

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
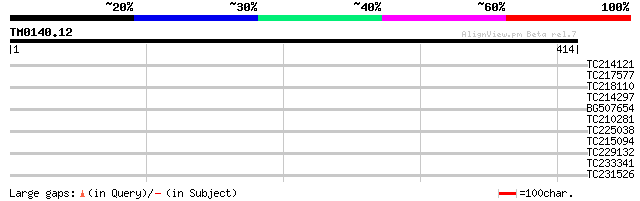
Query= TM0140.12
(414 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC214121 similar to UP|Q8S2S7 (Q8S2S7) Ethylene responsive eleme... 35 0.081
TC217577 similar to UP|Q9ASS7 (Q9ASS7) AT5g36940/MLF18_60, parti... 34 0.14
TC218110 homologue to UP|SUI1_SALBA (O48650) Protein translation... 33 0.24
TC214297 similar to UP|Q8S2S7 (Q8S2S7) Ethylene responsive eleme... 32 0.52
BG507654 30 1.5
TC210281 weakly similar to UP|Q8GU29 (Q8GU29) Glycin-rich RNA bi... 30 2.6
TC225038 similar to UP|Q41025 (Q41025) Histone H1, partial (67%) 29 3.4
TC215094 weakly similar to UP|Q6RJY7 (Q6RJY7) Elicitor-inducible... 29 4.4
TC229132 weakly similar to UP|Q944E4 (Q944E4) Cellulose synthase... 28 5.8
TC233341 similar to UP|Q8PWH4 (Q8PWH4) Conserved protein, partia... 28 5.8
TC231526 similar to UP|PME_MEDSA (Q42920) Pectinesterase precurs... 28 5.8
>TC214121 similar to UP|Q8S2S7 (Q8S2S7) Ethylene responsive element binding
factor 4-like protein, partial (50%)
Length = 965
Score = 34.7 bits (78), Expect = 0.081
Identities = 26/74 (35%), Positives = 38/74 (51%)
Frame = -1
Query: 63 EVGVSIGLCFFLGNAVAGALYVLGAVETFLKAVPAAGIFRETITQVNGTTIAQPIESPSS 122
EVG G G V GA VLG VE A P AG+ +T + G +A P P +
Sbjct: 278 EVGFGFGAAEVAGGGVIGAGGVLGGVE---GAEPDAGLL-AGVTDLGG--VAAPWSLPDA 117
Query: 123 HDLQIYGIVVTIVL 136
++++ GIVV +++
Sbjct: 116 AEVKLVGIVVWLLV 75
>TC217577 similar to UP|Q9ASS7 (Q9ASS7) AT5g36940/MLF18_60, partial (44%)
Length = 1475
Score = 33.9 bits (76), Expect = 0.14
Identities = 34/164 (20%), Positives = 68/164 (40%), Gaps = 5/164 (3%)
Frame = +3
Query: 229 GIMAGSNRSSSLKDTQRSIPLGTLAATLVTTFMYLVSVIMFGALATREKLLTDRLLTATV 288
G A ++ + +K+ QR +PLG + + +Y++ I+ L + D +++
Sbjct: 9 GFDAVASTAEEVKNPQRDLPLGIGGSLFLCCGLYMLVSIVIVGLVPYYAINPDTPISSAF 188
Query: 289 A-----WPFPSLIKIGIILSTMGAALQSLTGAPRLLAAIANDDILPILKYFKVADGSEPH 343
A W +I G + + + + PR+L A+A D +LP P
Sbjct: 189 ADNGMQWA-AYVINGGAFTALCASLMGGILPQPRILMAMARDGLLPPFFSDINKCSQVPV 365
Query: 344 VATLFTAFLCSGCVVIGNLDLITPTVTMFFLLCYAGVNLSCFLL 387
+T+ T + S + + V++ LL + V +S +L
Sbjct: 366 KSTIVTGLVASLLAFSMEVSELAGMVSVGTLLAFTMVAISVLIL 497
>TC218110 homologue to UP|SUI1_SALBA (O48650) Protein translation factor SUI1
homolog, complete
Length = 876
Score = 33.1 bits (74), Expect = 0.24
Identities = 21/57 (36%), Positives = 29/57 (50%), Gaps = 5/57 (8%)
Frame = +3
Query: 346 TLFTAFLCSGCVVIGNLDLI----TPTVTMFFLLCY-AGVNLSCFLLDLLDAPSWRP 397
T+F L S ++I D + TP + F LL + V L CFL+ LD P+W P
Sbjct: 597 TVFLPILDSYYLIIYGKDNMSASKTPVLNKFVLLFFHVQVTLGCFLIWALDVPNWEP 767
>TC214297 similar to UP|Q8S2S7 (Q8S2S7) Ethylene responsive element binding
factor 4-like protein, partial (51%)
Length = 996
Score = 32.0 bits (71), Expect = 0.52
Identities = 38/126 (30%), Positives = 51/126 (40%), Gaps = 2/126 (1%)
Frame = -2
Query: 18 GGTLLLVALCGTCTFLTAISLSAIATNGAMKGGGPYYLIGRALGPE--VGVSIGLCFFLG 75
GG L AL GT F + + + N +G +G E VG G G
Sbjct: 398 GGGLHGAALAGTVRFNVNVVVLLVEVN-----------VGVVVGEEGEVGFGFGATEVSG 252
Query: 76 NAVAGALYVLGAVETFLKAVPAAGIFRETITQVNGTTIAQPIESPSSHDLQIYGIVVTIV 135
V GA +LG VE A P AG+ + + G IA P P S ++ + I +V
Sbjct: 251 GDVVGAGGILGGVE---GAEPDAGLL-SRVADLGG--IAAPGALPDSAEVNLVCIAWLLV 90
Query: 136 LCFIVF 141
L VF
Sbjct: 89 LVGQVF 72
>BG507654
Length = 411
Score = 30.4 bits (67), Expect = 1.5
Identities = 12/50 (24%), Positives = 26/50 (52%)
Frame = +3
Query: 229 GIMAGSNRSSSLKDTQRSIPLGTLAATLVTTFMYLVSVIMFGALATREKL 278
G + S + +KD +S+P+G + + L+TT +Y + + + K+
Sbjct: 156 GYDSASTMAEEVKDPFKSLPIGIVGSVLITTLLYCLMALSLCMMVPYNKI 305
>TC210281 weakly similar to UP|Q8GU29 (Q8GU29) Glycin-rich RNA binding
protein, partial (16%)
Length = 680
Score = 29.6 bits (65), Expect = 2.6
Identities = 26/75 (34%), Positives = 31/75 (40%), Gaps = 7/75 (9%)
Frame = -3
Query: 17 IGGTLLLVALCGTCTFLTAISLSAIATNGAMKGGGPY-------YLIGRALGPEVGVSIG 69
IGG L +A G C + I +AI T A GG P Y L P V
Sbjct: 309 IGGLELTLAFPGCCIYPGFIGNAAIGTTPAGNGGWPVLPFWESAYFTTIFLPPSVFSRSS 130
Query: 70 LCFFLGNAVAGALYV 84
+ FF +AV LYV
Sbjct: 129 MAFFAASAV---LYV 94
>TC225038 similar to UP|Q41025 (Q41025) Histone H1, partial (67%)
Length = 862
Score = 29.3 bits (64), Expect = 3.4
Identities = 28/91 (30%), Positives = 43/91 (46%), Gaps = 3/91 (3%)
Frame = -2
Query: 14 MGGIGGTLLLVALCGTCTFLTAISLSAIATNGAMKGGGPYYLIGRALGPEVGVSIGLCFF 73
+ G+ T L L G + L+ +A A+ ++L ALG E G+ L F
Sbjct: 600 LAGLAATFLAATLAGL-----DLGLAVVAAALALGLATAFFLGVAALGLEAGLVAALVGF 436
Query: 74 LG-NAVAGALYVLGAVETF--LKAVPAAGIF 101
LG A+ GA V +++ F L ++PAA F
Sbjct: 435 LGAAALTGAGLVGASLKEFFTLTSLPAATDF 343
>TC215094 weakly similar to UP|Q6RJY7 (Q6RJY7) Elicitor-inducible protein
EIG-J7, partial (71%)
Length = 995
Score = 28.9 bits (63), Expect = 4.4
Identities = 18/53 (33%), Positives = 26/53 (48%), Gaps = 6/53 (11%)
Frame = -1
Query: 95 VPAAGIFRETITQVNGTTIAQPIESPSSHDLQIYGIVVTIVL------CFIVF 141
VPA R + VNGT + SPS+ D+++ + V + L CF VF
Sbjct: 371 VPAVVRTRSVRSNVNGTDGSVKTGSPSAEDVEVAALEVVVALAHQAAPCFEVF 213
>TC229132 weakly similar to UP|Q944E4 (Q944E4) Cellulose synthase-like
protein OsCslE1, partial (29%)
Length = 824
Score = 28.5 bits (62), Expect = 5.8
Identities = 18/60 (30%), Positives = 26/60 (43%)
Frame = -1
Query: 293 PSLIKIGIILSTMGAALQSLTGAPRLLAAIANDDILPILKYFKVADGSEPHVATLFTAFL 352
PSL+ ++ST+G L P L I N + PI+ + EP + L A L
Sbjct: 731 PSLVGCSTLISTLGTV*TILLSHPPLDTHILNPSLFPIVCFPHSCSLQEPLIVVLVLAGL 552
>TC233341 similar to UP|Q8PWH4 (Q8PWH4) Conserved protein, partial (7%)
Length = 714
Score = 28.5 bits (62), Expect = 5.8
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = -1
Query: 376 CYAGVNLSCFLLDLLDAPSWRPRWKFHHWSL 406
C++ + S LL LLD+ + R + HHW L
Sbjct: 117 CFSALGFSRHLLPLLDSDFPQQRQRHHHWVL 25
>TC231526 similar to UP|PME_MEDSA (Q42920) Pectinesterase precursor (Pectin
methylesterase) (PE) (P65) , partial (56%)
Length = 966
Score = 28.5 bits (62), Expect = 5.8
Identities = 29/113 (25%), Positives = 48/113 (41%), Gaps = 3/113 (2%)
Frame = +3
Query: 197 YQKTNDAGIPEPDGSVSWNFNALVGLFFPAVTGIMAGSNRSSSLKDTQRS---IPLGTLA 253
Y N +P P ++W + G F P ++ S S K T R +
Sbjct: 411 YSPANPTCLP*PPRLLTWEDH---GGFTPRLSSWTPKSTTSLFPKGTWRGWEVLSRTQAP 581
Query: 254 ATLVTTFMYLVSVIMFGALATREKLLTDRLLTATVAWPFPSLIKIGIILSTMG 306
T TT + ++++++ L+ RLL T A F SLI++ +IL + G
Sbjct: 582 TTSSTTGVLVLTLLVESPGLVLRFLILLRLLNTTQANSFRSLIRLSVILGS*G 740
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.142 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,325,790
Number of Sequences: 63676
Number of extensions: 320132
Number of successful extensions: 2418
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 2403
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2417
length of query: 414
length of database: 12,639,632
effective HSP length: 100
effective length of query: 314
effective length of database: 6,272,032
effective search space: 1969418048
effective search space used: 1969418048
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0140.12


