

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.2
(123 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
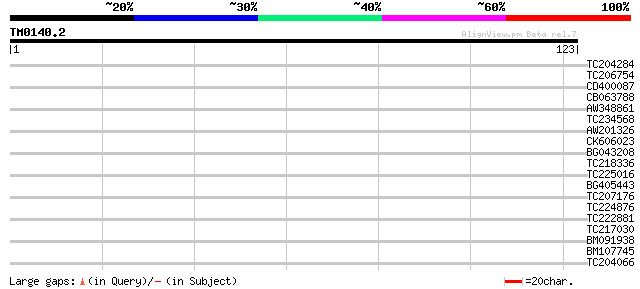
Score E
Sequences producing significant alignments: (bits) Value
TC204284 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, ... 31 0.16
TC206754 weakly similar to UP|GUN4_ARATH (Q9LX31) Tetrapyrrole-b... 30 0.28
CD400087 30 0.28
CB063788 similar to PIR|T00795|T00 26S proteasome regulatory sub... 30 0.36
AW348861 similar to SP|Q9UMN6|TRX2 Trithorax homolog 2 (Mixed li... 29 0.47
TC234568 similar to UP|Q9SML5 (Q9SML5) Knolle, partial (33%) 29 0.47
AW201326 29 0.62
CK606023 28 1.1
BG043208 similar to GP|11139266|gb| PRLI-interacting factor K {A... 27 1.8
TC218336 similar to UP|Q43085 (Q43085) Phosphoribosylanthranilat... 27 1.8
TC225016 similar to UP|Q39835 (Q39835) Extensin, partial (32%) 27 2.4
BG405443 27 2.4
TC207176 27 2.4
TC224876 weakly similar to UP|Q39835 (Q39835) Extensin, partial ... 27 2.4
TC222881 similar to UP|Q9LMT6 (Q9LMT6) F2H15.16, partial (61%) 27 2.4
TC217030 similar to UP|Q9FQF3 (Q9FQF3) Glutathione S-transferase... 27 2.4
BM091938 27 2.4
BM107745 similar to GP|16797791|gb| bZIP transcription factor {N... 27 3.1
TC204066 similar to UP|Q9S9A4 (Q9S9A4) Extensin (Fragment), part... 25 8.9
>TC204284 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, partial
(60%)
Length = 1202
Score = 30.8 bits (68), Expect = 0.16
Identities = 17/68 (25%), Positives = 30/68 (44%), Gaps = 1/68 (1%)
Frame = +1
Query: 33 HLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHHHSCKPTVN-RAPGWFPEKRGVFPEPPK 91
H QP+L C P + + + PP+ H + + + + R P V P PP+
Sbjct: 97 HANQPLLPCTAAPTRPSSAAPVTLRFMPPTSSHRATRASCSARCASRRPRM*PVKPTPPR 276
Query: 92 TALHNVTL 99
+AL + +
Sbjct: 277 SALPAIAI 300
>TC206754 weakly similar to UP|GUN4_ARATH (Q9LX31) Tetrapyrrole-binding
protein, chloroplast precursor (Genomes uncoulped 4),
partial (62%)
Length = 1115
Score = 30.0 bits (66), Expect = 0.28
Identities = 18/61 (29%), Positives = 25/61 (40%), Gaps = 2/61 (3%)
Frame = +3
Query: 62 SQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTALHNVTLLQHLRNTT--QSDIHRTGSSDT 119
S HH C T P P +F +PP +H+ L H + + I +T S T
Sbjct: 75 SIQHHQCSLTKRHHPSDTPPPTSLFLKPPTPPIHHNITLSHTPSNSYITYSISQTTPSST 254
Query: 120 T 120
T
Sbjct: 255 T 257
>CD400087
Length = 576
Score = 30.0 bits (66), Expect = 0.28
Identities = 13/36 (36%), Positives = 17/36 (47%)
Frame = -1
Query: 48 PRTPCVIMIASTPPSQHHHSCKPTVNRAPGWFPEKR 83
PR PC A++PP QH C +P W +R
Sbjct: 456 PRAPCPA*AAASPPMQHGSPCGALRAGSPRWRSRRR 349
>CB063788 similar to PIR|T00795|T00 26S proteasome regulatory subunit
[imported] - Arabidopsis thaliana, partial (4%)
Length = 397
Score = 29.6 bits (65), Expect = 0.36
Identities = 24/80 (30%), Positives = 32/80 (40%), Gaps = 2/80 (2%)
Frame = -1
Query: 25 RTLSPHILHLYQPMLVCHLDNLMPRT--PCVIMIASTPPSQHHHSCKPTVNRAPGWFPEK 82
+ L +L YQP L H N+ PR+ C+I P +HS P P + P
Sbjct: 280 QALMSSLLLGYQPSLCPHAKNIQPRSIRDCLISRIKPQPFYKNHSYPP----PPLYIPSL 113
Query: 83 RGVFPEPPKTALHNVTLLQH 102
P H +TLL H
Sbjct: 112 GPALSLHPS---HRITLLFH 62
>AW348861 similar to SP|Q9UMN6|TRX2 Trithorax homolog 2 (Mixed lineage
leukemia gene homolog 2 protein). [Human] {Homo
sapiens}, partial (1%)
Length = 642
Score = 29.3 bits (64), Expect = 0.47
Identities = 22/74 (29%), Positives = 26/74 (34%), Gaps = 6/74 (8%)
Frame = +1
Query: 46 LMPRTPCVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPK------TALHNVTL 99
L+P TP I +TPP +HHH P + P P P T H T
Sbjct: 40 LIPGTP----IHTTPPLRHHHH*HPQATKQQAQSPPSGAT*PPSPSRTP*PPTPCHPATR 207
Query: 100 LQHLRNTTQSDIHR 113
R T S R
Sbjct: 208 ASTARATNPSSAPR 249
>TC234568 similar to UP|Q9SML5 (Q9SML5) Knolle, partial (33%)
Length = 446
Score = 29.3 bits (64), Expect = 0.47
Identities = 14/40 (35%), Positives = 19/40 (47%)
Frame = -3
Query: 50 TPCVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEP 89
TP + S+PP HH+C PE + +FPEP
Sbjct: 99 TPGAAPVDSSPPPWQHHAC-----------PESQPLFPEP 13
>AW201326
Length = 520
Score = 28.9 bits (63), Expect = 0.62
Identities = 14/48 (29%), Positives = 18/48 (37%), Gaps = 8/48 (16%)
Frame = +3
Query: 65 HHSCKPTV--------NRAPGWFPEKRGVFPEPPKTALHNVTLLQHLR 104
H CKP + N P W PE P PP + + + LR
Sbjct: 78 HTQCKPKLSFLRRNLKNVPPDWLPESTDALPPPPNCSYTDAAITNTLR 221
>CK606023
Length = 538
Score = 28.1 bits (61), Expect = 1.1
Identities = 17/42 (40%), Positives = 20/42 (47%), Gaps = 6/42 (14%)
Frame = +3
Query: 60 PPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPK------TALH 95
PPSQHH+ P+ P P GV P P + TALH
Sbjct: 141 PPSQHHNQALPSGRGPP---PPLNGVLPLPVQK*TWSLTALH 257
>BG043208 similar to GP|11139266|gb| PRLI-interacting factor K {Arabidopsis
thaliana}, partial (4%)
Length = 454
Score = 27.3 bits (59), Expect = 1.8
Identities = 12/37 (32%), Positives = 17/37 (45%)
Frame = -1
Query: 41 CHLDNLMPRTPCVIMIASTPPSQHHHSCKPTVNRAPG 77
C LD+L P +++ P H +C T N PG
Sbjct: 328 CQLDSLSPPVTVNVVLGLFNPKNHSTACMLTDNIVPG 218
>TC218336 similar to UP|Q43085 (Q43085) Phosphoribosylanthranilate
transferase (Fragment) , partial (89%)
Length = 1156
Score = 27.3 bits (59), Expect = 1.8
Identities = 17/73 (23%), Positives = 33/73 (44%)
Frame = +3
Query: 1 MLAFSGSSSAIDITTYLASMSTNARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTP 60
M FSG + + +L +ST ++ ++H+ MLVC + ++P + + S
Sbjct: 390 MTVFSG---ILSVVRWLGEVSTWKHPITTVLVHILFLMLVCFPELILPTVFLYMFVISMW 560
Query: 61 PSQHHHSCKPTVN 73
+ C P +N
Sbjct: 561 NWRFRPRCPPHMN 599
>TC225016 similar to UP|Q39835 (Q39835) Extensin, partial (32%)
Length = 973
Score = 26.9 bits (58), Expect = 2.4
Identities = 14/35 (40%), Positives = 18/35 (51%), Gaps = 2/35 (5%)
Frame = +2
Query: 58 STPPSQHH--HSCKPTVNRAPGWFPEKRGVFPEPP 90
S PP HH H P V+ +P P+K +P PP
Sbjct: 395 SPPPPVHHYPHPHPPFVHYSPPPSPKKPYKYPSPP 499
>BG405443
Length = 339
Score = 26.9 bits (58), Expect = 2.4
Identities = 27/95 (28%), Positives = 42/95 (43%), Gaps = 8/95 (8%)
Frame = +1
Query: 36 QPMLVCHLDNLMPRTPCVI--MIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTA 93
Q +L HL P+TP + ++ SQHH S P ++R GW + PP+
Sbjct: 46 QELLGVHLPLC*PQTPPISEPWFRNSLASQHHPS-HPRLSREQGW------ISLLPPQHQ 204
Query: 94 LHNVTL------LQHLRNTTQSDIHRTGSSDTTVR 122
L + TL LQH T + +H+ + + R
Sbjct: 205 L*DPTLTLTLSTLQHNLLTFFAPLHKNSNFEVFTR 309
>TC207176
Length = 1616
Score = 26.9 bits (58), Expect = 2.4
Identities = 10/29 (34%), Positives = 16/29 (54%)
Frame = -3
Query: 50 TPCVIMIASTPPSQHHHSCKPTVNRAPGW 78
+P + + A +PP HH C+ +R GW
Sbjct: 966 SPSLSLTALSPPVAHHSRCRQ*GHRQRGW 880
>TC224876 weakly similar to UP|Q39835 (Q39835) Extensin, partial (66%)
Length = 1340
Score = 26.9 bits (58), Expect = 2.4
Identities = 14/35 (40%), Positives = 18/35 (51%), Gaps = 2/35 (5%)
Frame = +3
Query: 58 STPPSQHH--HSCKPTVNRAPGWFPEKRGVFPEPP 90
S PP HH H P V+ +P P+K +P PP
Sbjct: 987 SPPPPVHHYPHPHPPFVHYSPPPSPKKPYKYPSPP 1091
>TC222881 similar to UP|Q9LMT6 (Q9LMT6) F2H15.16, partial (61%)
Length = 739
Score = 26.9 bits (58), Expect = 2.4
Identities = 11/24 (45%), Positives = 16/24 (65%)
Frame = -2
Query: 55 MIASTPPSQHHHSCKPTVNRAPGW 78
+I S P+ HH +CKPT+N G+
Sbjct: 732 VILSLLPAFHHLTCKPTLNCKSGF 661
>TC217030 similar to UP|Q9FQF3 (Q9FQF3) Glutathione S-transferase GST 5 ,
partial (63%)
Length = 614
Score = 26.9 bits (58), Expect = 2.4
Identities = 14/31 (45%), Positives = 19/31 (61%)
Frame = -2
Query: 4 FSGSSSAIDITTYLASMSTNARTLSPHILHL 34
FS +SSA ++ AS S+ ARTL + HL
Sbjct: 187 FSSTSSACSKCSFFASSSSVARTLFKDVSHL 95
>BM091938
Length = 421
Score = 26.9 bits (58), Expect = 2.4
Identities = 14/35 (40%), Positives = 18/35 (51%), Gaps = 2/35 (5%)
Frame = +2
Query: 58 STPPSQHH--HSCKPTVNRAPGWFPEKRGVFPEPP 90
S PP HH H P V+ +P P+K +P PP
Sbjct: 269 SPPPPVHHYPHPHPPFVHYSPPPSPKKPYKYPSPP 373
>BM107745 similar to GP|16797791|gb| bZIP transcription factor {Nicotiana
tabacum}, partial (10%)
Length = 569
Score = 26.6 bits (57), Expect = 3.1
Identities = 11/40 (27%), Positives = 18/40 (44%)
Frame = -1
Query: 39 LVCHLDNLMPRTPCVIMIASTPPSQHHHSCKPTVNRAPGW 78
L C+L+ P P +A P S+ +C P + G+
Sbjct: 362 LFCNLEGSFPEGPDPFKVACNPSSEELDACGPLSEKVAGF 243
>TC204066 similar to UP|Q9S9A4 (Q9S9A4) Extensin (Fragment), partial (64%)
Length = 631
Score = 25.0 bits (53), Expect = 8.9
Identities = 14/46 (30%), Positives = 20/46 (43%)
Frame = +3
Query: 22 TNARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHHHS 67
T+ + +PH HL P HL + T + ST HHH+
Sbjct: 15 THXQCSTPHHQHLRSPTNTHHLRHQFTPTLPMFPPQSTTLLHHHHT 152
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.131 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,096,380
Number of Sequences: 63676
Number of extensions: 130773
Number of successful extensions: 998
Number of sequences better than 10.0: 83
Number of HSP's better than 10.0 without gapping: 978
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 996
length of query: 123
length of database: 12,639,632
effective HSP length: 85
effective length of query: 38
effective length of database: 7,227,172
effective search space: 274632536
effective search space used: 274632536
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0140.2


