8%)
Frame = +1
Query: 24 PYVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDDFKAVDRKTWLQEAI--------- 71
P+V A +GGL+FGYD G+ G ++ F +V RK + +
Sbjct: 157 PFVTVTCIVAAMGGLIFGYDIGISGGVTSMDPFLLKFFPSVFRKKNSDKTVNQYCQYDSQ 336
Query: 72 -----VSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGR 126
S+ A++ + V + RFGR+++++ LFL+G++I A + +L+ GR
Sbjct: 337 TLTMFTSSLYLAALLSSLVASTVTRRFGRKLSMLFGGLLFLVGALINGFAQHVWMLIVGR 516
Query: 127 VFVGFGVGMASMAS---PLYISEASPTRVRGALV--SLNSFLITGGQFLSYLINLAFTNA 181
+ +GFG+G A+ P+ R G L + F I GQ + L+ L A
Sbjct: 517 ILLGFGIGFANQVCATLPI*NGFIQI*RSIGTLAFNCQSQFGIPRGQCVELLLWLKSMVA 696
Query: 182 PGTWRWMLGV---AAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAE 238
W W + V A PA+I V L LP++P + +G E++K+ L++I ++VD E
Sbjct: 697 ---WGWKIEVWEGAMVPALIITVGSLVLPDTPNSMIERGDREKAKAQLQRIRGIDNVDEE 867
Query: 239 IEALKESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTI 298
L + ES S + LL+ R L + + FFQQ GIN +M+Y+P +
Sbjct: 868 FNDLVAASES----SSQVEHPWRNLLQRK-YRPHLTMAVLIPFFQQLTGINVIMFYAPVL 1032
Query: 299 VQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLAL-----ISLCGVVLSLA 353
GF + AL+ ++IT +N + +SIY +DK GR+ L L + +C V++ A
Sbjct: 1033FSSIGFKDD-AALMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGVQMLICQAVVAAA 1209
Query: 354 L 354
+
Sbjct: 1210I 1212
Score = 64.7 bits (156), Expect = 1e-10
Identities = 32/125 (25%), Positives = 64/125 (50%)
Frame = +1
Query: 444 PSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQ 503
P + ++ + +Y+ F+ G + W++ SEI+PL R I + + +++Q
Sbjct: 1252 PKWYAIVVVLFICIYVSAFAWSWGPLGWLVPSEIFPLEIRSAAQSINVSVNMLFTFLIAQ 1431
Query: 504 SFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQK 563
FL++ + F+ F ++ FFV F+PETKG+P+EE+ ++ + +F +
Sbjct: 1432 VFLTMLCHMKFG-LFLFFAFFVLIMTFFVYFFLPETKGIPIEEMGQVWQAHPFWSRFVEH 1608
Query: 564 RDSGS 568
D G+
Sbjct: 1609 DDYGN 1623
>TC206001 similar to UP|Q8VZI5 (Q8VZI5) AT5g18840/F17K4_90, partial (66%)
Length = 951
Score = 110 bits (274), Expect = 2e-24
Identities = 76/240 (31%), Positives = 118/240 (48%), Gaps = 5/240 (2%)
Frame = +3
Query: 125 GRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGT 184
GR F G+G+G+ S P+YI+E +P +RG L + N LI G +S+L+
Sbjct: 48 GRFFTGYGIGVISYVVPVYIAEIAPKNLRGGLATTNQLLIVTGGSVSFLLGSVI-----N 212
Query: 185 WRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAP----EDVDAEIE 240
WR + P I +V + +PESPRWL + GRE+E + L ++ D AEI
Sbjct: 213 WRELALAGLVPCICLLVGLCFIPESPRWLAKVGREKEFQLALSRLRGKHADISDEAAEIL 392
Query: 241 ALKESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQ 300
E++ES K ++ L ++ V + G+GL QQ VGIN + +Y+ I
Sbjct: 393 DYIETLES------LPKTKLLDLFQSKYV-HSVVIGVGLMACQQSVGINGIGFYTAEIFV 551
Query: 301 LAGFASNRT-ALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAF 359
AG +S + + + I G+IL +DK+GR+ L ++S G L + AF
Sbjct: 552 AAGLSSGKAGTIAYACIQIPFTLLGAIL----MDKSGRRPLVMVSAAGTFLGCFVAAFAF 719
Score = 56.6 bits (135), Expect = 3e-08
Identities = 23/55 (41%), Positives = 34/55 (61%)
Frame = +3
Query: 451 ALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSF 505
A G+ +YI FS G+G+VPWVI SEI+P+ +G G + W+ +VS +F
Sbjct: 765 AFAGVLIYIAAFSIGLGSVPWVIMSEIFPIHLKGTAGSLVVLVAWLGAWVVSYTF 929
>AW620575
Length = 377
Score = 102 bits (254), Expect = 5e-22
Identities = 52/86 (60%), Positives = 60/86 (69%)
Frame = +2
Query: 6 PEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKT 65
PE +SAF+ NPY++ A IGGLLFGYDTGVISGALLYI+DDF V
Sbjct: 143 PERKVSAFQ--------NPYIMGFTAVASIGGLLFGYDTGVISGALLYIKDDFPEVRHSN 298
Query: 66 WLQEAIVSTAIAGAIIGAAVGGWIND 91
+LQE IVS A+ GAI+GAA GGWIND
Sbjct: 299 FLQETIVSMAVTGAIVGAAAGGWIND 376
>TC226707 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 2338
Score = 101 bits (252), Expect = 8e-22
Identities = 65/201 (32%), Positives = 103/201 (50%), Gaps = 15/201 (7%)
Frame = +3
Query: 170 LSYLINLAFTNAPGT--WRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILK 227
+ Y+ N F+ WR MLGV A P+I+ V +L++PESPRWL KGR E+K +L
Sbjct: 9 IGYISNYGFSKLALRLGWRLMLGVGAIPSILIGVAVLAMPESPRWLVAKGRLGEAKRVLY 188
Query: 228 KIYAPEDVDAEIEALKESVESEIEESKTSKISMMT-------------LLKTTTVRRGLY 274
KI E+ +A + + I + + +++ L T VR
Sbjct: 189 KISESEE-EARLRLADIKDTAGIPQDCDDDVVLVSKQTHGHGVWRELFLHPTPAVRHIFI 365
Query: 275 AGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDK 334
A +G+ FF Q GI+ V+ YSP I + AG S+ LL ++ + +++ +F+D+
Sbjct: 366 ASLGIHFFAQATGIDAVVLYSPRIFEKAGIKSDNYRLLATVAVGFVKTVSILVATFFLDR 545
Query: 335 TGRKKLALISLCGVVLSLALL 355
GR+ L L S+ G++LSL L
Sbjct: 546 AGRRVLLLCSVSGLILSLLTL 608
Score = 71.6 bits (174), Expect = 9e-13
Identities = 38/104 (36%), Positives = 59/104 (56%), Gaps = 3/104 (2%)
Frame = +3
Query: 449 WAALVGLAL---YIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSF 505
WA + +A Y+ TFS G G + WV +SEI+PLR R I + +++ +++ +F
Sbjct: 654 WAVGLSIAAVLSYVATFSIGSGPITWVYSSEIFPLRLRAQGVAIGAAVNRVTSGVIAMTF 833
Query: 506 LSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEK 549
LSL + I F +F +A VA F +PET+G +EE+EK
Sbjct: 834 LSLQKAITIGGAFFLFAGVAAVAWIFHYTLLPETRGKTLEEIEK 965
>TC229298 weakly similar to UP|Q8MKK4 (Q8MKK4) AT19440p (CG8234-PA), partial
(6%)
Length = 455
Score = 98.6 bits (244), Expect = 7e-21
Identities = 46/52 (88%), Positives = 51/52 (97%)
Frame = +2
Query: 520 MFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQKRDSGSEKH 571
+FGI+A+VAIFFVIVFVPETKGV MEEVEKMLEQRS+QFKFW+KRDSGSEKH
Sbjct: 2 LFGIVAIVAIFFVIVFVPETKGVSMEEVEKMLEQRSVQFKFWEKRDSGSEKH 157
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.138 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,183,458
Number of Sequences: 63676
Number of extensions: 384645
Number of successful extensions: 2063
Number of sequences better than 10.0: 128
Number of HSP's better than 10.0 without gapping: 1965
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2012
length of query: 571
length of database: 12,639,632
effective HSP length: 102
effective length of query: 469
effective length of database: 6,144,680
effective search space: 2881854920
effective search space used: 2881854920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0140.7


