

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0137.5
(214 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
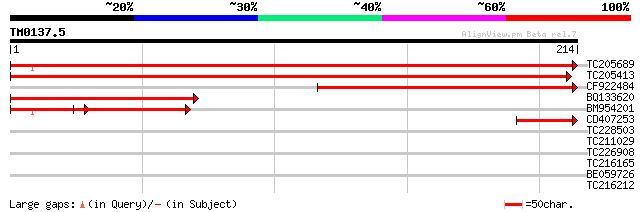
Score E
Sequences producing significant alignments: (bits) Value
TC205689 homologue to UP|Q949G5 (Q949G5) Mob1-like protein, comp... 432 e-122
TC205413 homologue to GB|AAP12863.1|30017259|BT006214 At4g19050 ... 399 e-112
CF922484 171 2e-43
BQ133620 122 9e-29
BM954201 homologue to GP|19309913|emb Mob1-like protein {Medicag... 93 1e-25
CD407253 homologue to GP|19309913|emb Mob1-like protein {Medicag... 45 3e-05
TC228503 similar to PIR|H96770|H96770 protein heat shock protein... 30 0.82
TC211029 similar to UP|Q96VI4 (Q96VI4) Protease 1, partial (3%) 28 3.1
TC226908 similar to UP|Q8RXD9 (Q8RXD9) 4-alpha-glucanotransferas... 27 5.3
TC216165 similar to UP|O24135 (O24135) Citrate synthase, partial... 27 5.3
BE059726 weakly similar to SP|P00870|RBS1_ Ribulose bisphosphate... 27 7.0
TC216212 27 9.1
>TC205689 homologue to UP|Q949G5 (Q949G5) Mob1-like protein, complete
Length = 1001
Score = 432 bits (1111), Expect = e-122
Identities = 210/215 (97%), Positives = 213/215 (98%), Gaps = 1/215 (0%)
Frame = +1
Query: 1 MSLFGLG-RNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAV 59
MSLFGLG RNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAV
Sbjct: 121 MSLFGLGNRNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAV 300
Query: 60 NTVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLM 119
NTVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLM
Sbjct: 301 NTVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLM 480
Query: 120 DWIESQLDDETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHL 179
DWIESQLDDETIFPQ+LGAPFP NFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHL
Sbjct: 481 DWIESQLDDETIFPQRLGAPFPTNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHL 660
Query: 180 NTCFKHFVLFTWEFRLIDKAELAPLEDLVDSIIQL 214
NTCFKHFVLFTWEFRLIDKAELAPLEDLV+SI+QL
Sbjct: 661 NTCFKHFVLFTWEFRLIDKAELAPLEDLVESIVQL 765
>TC205413 homologue to GB|AAP12863.1|30017259|BT006214 At4g19050 {Arabidopsis
thaliana;} , complete
Length = 1073
Score = 399 bits (1026), Expect = e-112
Identities = 186/212 (87%), Positives = 205/212 (95%)
Frame = +2
Query: 1 MSLFGLGRNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVN 60
MSLFG+GRNQ+TFRPKKS PSGSKGAQL+KHIDATLGSGNLREAVKLPPGED+NEWLAVN
Sbjct: 113 MSLFGIGRNQRTFRPKKSTPSGSKGAQLRKHIDATLGSGNLREAVKLPPGEDLNEWLAVN 292
Query: 61 TVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMD 120
TVDFFNQVN+L+GTLTEFCTP NC +M+AGPKYEYRWADGV IKKPIEVSAPKYVEYLMD
Sbjct: 293 TVDFFNQVNLLYGTLTEFCTPENCRTMSAGPKYEYRWADGVQIKKPIEVSAPKYVEYLMD 472
Query: 121 WIESQLDDETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLN 180
WIE+QLDDE+IFPQKLG+PFPPNF++VVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLN
Sbjct: 473 WIEAQLDDESIFPQKLGSPFPPNFKEVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLN 652
Query: 181 TCFKHFVLFTWEFRLIDKAELAPLEDLVDSII 212
TCFKHF+LFT EF LIDK ELAPL++L+++II
Sbjct: 653 TCFKHFILFTCEFGLIDKKELAPLQELIETII 748
>CF922484
Length = 535
Score = 171 bits (433), Expect = 2e-43
Identities = 83/98 (84%), Positives = 86/98 (87%)
Frame = -2
Query: 117 YLMDWIESQLDDETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEE 176
YLMDWIE QLDDETIFPQ+LG PFP NFRDVVKTIFK FRVYA IYHSHFQKIVSLKEE
Sbjct: 531 YLMDWIEFQLDDETIFPQRLGGPFPTNFRDVVKTIFKGFFRVYAPIYHSHFQKIVSLKEE 352
Query: 177 AHLNTCFKHFVLFTWEFRLIDKAELAPLEDLVDSIIQL 214
A LNT FKHFV FTWEFRLIDKAELAP DLV+ I+QL
Sbjct: 351 APLNTWFKHFVFFTWEFRLIDKAELAPFGDLVEFIVQL 238
>BQ133620
Length = 391
Score = 122 bits (307), Expect = 9e-29
Identities = 58/71 (81%), Positives = 66/71 (92%)
Frame = +3
Query: 1 MSLFGLGRNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVN 60
MSLFG+GRNQ+TFRPKKS PSGSKGAQL+KHIDATLGSGNLREAVKLPPGED+NEWLAVN
Sbjct: 6 MSLFGIGRNQRTFRPKKSTPSGSKGAQLRKHIDATLGSGNLREAVKLPPGEDLNEWLAVN 185
Query: 61 TVDFFNQVNIL 71
+ FF+ ++ L
Sbjct: 186SNYFFSVMHCL 218
Score = 30.0 bits (66), Expect = 0.82
Identities = 11/16 (68%), Positives = 14/16 (86%)
Frame = +2
Query: 59 VNTVDFFNQVNILFGT 74
+ VDFFNQVN+L+GT
Sbjct: 344 ITAVDFFNQVNLLYGT 391
>BM954201 homologue to GP|19309913|emb Mob1-like protein {Medicago sativa
subsp. falcata}, partial (30%)
Length = 326
Score = 93.2 bits (230), Expect(2) = 1e-25
Identities = 44/44 (100%), Positives = 44/44 (100%)
Frame = +3
Query: 25 GAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVNTVDFFNQV 68
GAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVNTVDFFNQV
Sbjct: 195 GAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVNTVDFFNQV 326
Score = 40.4 bits (93), Expect(2) = 1e-25
Identities = 22/31 (70%), Positives = 22/31 (70%), Gaps = 1/31 (3%)
Frame = +2
Query: 1 MSLFGLG-RNQKTFRPKKSAPSGSKGAQLQK 30
MSLFGLG RNQKTFRPKKSAP G K
Sbjct: 119 MSLFGLGNRNQKTFRPKKSAPFFFLGCSTSK 211
>CD407253 homologue to GP|19309913|emb Mob1-like protein {Medicago sativa
subsp. falcata}, partial (15%)
Length = 455
Score = 44.7 bits (104), Expect = 3e-05
Identities = 21/23 (91%), Positives = 23/23 (99%)
Frame = -1
Query: 192 EFRLIDKAELAPLEDLVDSIIQL 214
EFRLIDKAELAPLEDLV+SI+QL
Sbjct: 185 EFRLIDKAELAPLEDLVESIVQL 117
Score = 27.3 bits (59), Expect = 5.3
Identities = 10/10 (100%), Positives = 10/10 (100%)
Frame = -3
Query: 180 NTCFKHFVLF 189
NTCFKHFVLF
Sbjct: 447 NTCFKHFVLF 418
>TC228503 similar to PIR|H96770|H96770 protein heat shock protein F1O17.8
[imported] - Arabidopsis thaliana {Arabidopsis thaliana;}
, partial (6%)
Length = 1165
Score = 30.0 bits (66), Expect = 0.82
Identities = 11/18 (61%), Positives = 13/18 (72%)
Frame = -2
Query: 178 HLNTCFKHFVLFTWEFRL 195
H +T FKHF LF WE +L
Sbjct: 1119 HHSTSFKHFCLFVWELKL 1066
>TC211029 similar to UP|Q96VI4 (Q96VI4) Protease 1, partial (3%)
Length = 858
Score = 28.1 bits (61), Expect = 3.1
Identities = 19/45 (42%), Positives = 25/45 (55%), Gaps = 4/45 (8%)
Frame = +3
Query: 105 KPIEVSAPKYVEYLMDWIESQLDDETIFPQ----KLGAPFPPNFR 145
K I V K++ L+D IE +LD E P +LG P PP+FR
Sbjct: 609 KFIVVGGVKFIGNLVD-IEDRLDVEIYDPLVGSWELGPPLPPDFR 740
>TC226908 similar to UP|Q8RXD9 (Q8RXD9) 4-alpha-glucanotransferase
(At2g40840), partial (60%)
Length = 2257
Score = 27.3 bits (59), Expect = 5.3
Identities = 17/59 (28%), Positives = 29/59 (48%), Gaps = 1/59 (1%)
Frame = +3
Query: 145 RDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNT-CFKHFVLFTWEFRLIDKAELA 202
RD +T + + +AH +K+VS K+ H CF ++V + +L + AE A
Sbjct: 195 RDFFETSDRTQWGCFAHYSEDKLEKLVS-KDSLHYEIICFHYYVQYHLHLQLSEAAEYA 368
>TC216165 similar to UP|O24135 (O24135) Citrate synthase, partial (70%)
Length = 1419
Score = 27.3 bits (59), Expect = 5.3
Identities = 27/115 (23%), Positives = 46/115 (39%), Gaps = 6/115 (5%)
Frame = +1
Query: 81 PSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFPQKLGAP- 139
P+NC A P Y Y+ D + VSA ++ + Q+ E + G
Sbjct: 91 PTNCEDEQAIPDYAYKAIDA------LPVSAHAMTQFTTGVMALQVQSEFQKAYEGGITK 252
Query: 140 ---FPPNFRDVVKTIFKRLFRVYAHIYHSHFQ--KIVSLKEEAHLNTCFKHFVLF 189
+ P + D + I RL + A+IY ++ KI+ L + + H + F
Sbjct: 253 ARYWEPTYEDTMNLI-ARLPAIAAYIYRRKYKDGKIIPLDDSLDYGANYAHMLGF 414
>BE059726 weakly similar to SP|P00870|RBS1_ Ribulose bisphosphate carboxylase
small chain (EC 4.1.1.39) (RuBisCO small subunit).
[Spinach], partial (39%)
Length = 262
Score = 26.9 bits (58), Expect = 7.0
Identities = 12/32 (37%), Positives = 19/32 (58%)
Frame = -1
Query: 141 PPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVS 172
PP V+KT+F F+ ++ +HSH I+S
Sbjct: 187 PPVVLPVLKTVFLTEFQAWSPSFHSHVLVILS 92
>TC216212
Length = 731
Score = 26.6 bits (57), Expect = 9.1
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = -1
Query: 118 LMDWIESQLDDETIFPQKLGAPFPPN 143
+ DW E D+++ F Q L +PFPPN
Sbjct: 542 IYDWSE---DNQSPFSQTLLSPFPPN 474
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.139 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,894,449
Number of Sequences: 63676
Number of extensions: 137305
Number of successful extensions: 702
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 700
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 701
length of query: 214
length of database: 12,639,632
effective HSP length: 93
effective length of query: 121
effective length of database: 6,717,764
effective search space: 812849444
effective search space used: 812849444
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0137.5


