

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.7
(841 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
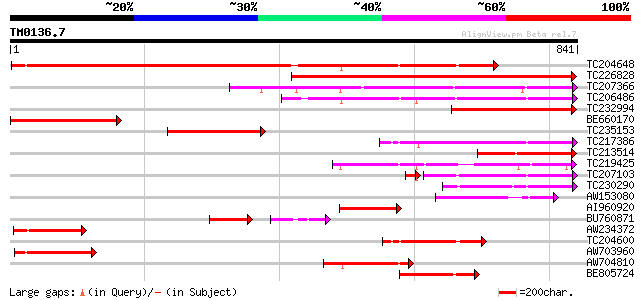
Score E
Sequences producing significant alignments: (bits) Value
TC204648 homologue to UP|Q9M5J4 (Q9M5J4) Beta galactosidase, com... 868 0.0
TC226828 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , par... 692 0.0
TC207366 similar to UP|Q94BZ3 (Q94BZ3) At2g32810/F24L7.5, partia... 441 e-124
TC206486 similar to UP|O65761 (O65761) Beta-galactosidase (Frag... 371 e-103
TC232994 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , par... 319 4e-87
BE660170 310 1e-84
TC235153 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , par... 262 4e-70
TC217386 similar to GB|AAF70823.1|7939621|AF154422 beta-galactos... 257 2e-68
TC213514 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , par... 253 2e-67
TC219425 224 1e-58
TC207103 homologue to UP|O65761 (O65761) Beta-galactosidase (Fr... 214 7e-56
TC230290 weakly similar to UP|Q93X58 (Q93X58) Beta-galactosidase... 176 3e-44
AW153080 weakly similar to GP|14517399|gb| At2g32810/F24L7.5 {Ar... 155 9e-38
AI960920 similar to GP|14970841|emb beta-galactosidase {Fragaria... 155 9e-38
BU760871 homologue to GP|3204134|emb beta-galactosidase {Cicer a... 108 5e-37
AW234372 homologue to GP|7682680|gb| beta galactosidase {Vigna r... 151 1e-36
TC204600 homologue to UP|Q9M5J3 (Q9M5J3) Beta galactosidase, par... 151 1e-36
AW703960 similar to GP|7682677|gb| beta galactosidase {Vigna rad... 140 2e-33
AW704810 similar to GP|7682677|gb| beta galactosidase {Vigna rad... 134 1e-31
BE805724 122 5e-28
>TC204648 homologue to UP|Q9M5J4 (Q9M5J4) Beta galactosidase, complete
Length = 2817
Score = 868 bits (2243), Expect = 0.0
Identities = 426/730 (58%), Positives = 513/730 (69%), Gaps = 8/730 (1%)
Frame = +3
Query: 3 MRRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDL 62
M + +F ++L +LC++ A+VTYDH+A+V+DGKRR+LISGSIHYPRSTP+MWPDL
Sbjct: 72 MGKREFNGVVLMMLCLWVCGV-TASVTYDHKAIVVDGKRRILISGSIHYPRSTPQMWPDL 248
Query: 63 IQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACA 122
IQKAKDGGLDVI+TYVFWN HEP GQY FE R DLVKFVK AGLYVHLRIGPY CA
Sbjct: 249 IQKAKDGGLDVIQTYVFWNGHEPSPGQYYFEDRFDLVKFVKLAQQAGLYVHLRIGPYICA 428
Query: 123 EWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIE 182
EWN GGFP+WL ++PGI FRTDNEPFKA MQ+FTAKIV +MK+ L+ SQGGPIILSQIE
Sbjct: 429 EWNLGGFPVWLKYVPGIAFRTDNEPFKAAMQKFTAKIVSLMKENRLFQSQGGPIILSQIE 608
Query: 183 NEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTP 242
NEYG VE G Y WAA MA LDTGVPWVMC+QE+APDP+I+TCNGFYC+ F P
Sbjct: 609 NEYGPVEWEIGAPGKAYTKWAAQMAVGLDTGVPWVMCKQEDAPDPVIDTCNGFYCENFKP 788
Query: 243 NSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGR 302
N KPK WTE + GW FGGAVP RP EDLAF+VARF Q GG+ NYYMYHGGTNFGR
Sbjct: 789 NKNTKPKMWTENWTGWYTDFGGAVPRRPAEDLAFSVARFIQNGGSFVNYYMYHGGTNFGR 968
Query: 303 TSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLE 362
TSGG F+ATSYDYDA +DEYG +PK+ HL+ +HKAIK E AL+ATDP + SLG NLE
Sbjct: 969 TSGGLFIATSYDYDAPLDEYGLENEPKYEHLRALHKAIKQSEPALVATDPKVQSLGYNLE 1148
Query: 363 AAVYKTEAECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESM 422
A V+ C AF+AN D S A KF Y+LP WS+SILPDCK VV NTAK+
Sbjct: 1149AHVFSAPGACAAFIANYDTKSYAKAKFGNGQYDLPPWSISILPDCKTVVYNTAKVG---- 1316
Query: 423 ISSFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSL 482
LK++ + S+ EP S+ADS + + L EQ+N T D SDYLWY
Sbjct: 1317-----YGWLKKMTPVNSAFAWQSYNEEPASSSQADSIAAYALWEQVNVTRDSSDYLWYMT 1481
Query: 483 RIDLEDDA-----GAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAG 537
+++ + G +L + S GH LH FING+LAG+ G N K+ + L AG
Sbjct: 1482DVNVNANEGFLKNGQSPLLTVMSAGHVLHVFINGQLAGTVWGGLGNPKLTFSDNVKLRAG 1661
Query: 538 KNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDL 597
N + LLS+ VGL + G F+TW AG+ GPV LKGL G T D S ++W+Y++GLKGE L
Sbjct: 1662NNKLSLLSVAVGLPNVGVHFETWNAGVLGPVTLKGLNEG-TRDLSRQKWSYKVGLKGESL 1838
Query: 598 GL--PSGTSG-QWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSI 654
L SG+S +W S + K QPLTWYK FSAP+GNDP+A+D MGKGE WVNG+SI
Sbjct: 1839SLHTESGSSSVEWIQGSLVAKKQPLTWYKTTFSAPAGNDPLALDLGSMGKGEVWVNGRSI 2018
Query: 655 GRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEE 714
GR+WP YI+ G ++CNY G Y +KC NCG+PSQ YHVPRSWL N+LV+FEE
Sbjct: 2019GRHWPGYIA--HGSCNACNYAGYYTDTKCRTNCGQPSQRWYHVPRSWLSSGGNSLVVFEE 2192
Query: 715 SGGDPTQISI 724
GGDP I++
Sbjct: 2193WGGDPNGIAL 2222
>TC226828 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , partial (47%)
Length = 1488
Score = 692 bits (1786), Expect = 0.0
Identities = 330/424 (77%), Positives = 364/424 (85%), Gaps = 1/424 (0%)
Frame = +1
Query: 418 NSESMISSFTTESLKE-VGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSD 476
NS S ISSFTTESLKE +GS E S GWSWISEP+GISKADSF + GLLEQINTTAD+SD
Sbjct: 4 NSASAISSFTTESLKEDIGSSEASSTGWSWISEPVGISKADSFPQTGLLEQINTTADKSD 183
Query: 477 YLWYSLRIDLEDDAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVA 536
YLWYSL ID + DAG+QTVLHIESLGHALHAFINGKLAGS GN K V+IP+TLVA
Sbjct: 184 YLWYSLSIDYKGDAGSQTVLHIESLGHALHAFINGKLAGSQTGNSGKYKFTVDIPVTLVA 363
Query: 537 GKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGED 596
GKNTIDLLSLTVGLQ++G FFDTWGAGITGPVILKGL NG+TLD S ++WTYQ+GLKGED
Sbjct: 364 GKNTIDLLSLTVGLQNYGAFFDTWGAGITGPVILKGLANGNTLDLSYQKWTYQVGLKGED 543
Query: 597 LGLPSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGR 656
LGL SG+SGQWNSQST PKNQPL WYK F+APSG+DP+AIDFTGMGKGEAWVNGQSIGR
Sbjct: 544 LGLSSGSSGQWNSQSTFPKNQPLIWYKTTFAAPSGSDPVAIDFTGMGKGEAWVNGQSIGR 723
Query: 657 YWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESG 716
YWPTY++ GCT SCNYRGPY +SKC +NCGKPSQTLYHVPRSWL+P N LVLFEE G
Sbjct: 724 YWPTYVASDAGCTDSCNYRGPYSASKCRRNCGKPSQTLYHVPRSWLKPSGNILVLFEEKG 903
Query: 717 GDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTESDRSGGPVLSLECPYPNEVITTIK 776
GDPTQIS K S C+HVS+SHPPPVDLWNSDTES R GPVLSL CP+ N+VI++IK
Sbjct: 904 GDPTQISFVTKQTESLCAHVSDSHPPPVDLWNSDTESGRKVGPVLSLTCPHDNQVISSIK 1083
Query: 777 FASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTFGDPCGGVTKSLAV 836
FAS+GTP G CGNF HG CSS +ALSIVQKACIGSSSCS+GVS TFG+PC GV KSLAV
Sbjct: 1084FASYGTPLGTCGNFYHGRCSSNKALSIVQKACIGSSSCSVGVSSETFGNPCRGVAKSLAV 1263
Query: 837 EASC 840
EA+C
Sbjct: 1264EATC 1275
>TC207366 similar to UP|Q94BZ3 (Q94BZ3) At2g32810/F24L7.5, partial (82%)
Length = 2012
Score = 441 bits (1134), Expect = e-124
Identities = 242/556 (43%), Positives = 322/556 (57%), Gaps = 41/556 (7%)
Frame = +2
Query: 327 QPKWGHLKDVHKAIKLCEEALIATD-PTITSLGSNLEAAVYKTEAE-------------- 371
+PKWGHLKD+H A+KLCE AL+ATD PT LG EA VY+
Sbjct: 2 EPKWGHLKDLHAALKLCEPALVATDSPTYIKLGPKQEAHVYQANVHLEGLNLSMFESSSI 181
Query: 372 CVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMI-------- 423
C AFLANID +ATV F G Y +P WSVS+LPDC+N V NTAK+ +++ +
Sbjct: 182 CSAFLANIDEWKEATVTFRGQRYTIPPWSVSVLPDCRNTVFNTAKVRAQTSVKLVESYLP 361
Query: 424 ---SSFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWY 480
+ F + L+ S W EP+ I SF+ G+ E +N T D+SDYLWY
Sbjct: 362 TVSNIFPAQQLRHQNDFYYISKSWMTTKEPLNIWSKSSFTVEGIWEHLNVTKDQSDYLWY 541
Query: 481 SLRIDLED-------DAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPIT 533
S R+ + D + L I+ + L FING+L G+ G+ + V +
Sbjct: 542 STRVYVSDSDILFWEENDVHPKLTIDGVRDILRVFINGQLIGNVVGHW----IKVVQTLQ 709
Query: 534 LVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLK 593
+ G N + LL+ TVGLQ++G F + GAGI G + + G +NG +D S WTYQ+GL+
Sbjct: 710 FLPGYNDLTLLTQTVGLQNYGAFLEKDGAGIRGKIKITGFENGD-IDLSKSLWTYQVGLQ 886
Query: 594 GEDLGLPS--GTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNG 651
GE L S + +W + TWYK F P G DP+A+DF MGKG+AWVNG
Sbjct: 887 GEFLKFYSEENENSEWVELTPDAIPSTFTWYKTYFDVPGGIDPVALDFKSMGKGQAWVNG 1066
Query: 652 QSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVL 711
Q IGRYW T +SP GC C+YRG Y+S KC NCGKP+QTLYHVPRSWL+ +N LV+
Sbjct: 1067QHIGRYW-TRVSPKSGCQQVCDYRGAYNSDKCSTNCGKPTQTLYHVPRSWLKATNNLLVI 1243
Query: 712 FEESGGDPTQISIAIKLIGSSCSHVSESHPPPVD-LWNSDTESDRSGG----PVLSLECP 766
EE+GG+P +IS+ + C+ VSES+ PP+ L N+D + P L L C
Sbjct: 1244LEETGGNPFEISVKLHSSRIICAQVSESNYPPLQKLVNADLIGEEVSANNMIPELHLHC- 1420
Query: 767 YPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTFG-D 825
I+++ FASFGTP G+C NFS G+C + ++SIV +AC G SCSI +S + FG D
Sbjct: 1421QQGHTISSVAFASFGTPGGSCQNFSRGNCHAPSSMSIVSEACQGKRSCSIKISDSAFGVD 1600
Query: 826 PCGGVTKSLAVEASCT 841
PC GV K+L+VEA CT
Sbjct: 1601PCPGVVKTLSVEARCT 1648
>TC206486 similar to UP|O65761 (O65761) Beta-galactosidase (Fragment) ,
partial (61%)
Length = 1763
Score = 371 bits (952), Expect = e-103
Identities = 197/450 (43%), Positives = 267/450 (58%), Gaps = 11/450 (2%)
Frame = +2
Query: 403 ILPDCKNVVLNTAKINSESMISSFTTESLKEVGSLEGPSPGWSWIS--EPIGISKADSFS 460
ILPDCKN V NTA++ S+S T + G+SW+S E + SF+
Sbjct: 2 ILPDCKNTVYNTARVGSQSAQMKMTRVPIHG---------GFSWLSFNEETTTTDDSSFT 154
Query: 461 RFGLLEQINTTADRSDYLWYSLRIDLEDDA-----GAQTVLHIESLGHALHAFINGKLAG 515
GLLEQ+NTT D SDYLWYS + L+ + G VL + S GHALH FING+L+G
Sbjct: 155 MTGLLEQLNTTRDLSDYLWYSTDVVLDPNERFLRNGKDPVLTVFSAGHALHVFINGQLSG 334
Query: 516 SGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKN 575
+ G+ + K+ + L AG N I LLS+ VGL + GP F+TW AG+ GP+ L GL
Sbjct: 335 TAYGSLEFPKLTFNEGVKLRAGVNKISLLSVAVGLPNVGPHFETWNAGVLGPISLSGLNE 514
Query: 576 GSTLDFSSKQWTYQIGLKGEDLGLPS---GTSGQWNSQSTLPKNQPLTWYKINFSAPSGN 632
G D S ++W+Y++GLKGE L L S +S +W S + + QPLTWYK F AP+G
Sbjct: 515 GRR-DLSWQKWSYKVGLKGETLSLHSLSGSSSVEWIQGSLVSQRQPLTWYKTTFDAPAGT 691
Query: 633 DPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQ 692
P+A+D MGKG+ W+NGQ++GRYWP Y G C+Y G Y+ +KC NCG+ SQ
Sbjct: 692 APLALDMDSMGKGQVWLNGQNLGRYWPAY--KASGTCDYCDYAGTYNENKCRSNCGEASQ 865
Query: 693 TLYHVPRSWLQPDSNTLVLFEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTE 752
YHVP+SWL+P N LV+FEE GGDP I + + I S C+ + E P + + T
Sbjct: 866 RWYHVPQSWLKPTGNLLVVFEELGGDPNGIFLVRRDIDSVCADIYEWQPNLIS-YQMQTS 1042
Query: 753 SDRSGGPVLSLECPYPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSS 812
P + L C P + I++IKFASFGTP G+CGNF G C + ++ ++ C+G +
Sbjct: 1043GKAPVRPKVHLSCS-PGQKISSIKFASFGTPAGSCGNFHEGSCHAHKSYDAFERNCVGQN 1219
Query: 813 SCSIGVSINTF-GDPCGGVTKSLAVEASCT 841
C++ VS F GDPC V K L+VEA C+
Sbjct: 1220WCTVTVSPENFGGDPCPNVLKKLSVEAICS 1309
>TC232994 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , partial (22%)
Length = 735
Score = 319 bits (817), Expect = 4e-87
Identities = 148/185 (80%), Positives = 160/185 (86%)
Frame = +1
Query: 656 RYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEES 715
RYWPTY SP GCT SCNYRG YD+SKCLKNCGKPSQTLYHVPRSWL+PD NTLVLFEES
Sbjct: 1 RYWPTYASPKGGCTDSCNYRGAYDASKCLKNCGKPSQTLYHVPRSWLRPDRNTLVLFEES 180
Query: 716 GGDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTESDRSGGPVLSLECPYPNEVITTI 775
GG+P QIS A K IGS CSHVSESHPPPVD WNS+TES R PV+SLECPYPN+V+++I
Sbjct: 181 GGNPKQISFATKQIGSVCSHVSESHPPPVDSWNSNTESGRKVVPVVSLECPYPNQVVSSI 360
Query: 776 KFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTFGDPCGGVTKSLA 835
KFASFGTP G CGNF HG CSS +ALSIVQKACIGSSSC I +S+NTFGDPC GV KSLA
Sbjct: 361 KFASFGTPLGTCGNFKHGLCSSNKALSIVQKACIGSSSCRIELSVNTFGDPCKGVAKSLA 540
Query: 836 VEASC 840
VEASC
Sbjct: 541 VEASC 555
>BE660170
Length = 512
Score = 310 bits (795), Expect = 1e-84
Identities = 142/164 (86%), Positives = 155/164 (93%)
Frame = +1
Query: 2 AMRRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPD 61
AMR +Q LL+LLW C+YAP+ F ANVTYDHRALVIDGKRRVL+SGSIHYPRSTPEMWPD
Sbjct: 16 AMRTSQILLVLLWFFCIYAPSSFGANVTYDHRALVIDGKRRVLVSGSIHYPRSTPEMWPD 195
Query: 62 LIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYAC 121
LIQK+KDGGLDVIETYVFWNLHEPV+GQYNFEGRGDLVKFVK VA AGLYVHLRIGPYAC
Sbjct: 196 LIQKSKDGGLDVIETYVFWNLHEPVRGQYNFEGRGDLVKFVKVVAAAGLYVHLRIGPYAC 375
Query: 122 AEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQ 165
AEWNYGGFPLWLHFIPGIQFRTDN+PF+AEM++FTAKIVD+M +
Sbjct: 376 AEWNYGGFPLWLHFIPGIQFRTDNKPFEAEMKQFTAKIVDLMNK 507
>TC235153 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , partial (18%)
Length = 445
Score = 262 bits (670), Expect = 4e-70
Identities = 121/145 (83%), Positives = 128/145 (87%)
Frame = +3
Query: 235 FYCDQFTPNSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMY 294
FY D+FTPNS KPK WTE ++GW L FGGAVPYRPVEDLAFAVARFFQ GGT QNYYMY
Sbjct: 3 FYGDEFTPNSNTKPKMWTENWSGWFLVFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMY 182
Query: 295 HGGTNFGRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTI 354
HGGTNF R SGGPF+ATSYDYDA IDEYG IRQPKWGHLK+VHKAIKLCEEALIATDPTI
Sbjct: 183 HGGTNFDRASGGPFIATSYDYDAPIDEYGIIRQPKWGHLKEVHKAIKLCEEALIATDPTI 362
Query: 355 TSLGSNLEAAVYKTEAECVAFLANI 379
TSLG NLEAAVYKT + C AFLAN+
Sbjct: 363 TSLGPNLEAAVYKTGSVCAAFLANV 437
>TC217386 similar to GB|AAF70823.1|7939621|AF154422 beta-galactosidase
{Lycopersicon esculentum;} , partial (21%)
Length = 1267
Score = 257 bits (656), Expect = 2e-68
Identities = 134/299 (44%), Positives = 178/299 (58%), Gaps = 6/299 (2%)
Frame = +1
Query: 549 GLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGLPSGTS---G 605
GLQ GPF+D GAG+T V +KGLKNG T+D SS WTY+IG++GE L L G
Sbjct: 67 GLQTAGPFYDFIGAGLTS-VKIKGLKNG-TIDLSSYAWTYKIGVQGEYLRLYQGNGLNKV 240
Query: 606 QWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYIS-P 664
W S S K QPLTWYK AP G++P+ +D MGKG AW+NG+ IGRYWP
Sbjct: 241 NWTSTSEPQKMQPLTWYKAIVDAPPGDEPVGLDMLHMGKGLAWLNGEEIGRYWPRKSEFK 420
Query: 665 IDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISI 724
+ C C+YRG ++ KC CG+P+Q YHVPRSW +P N LVLFEE GGDP +I
Sbjct: 421 SEDCVKECDYRGKFNPDKCDTGCGEPTQRWYHVPRSWFKPSGNILVLFEEKGGDPEKIKF 600
Query: 725 AIKLIGSSCSHVSESHPP-PVDLWNSDTESDRSGGPVLSLECPYPNEVITTIKFASFGTP 783
+ + +C+ V+E +P + D P L CP N I+ +KFASFGTP
Sbjct: 601 VRRKVSGACALVAEDYPSVALVSQGEDKIQSNKNIPFAHLTCP-SNTRISAVKFASFGTP 777
Query: 784 HGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTF-GDPCGGVTKSLAVEASCT 841
G CG++ GDC + +IV+KAC+ + C I ++ F + C G+++ LAVEA C+
Sbjct: 778 SGTCGSYLKGDCHDPNSSTIVEKACLNKNDCVIKLTEENFKSNLCPGLSRKLAVEAVCS 954
>TC213514 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , partial (17%)
Length = 661
Score = 253 bits (647), Expect = 2e-67
Identities = 120/146 (82%), Positives = 128/146 (87%)
Frame = +2
Query: 695 YHVPRSWLQPDSNTLVLFEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTESD 754
YH+PRSWLQPDSNTLVLFEESGGDPTQIS A K IGS CSHVSESHPPPVDLWNSD
Sbjct: 80 YHIPRSWLQPDSNTLVLFEESGGDPTQISFATKQIGSMCSHVSESHPPPVDLWNSD--KG 253
Query: 755 RSGGPVLSLECPYPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSC 814
R GPVLSLECPYPN++I++IKFASFGTP+G CGNF HG C S +AL IVQKACIGSSSC
Sbjct: 254 RKVGPVLSLECPYPNQLISSIKFASFGTPYGTCGNFKHGRCRSNKALFIVQKACIGSSSC 433
Query: 815 SIGVSINTFGDPCGGVTKSLAVEASC 840
IG+SINTFGDPC GVTKSLAVEASC
Sbjct: 434 RIGISINTFGDPCKGVTKSLAVEASC 511
>TC219425
Length = 1170
Score = 224 bits (571), Expect = 1e-58
Identities = 132/378 (34%), Positives = 191/378 (49%), Gaps = 16/378 (4%)
Frame = +3
Query: 479 WYSLRIDLEDD-----AGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPIT 533
WY+ +L + G VL + SLGH++ AF+NG + G+ G + + P+
Sbjct: 6 WYTTSFELSQEDMSMKPGVLPVLRVMSLGHSMVAFVNGDIVGTAHGTHEEKSFEFQTPVL 185
Query: 534 LVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLK 593
L G N I LLS TVGL G + + AG IL GL G TLD + W +++GLK
Sbjct: 186 LRVGTNYISLLSSTVGLPDSGAYMEHRYAGPKSINIL-GLNRG-TLDLTRNGWGHRVGLK 359
Query: 594 GEDLGLPS---GTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVN 650
GE + S TS +W +P+ L+WY+ F P G P+AI +GM KG WVN
Sbjct: 360 GEGKKVFSEEGSTSVKWKPLGAVPR--ALSWYRTRFGTPEGTGPVAIRMSGMAKGMVWVN 533
Query: 651 GQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLV 710
G +IGRYW +Y+SP+ GKP+Q+ YH+PRS+L P N LV
Sbjct: 534 GNNIGRYWMSYLSPL----------------------GKPTQSEYHIPRSFLNPQDNLLV 647
Query: 711 LFEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTES----DRSGGPVLSLECP 766
+FEE P Q+ I + CS V E P V+ W S + +S G S+ C
Sbjct: 648 IFEEEARVPAQVEILNVNRDTICSVVGERDPANVNSWVSRRGNFHPVVKSVGAAASMACA 827
Query: 767 YPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTFG-- 824
++ ++FASFG P G CG+F+ G C++ + IV++ C+G +C++ + F
Sbjct: 828 TGKRIV-AVEFASFGNPSGYCGDFAMGSCNAAASKQIVERECLGQEACTLALDRAVFNNN 1004
Query: 825 --DPCGGVTKSLAVEASC 840
D C + K LAV+ C
Sbjct: 1005GVDACPDLVKQLAVQVRC 1058
>TC207103 homologue to UP|O65761 (O65761) Beta-galactosidase (Fragment) ,
partial (36%)
Length = 1230
Score = 214 bits (544), Expect(2) = 7e-56
Identities = 109/229 (47%), Positives = 138/229 (59%), Gaps = 2/229 (0%)
Frame = +2
Query: 615 KNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNY 674
+ QPLTWYK F A +G P+A+D MGKG+ W+NGQS+GRYWP Y + G CNY
Sbjct: 92 RRQPLTWYKTTFDAQAGVAPLALDMGSMGKGQVWINGQSLGRYWPAYKA--SGSCGYCNY 265
Query: 675 RGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISIAIKLIGSSCS 734
G Y+ KC NCG+ SQ YHVP SWL+P N LV+FEE GGDP I + + I S C+
Sbjct: 266 AGTYNEKKCGSNCGQASQRWYHVPHSWLKPTGNLLVVFEELGGDPNGIFLVRRDIDSVCA 445
Query: 735 HVSESHPPPVDLWNSDTESDRSG-GPVLSLECPYPNEVITTIKFASFGTPHGNCGNFSHG 793
+ E P V + RS P L C P + I++IKFASFGTP G+CGN+ G
Sbjct: 446 DIYEWQPNLVSYDMQASGKVRSPVRPKAHLSCG-PGQKISSIKFASFGTPVGSCGNYREG 622
Query: 794 DCSSKQALSIVQKACIGSSSCSIGVSINTF-GDPCGGVTKSLAVEASCT 841
C + ++ QK C+G S C++ VS F GDPC V K L+VEA CT
Sbjct: 623 SCHAHKSYDAFQKNCVGQSWCTVTVSPEIFGGDPCPSVMKKLSVEAICT 769
Score = 23.1 bits (48), Expect(2) = 7e-56
Identities = 12/25 (48%), Positives = 16/25 (64%), Gaps = 3/25 (12%)
Frame = +1
Query: 588 YQIGLKGEDLGLPS---GTSGQWNS 609
Y++GLKGE L L S +S +W S
Sbjct: 1 YKVGLKGEALNLHSLTGSSSVEWAS 75
>TC230290 weakly similar to UP|Q93X58 (Q93X58) Beta-galactosidase , partial
(23%)
Length = 585
Score = 176 bits (447), Expect = 3e-44
Identities = 88/200 (44%), Positives = 117/200 (58%), Gaps = 1/200 (0%)
Frame = +1
Query: 643 GKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWL 702
GKG+ W+NG+SIGR+WP YI+ G C+Y G Y KC NCG+PSQ YH+PRSWL
Sbjct: 1 GKGQVWINGRSIGRHWPGYIAR--GSCGDCDYAGTYTDKKCRTNCGEPSQRWYHIPRSWL 174
Query: 703 QPDSNTLVLFEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTESDRSGGPVLS 762
P N LV+FEE GGDPT I++ + S C+ + + P + +S + P
Sbjct: 175 NPSGNYLVVFEEWGGDPTGITLVKRTTASVCADIYQGQPTLKN--RQMLDSGKVIRPKAH 348
Query: 763 LECPYPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINT 822
L CP P + I+ IKFAS+G PHG CGNF G C + ++ QK CIG S + V
Sbjct: 349 LWCP-PGKNISKIKFASYGLPHGTCGNFREGSCHAHKSYDAPQKNCIGKQSSLVTVPPEV 525
Query: 823 F-GDPCGGVTKSLAVEASCT 841
F GDPC G+ K ++ A C+
Sbjct: 526 FGGDPCPGIAKKSSL*ALCS 585
>AW153080 weakly similar to GP|14517399|gb| At2g32810/F24L7.5 {Arabidopsis
thaliana}, partial (16%)
Length = 493
Score = 155 bits (391), Expect = 9e-38
Identities = 80/184 (43%), Positives = 107/184 (57%), Gaps = 1/184 (0%)
Frame = +2
Query: 632 NDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPY-DSSKCLKNCGKP 690
NDP+ +D G+GKGEAWVNG SIGRYW ++I+ +GC+ +C+YRG Y + KC NCG P
Sbjct: 2 NDPVVVDLQGLGKGEAWVNGLSIGRYWTSWITATNGCSDTCDYRG*YVPAQKCNTNCGDP 181
Query: 691 SQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSD 750
SQ HVPRS+L+ D NTLVLFEE GG+P +S + G+ C+ V E
Sbjct: 182 SQMWVHVPRSFLKNDKNTLVLFEEIGGNP*NVSFRTVITGTICARVQEC----------- 328
Query: 751 TESDRSGGPVLSLECPYPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIG 810
+L L C N I+ I+F+SFG P NCG F+ + V+ AC+G
Sbjct: 329 --------ALLELSCQGGN-TISQIQFSSFGNPTRNCGLFTKCAWEATDGQPAVEAACVG 481
Query: 811 SSSC 814
+SC
Sbjct: 482 MNSC 493
>AI960920 similar to GP|14970841|emb beta-galactosidase {Fragaria x
ananassa}, partial (10%)
Length = 285
Score = 155 bits (391), Expect = 9e-38
Identities = 75/92 (81%), Positives = 83/92 (89%)
Frame = +2
Query: 489 DAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTV 548
DAGAQT LHI+SLGHALHAFINGKLAGSG GN + + V V+IPITLV+GKNTIDLLSLTV
Sbjct: 5 DAGAQTFLHIKSLGHALHAFINGKLAGSGTGNHEKANVEVDIPITLVSGKNTIDLLSLTV 184
Query: 549 GLQHFGPFFDTWGAGITGPVILKGLKNGSTLD 580
GLQ++G FFDTWGAGITGPVILK LKNGS +D
Sbjct: 185 GLQNYGAFFDTWGAGITGPVILKCLKNGSNVD 280
>BU760871 homologue to GP|3204134|emb beta-galactosidase {Cicer arietinum},
partial (21%)
Length = 445
Score = 108 bits (270), Expect(2) = 5e-37
Identities = 46/63 (73%), Positives = 55/63 (87%)
Frame = +2
Query: 297 GTNFGRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITS 356
GTNFGRT+GGPF+ATSYDYDA +DEYG RQPKWGHLKD+H+AIKLCE AL++ D T+
Sbjct: 2 GTNFGRTAGGPFIATSYDYDAPLDEYGLARQPKWGHLKDLHRAIKLCEPALVSGDSTVQR 181
Query: 357 LGS 359
LG+
Sbjct: 182 LGN 190
Score = 65.5 bits (158), Expect(2) = 5e-37
Identities = 39/89 (43%), Positives = 49/89 (54%)
Frame = +1
Query: 387 VKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTESLKEVGSLEGPSPGWSW 446
V F YNLP WS+SILP+CK+ V NTA++ S+S TT + V G S W
Sbjct: 199 VAFGNQHYNLPPWSISILPNCKHTVYNTARVGSQS-----TTMKMTRVPIHGGLS--WKA 357
Query: 447 ISEPIGISKADSFSRFGLLEQINTTADRS 475
+E + SF+ GLLEQIN T D S
Sbjct: 358 FNEETTTTDDSSFTVTGLLEQINATRDLS 444
>AW234372 homologue to GP|7682680|gb| beta galactosidase {Vigna radiata},
partial (15%)
Length = 406
Score = 151 bits (382), Expect = 1e-36
Identities = 71/108 (65%), Positives = 88/108 (80%)
Frame = +1
Query: 6 TQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQK 65
++ LLL+ +L V + C+ VTYD +A++I+G+RR+LISGSIHYPRSTPEMW DLI+K
Sbjct: 85 SKLLLLVFSVLFVGSELIHCS-VTYDRKAIIINGQRRILISGSIHYPRSTPEMWEDLIRK 261
Query: 66 AKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVH 113
AKDGGLDVI+TYVFWN+HEP G YNFEGR DLV+F+K V GLYVH
Sbjct: 262 AKDGGLDVIDTYVFWNVHEPSPGNYNFEGRNDLVRFIKTVQRVGLYVH 405
>TC204600 homologue to UP|Q9M5J3 (Q9M5J3) Beta galactosidase, partial (21%)
Length = 567
Score = 151 bits (381), Expect = 1e-36
Identities = 78/158 (49%), Positives = 102/158 (64%), Gaps = 4/158 (2%)
Frame = +2
Query: 554 GPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGL--PSGTSG-QWNSQ 610
G F+T AGITG V+L GL +G D + ++W+YQIGL+GE + L P+G S W
Sbjct: 8 GFHFET*KAGITG-VLLNGLDHGQK-DLTWQKWSYQIGLRGEAMNLVAPNGVSSVDWEKD 181
Query: 611 STLPKNQP-LTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCT 669
S ++Q L W+K F+AP G +P+A+D + MGKG+ W+NGQSIGRYW Y G
Sbjct: 182 SLAVRSQSQLKWHKAYFNAPEGVEPLALDLSSMGKGQVWINGQSIGRYWMVYAK---GSC 352
Query: 670 SSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSN 707
SSCNY G Y +KC CG+P+Q YHVPRSWL+P N
Sbjct: 353 SSCNYAGTYRPAKCQLGCGQPTQRWYHVPRSWLRPTKN 466
>AW703960 similar to GP|7682677|gb| beta galactosidase {Vigna radiata},
partial (16%)
Length = 370
Score = 140 bits (353), Expect = 2e-33
Identities = 69/123 (56%), Positives = 88/123 (71%)
Frame = +2
Query: 7 QFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKA 66
+F ++L +LC++ A+VTYDH A+V+DG RR+LISGSIH+P TP+MWP LIQ+A
Sbjct: 5 EFNGVVLMMLCLWVCGV-TASVTYDHIAIVVDGNRRILISGSIHFPSCTPQMWPYLIQEA 181
Query: 67 KDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNY 126
KDGGLD I+TYVFWN+HEP + FE R DLV + K A LYV L IGP+ C+ WN
Sbjct: 182 KDGGLDAIQTYVFWNVHEPSPRRCYFEDRVDLV*YEKSAQ*ARLYVLLSIGPFICS*WNL 361
Query: 127 GGF 129
GF
Sbjct: 362 WGF 370
>AW704810 similar to GP|7682677|gb| beta galactosidase {Vigna radiata},
partial (19%)
Length = 425
Score = 134 bits (338), Expect = 1e-31
Identities = 68/139 (48%), Positives = 90/139 (63%), Gaps = 5/139 (3%)
Frame = +3
Query: 466 EQINTTADRSDYLWYSLRIDLEDDAG-----AQTVLHIESLGHALHAFINGKLAGSGKGN 520
EQIN T D +DYLWY ++++ + G VL + S GHALH FING+L+G+ G
Sbjct: 9 EQINVTRDSTDYLWYMTDVNIDANEGFIKNGQSPVLTVMSAGHALHVFINGQLSGTVYGG 188
Query: 521 RDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLD 580
D+ K+ + L G N I LLS+TVGL + GP+F+TW AG+ GPV LKGL G T D
Sbjct: 189 LDSHKLTFSDSVKLRVGNNKISLLSITVGLPNVGPYFETWNAGVLGPVTLKGLNEG-TRD 365
Query: 581 FSSKQWTYQIGLKGEDLGL 599
S ++W+Y+IGLKGE L L
Sbjct: 366 LSKQKWSYKIGLKGEALNL 422
>BE805724
Length = 370
Score = 122 bits (307), Expect = 5e-28
Identities = 61/123 (49%), Positives = 78/123 (62%), Gaps = 4/123 (3%)
Frame = +1
Query: 579 LDFSSKQWTYQIGLKGEDLGL--PSGTSGQWNSQSTL--PKNQPLTWYKINFSAPSGNDP 634
LD S ++WTYQ+GLKGE + L P+G S QS L KNQPLTW+K F AP G++P
Sbjct: 10 LDLSWQKWTYQVGLKGEAMNLASPNGISSVEWMQSALVSEKNQPLTWHKTYFDAPDGDEP 189
Query: 635 IAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTL 694
+A+D GMGKG+ W+NG SIGRYW +P G + C+Y G + KC G P+Q
Sbjct: 190 LALDMEGMGKGQIWINGLSIGRYW---TAPAAGICNGCSYAGTFRPPKCQVGXGXPTQQW 360
Query: 695 YHV 697
YHV
Sbjct: 361 YHV 369
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,483,128
Number of Sequences: 63676
Number of extensions: 723076
Number of successful extensions: 3401
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 3312
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3346
length of query: 841
length of database: 12,639,632
effective HSP length: 105
effective length of query: 736
effective length of database: 5,953,652
effective search space: 4381887872
effective search space used: 4381887872
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0136.7


