

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.17
(83 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
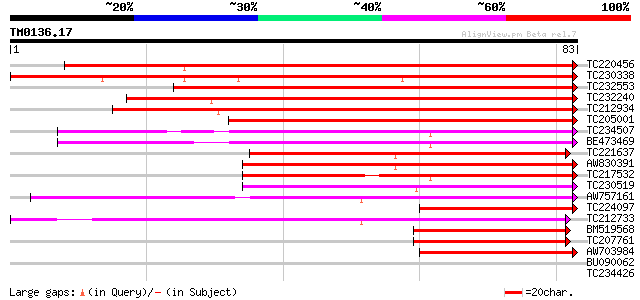
Sequences producing significant alignments: (bits) Value
TC220456 weakly similar to UP|O81262 (O81262) Early nodule-speci... 106 2e-24
TC230338 similar to UP|Q8W0Y6 (Q8W0Y6) Enod8.2, partial (49%) 100 1e-22
TC232553 similar to UP|Q9FXE5 (Q9FXE5) F12A21.4, partial (59%) 86 3e-18
TC232240 weakly similar to UP|Q9FXE5 (Q9FXE5) F12A21.4, partial ... 79 4e-16
TC212934 similar to UP|O82681 (O82681) Lanatoside 15'-O-acetyles... 75 8e-15
TC205001 similar to UP|O82681 (O82681) Lanatoside 15'-O-acetyles... 69 3e-13
TC234507 similar to UP|Q6H8W7 (Q6H8W7) Esterase precursor, parti... 60 2e-10
BE473469 similar to GP|10764858|gb| F1K23.17 {Arabidopsis thalia... 57 2e-09
TC221637 similar to UP|O82076 (O82076) Lipase homolog (Fragment)... 53 2e-08
AW830391 53 3e-08
TC217532 weakly similar to UP|Q9LHW8 (Q9LHW8) ESTs AU075812(E608... 53 3e-08
TC230519 weakly similar to PIR|T06696|T06696 lipase homolog T29H... 44 1e-05
AW757161 42 5e-05
TC224097 similar to UP|Q8L8G1 (Q8L8G1) Lipase SIL1, partial (32%) 41 1e-04
TC212733 weakly similar to UP|Q9FLN0 (Q9FLN0) GDSL-motif lipase/... 40 2e-04
BM519568 homologue to GP|14586378|emb calcium-dependent protein ... 40 2e-04
TC207761 weakly similar to UP|Q8L8G1 (Q8L8G1) Lipase SIL1, parti... 40 2e-04
AW703984 similar to GP|20146410|dbj lipase-like protein {Oryza s... 40 3e-04
BU090062 37 0.001
TC234426 36 0.004
>TC220456 weakly similar to UP|O81262 (O81262) Early nodule-specific protein,
partial (43%)
Length = 587
Score = 106 bits (265), Expect = 2e-24
Identities = 50/77 (64%), Positives = 58/77 (74%), Gaps = 2/77 (2%)
Frame = +1
Query: 9 HVSFVSLFVILFTTTL--NPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTY 66
H VSLF IL T+ NP FA K C FPAIFN G SNSDTGGLAA+ + P PYG+TY
Sbjct: 40 HNQHVSLFAILSIATIVPNPAFATKECVFPAIFNFGDSNSDTGGLAASLIAPTPPYGETY 219
Query: 67 FHRPAGRYSDGRIILDF 83
FHRPAGR+SDGR+++DF
Sbjct: 220 FHRPAGRFSDGRLVIDF 270
>TC230338 similar to UP|Q8W0Y6 (Q8W0Y6) Enod8.2, partial (49%)
Length = 737
Score = 100 bits (249), Expect = 1e-22
Identities = 55/96 (57%), Positives = 66/96 (68%), Gaps = 13/96 (13%)
Frame = +2
Query: 1 MSLMEFHRHVSF-------VSLFVILFTTTL--NPVFAKKR--CNFPAIFNLGASNSDTG 49
MS MEFH F SL ++ TT+ NP A K+ C+FPAIFN GASN+DTG
Sbjct: 11 MSPMEFHSIAKFHNIPLVSSSLVILCIATTILNNPAMATKQYYCDFPAIFNFGASNADTG 190
Query: 50 GLAAAFL--PPKSPYGDTYFHRPAGRYSDGRIILDF 83
GLAA+F PKSP G+TYFHRPAGR+SDGR+I+DF
Sbjct: 191 GLAASFFVAAPKSPNGETYFHRPAGRFSDGRLIIDF 298
>TC232553 similar to UP|Q9FXE5 (Q9FXE5) F12A21.4, partial (59%)
Length = 832
Score = 85.9 bits (211), Expect = 3e-18
Identities = 38/59 (64%), Positives = 47/59 (79%)
Frame = +2
Query: 25 NPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
N + A K+C+FPAIFN G SNSDTGGL+AAF P+G++YFH PAGRY DGR+I+DF
Sbjct: 5 NSLAASKQCHFPAIFNFGDSNSDTGGLSAAFGQAGPPHGESYFHHPAGRYCDGRLIVDF 181
>TC232240 weakly similar to UP|Q9FXE5 (Q9FXE5) F12A21.4, partial (38%)
Length = 469
Score = 79.0 bits (193), Expect = 4e-16
Identities = 39/67 (58%), Positives = 48/67 (71%), Gaps = 1/67 (1%)
Frame = +2
Query: 18 ILFTTTLNPVF-AKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSD 76
++ TT + V ++ C FPAIFNLG SNSDTGGL+AAF P G TYFH P GR+SD
Sbjct: 2 VISTTLMRSVSGSESECIFPAIFNLGDSNSDTGGLSAAFGQAPPPNGITYFHSPNGRFSD 181
Query: 77 GRIILDF 83
GR+I+DF
Sbjct: 182GRLIIDF 202
>TC212934 similar to UP|O82681 (O82681) Lanatoside 15'-O-acetylesterase
precursor, partial (30%)
Length = 442
Score = 74.7 bits (182), Expect = 8e-15
Identities = 38/70 (54%), Positives = 45/70 (64%), Gaps = 2/70 (2%)
Frame = +2
Query: 16 FVILFTTTLNPVFA--KKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGR 73
F ++ T L +F+ CNF AIFN G SNSDTGG AAF PYG TYF +PAGR
Sbjct: 26 FFVIVTIVLLCLFSLSHSECNFKAIFNFGDSNSDTGGFYAAFPGESGPYGMTYFKKPAGR 205
Query: 74 YSDGRIILDF 83
SDGR+I+ F
Sbjct: 206ASDGRLIIGF 235
>TC205001 similar to UP|O82681 (O82681) Lanatoside 15'-O-acetylesterase
precursor, partial (62%)
Length = 1384
Score = 69.3 bits (168), Expect = 3e-13
Identities = 31/51 (60%), Positives = 37/51 (71%)
Frame = +1
Query: 33 CNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
C+F AIFN G SNSDTGG +F +PYG TYF +P GR SDGR+I+DF
Sbjct: 151 CDFEAIFNFGDSNSDTGGFHTSFPAQPAPYGMTYFKKPVGRASDGRLIVDF 303
>TC234507 similar to UP|Q6H8W7 (Q6H8W7) Esterase precursor, partial (18%)
Length = 433
Score = 60.1 bits (144), Expect = 2e-10
Identities = 35/81 (43%), Positives = 48/81 (59%), Gaps = 5/81 (6%)
Frame = +3
Query: 8 RHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKS-----PY 62
R +S V+ VI ++ P+ A C + +IF+ G S +DTG L + PP PY
Sbjct: 45 RWISIVAFVVIASSSA--PLLAA--CPYTSIFSFGDSFADTGNLYLSSHPPTHHCFFPPY 212
Query: 63 GDTYFHRPAGRYSDGRIILDF 83
G+TYFHR GR SDGR+I+DF
Sbjct: 213 GETYFHRVTGRCSDGRLIIDF 275
>BE473469 similar to GP|10764858|gb| F1K23.17 {Arabidopsis thaliana},
partial (5%)
Length = 415
Score = 56.6 bits (135), Expect = 2e-09
Identities = 31/81 (38%), Positives = 44/81 (54%), Gaps = 5/81 (6%)
Frame = +1
Query: 8 RHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKS-----PY 62
R ++ V V+ + T+ C + +IF+ G S +DTG L + PP PY
Sbjct: 22 RWIAIVGFVVVFSSATILAA-----CPYKSIFSFGDSFADTGNLYFSSHPPSHHCFFPPY 186
Query: 63 GDTYFHRPAGRYSDGRIILDF 83
G T+FHR GR SDGR+I+DF
Sbjct: 187GQTFFHRVTGRCSDGRLIIDF 249
>TC221637 similar to UP|O82076 (O82076) Lipase homolog (Fragment), partial
(30%)
Length = 443
Score = 53.1 bits (126), Expect = 2e-08
Identities = 25/48 (52%), Positives = 31/48 (64%), Gaps = 1/48 (2%)
Frame = +1
Query: 36 PAIFNLGASNSDTGGLAAAF-LPPKSPYGDTYFHRPAGRYSDGRIILD 82
P +F G SNSDTGGLA+ P P G +FHR GR SDGR+++D
Sbjct: 148 PVLFVFGDSNSDTGGLASGLGFPINPPNGRNFFHRSTGRLSDGRLLID 291
>AW830391
Length = 419
Score = 52.8 bits (125), Expect = 3e-08
Identities = 26/50 (52%), Positives = 32/50 (64%), Gaps = 1/50 (2%)
Frame = +3
Query: 35 FPAIFNLGASNSDTGGLAAAF-LPPKSPYGDTYFHRPAGRYSDGRIILDF 83
+PA+FN G S SDTG LAA P G YF P+GR+ DGR+I+DF
Sbjct: 132 YPAVFNFGDSYSDTGELAAGLGFQVAPPNGQDYFKIPSGRFCDGRLIVDF 281
>TC217532 weakly similar to UP|Q9LHW8 (Q9LHW8) ESTs AU075812(E60850), partial
(54%)
Length = 1372
Score = 52.8 bits (125), Expect = 3e-08
Identities = 27/56 (48%), Positives = 36/56 (64%), Gaps = 7/56 (12%)
Frame = +1
Query: 35 FPAIFNLGASNSDTGGLAAAFLPPKS-------PYGDTYFHRPAGRYSDGRIILDF 83
+ ++F+ G S +DTG L F+ P+ PYG T+FHRP GR SDGR+ILDF
Sbjct: 112 YTSLFSFGDSLTDTGNLY--FISPRQSPDCLLPPYGQTHFHRPNGRCSDGRLILDF 273
>TC230519 weakly similar to PIR|T06696|T06696 lipase homolog T29H11.20 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(24%)
Length = 588
Score = 43.9 bits (102), Expect = 1e-05
Identities = 22/55 (40%), Positives = 30/55 (54%), Gaps = 6/55 (10%)
Frame = +2
Query: 35 FPAIFNLGASNSDTGGLAAAFLPP------KSPYGDTYFHRPAGRYSDGRIILDF 83
F ++ G S +DTG A P SPYG T+F+ RYSDGR+++DF
Sbjct: 149 FKRVYAFGDSFTDTGNTQNAEGPSGFGHVSNSPYGTTFFNHSTNRYSDGRLVIDF 313
>AW757161
Length = 395
Score = 42.0 bits (97), Expect = 5e-05
Identities = 30/85 (35%), Positives = 41/85 (47%), Gaps = 5/85 (5%)
Frame = +1
Query: 4 MEFHRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGG-----LAAAFLPP 58
++F V FV +++ T L + K A+F G S D G A
Sbjct: 13 LKFSFLVLFVCCGILIPTCCLGDMCQPKEN--AALFVFGDSLFDVGNNNYINTTADNQAN 186
Query: 59 KSPYGDTYFHRPAGRYSDGRIILDF 83
SPYG+T+F P GR+SDGR+I DF
Sbjct: 187YSPYGETFFKYPTGRFSDGRVIPDF 261
>TC224097 similar to UP|Q8L8G1 (Q8L8G1) Lipase SIL1, partial (32%)
Length = 617
Score = 40.8 bits (94), Expect = 1e-04
Identities = 16/23 (69%), Positives = 19/23 (82%)
Frame = +3
Query: 61 PYGDTYFHRPAGRYSDGRIILDF 83
PYG+TYF P GR+SDGR+I DF
Sbjct: 207 PYGETYFKFPTGRFSDGRLISDF 275
>TC212733 weakly similar to UP|Q9FLN0 (Q9FLN0) GDSL-motif
lipase/hydrolase-like protein, partial (25%)
Length = 464
Score = 40.0 bits (92), Expect = 2e-04
Identities = 27/87 (31%), Positives = 43/87 (49%), Gaps = 5/87 (5%)
Frame = +1
Query: 1 MSLMEFHRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGG-----LAAAF 55
+SL+EF + +F+ + T + + A+F LG S D G ++
Sbjct: 34 ISLLEFS-----LVIFIQIMTHCHSSITTCLPEKHAALFILGDSLFDNGNNNYINTTTSY 198
Query: 56 LPPKSPYGDTYFHRPAGRYSDGRIILD 82
PYG+T+F P+GR+SDGR+I D
Sbjct: 199 QANYPPYGETFFKYPSGRFSDGRMIPD 279
>BM519568 homologue to GP|14586378|emb calcium-dependent protein kinase-like
protein {Arabidopsis thaliana}, partial (7%)
Length = 407
Score = 40.0 bits (92), Expect = 2e-04
Identities = 15/23 (65%), Positives = 21/23 (91%)
Frame = +3
Query: 60 SPYGDTYFHRPAGRYSDGRIILD 82
SPYG+T+F+ P+GR+SDGR+I D
Sbjct: 192 SPYGETFFNYPSGRFSDGRVIPD 260
>TC207761 weakly similar to UP|Q8L8G1 (Q8L8G1) Lipase SIL1, partial (46%)
Length = 1609
Score = 40.0 bits (92), Expect = 2e-04
Identities = 15/23 (65%), Positives = 21/23 (91%)
Frame = +3
Query: 60 SPYGDTYFHRPAGRYSDGRIILD 82
SPYG+T+F+ P+GR+SDGR+I D
Sbjct: 504 SPYGETFFNYPSGRFSDGRVIPD 572
>AW703984 similar to GP|20146410|dbj lipase-like protein {Oryza sativa
(japonica cultivar-group)}, partial (15%)
Length = 241
Score = 39.7 bits (91), Expect = 3e-04
Identities = 15/23 (65%), Positives = 19/23 (82%)
Frame = +1
Query: 61 PYGDTYFHRPAGRYSDGRIILDF 83
PYG T+FH +GR SDGR+I+DF
Sbjct: 31 PYGQTFFHHVSGRCSDGRLIIDF 99
>BU090062
Length = 419
Score = 37.4 bits (85), Expect = 0.001
Identities = 14/25 (56%), Positives = 20/25 (80%)
Frame = +1
Query: 59 KSPYGDTYFHRPAGRYSDGRIILDF 83
K PYG T+ +PAGR+SDGR++ D+
Sbjct: 247 KDPYGVTFPGKPAGRFSDGRVLTDY 321
>TC234426
Length = 652
Score = 35.8 bits (81), Expect = 0.004
Identities = 29/84 (34%), Positives = 39/84 (45%), Gaps = 4/84 (4%)
Frame = +1
Query: 4 MEFHRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGG---LAAAFLPPKS 60
ME H + +F + F L+ V AK PA+ G S+ D G +A
Sbjct: 13 MEMHSSI----IFCMFFLPWLSMVGAK----VPAMIAFGDSSVDAGNNNYIATVARSNFQ 168
Query: 61 PYG-DTYFHRPAGRYSDGRIILDF 83
PYG D +P GR+S+GRI DF
Sbjct: 169PYGRDFVGGKPTGRFSNGRIATDF 240
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.143 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,935,251
Number of Sequences: 63676
Number of extensions: 47174
Number of successful extensions: 389
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 382
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 384
length of query: 83
length of database: 12,639,632
effective HSP length: 59
effective length of query: 24
effective length of database: 8,882,748
effective search space: 213185952
effective search space used: 213185952
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0136.17


