

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
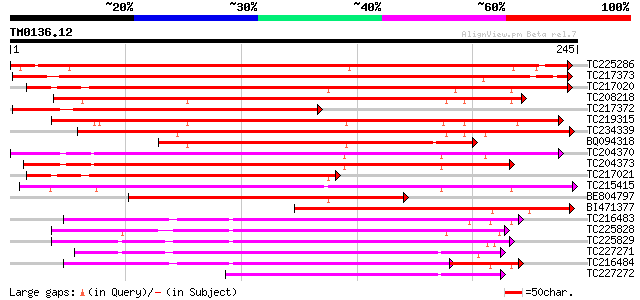
Query= TM0136.12
(245 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC225286 weakly similar to UP|Q9SP59 (Q9SP59) Arabinogalactan pr... 346 5e-96
TC217373 weakly similar to UP|Q7Y250 (Q7Y250) Arabinogalactan pr... 338 1e-93
TC217020 similar to UP|Q6J193 (Q6J193) Fasciclin-like AGP 11, pa... 221 2e-58
TC208218 weakly similar to UP|Q6J189 (Q6J189) Fasciclin-like AGP... 210 4e-55
TC217372 weakly similar to UP|Q6J193 (Q6J193) Fasciclin-like AGP... 204 2e-53
TC219315 weakly similar to UP|Q6J190 (Q6J190) Fasciclin-like AGP... 200 4e-52
TC234339 similar to UP|Q6J196 (Q6J196) Fasciclin-like AGP 8, par... 195 2e-50
BQ094318 weakly similar to GP|21553523|gb arabinogalactan protei... 169 1e-42
TC204370 weakly similar to GB|AAM19981.1|20466119|AY098971 AT5g4... 163 6e-41
TC204373 weakly similar to GB|AAM19981.1|20466119|AY098971 AT5g4... 159 1e-39
TC217021 similar to UP|Q6J193 (Q6J193) Fasciclin-like AGP 11, pa... 136 1e-32
TC215415 similar to UP|Q6J192 (Q6J192) Fasciclin-like AGP 12, pa... 120 7e-28
BE804797 101 3e-22
BI471377 99 2e-21
TC216483 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabin... 91 4e-19
TC225828 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like ara... 90 8e-19
TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like ara... 88 4e-18
TC227271 similar to GB|AAG24276.1|10880493|AF195889 fasciclin-li... 85 3e-17
TC216484 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabin... 67 3e-14
TC227272 similar to GB|AAK55685.1|14190409|AF378882 AT3g60900/T4... 69 1e-12
>TC225286 weakly similar to UP|Q9SP59 (Q9SP59) Arabinogalactan protein
Pop14A9, partial (64%)
Length = 1207
Score = 346 bits (888), Expect = 5e-96
Identities = 188/262 (71%), Positives = 217/262 (82%), Gaps = 19/262 (7%)
Frame = +1
Query: 1 MKH-QLFTLPLLALMFFFHSTTTLA---------------ASSPAPAPSSSPTDIIRILK 44
MKH LF+LPLL ++ F HSTTTLA A++PAP PS+ PTDIIRILK
Sbjct: 130 MKHIPLFSLPLL-IVLFSHSTTTLAQSPAAAPTAPATPSTAAAPAPVPSAPPTDIIRILK 306
Query: 45 KAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNE 104
KAGG+TTLIRLLQ TQV+ QIN+QL+ ++ GLT FAPNDNAFS+LKPGFLNSLNDQQKNE
Sbjct: 307 KAGGFTTLIRLLQATQVSNQINSQLLTTSGGLTLFAPNDNAFSSLKPGFLNSLNDQQKNE 486
Query: 105 LIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSS-GNTVNMTTGIVNVTVGGS 163
LIQFHLLPT+VS+SNFDTLSNPVRTQAGENPDRLALN+TSS GN VNMTTG+VNVT+GG+
Sbjct: 487 LIQFHLLPTYVSVSNFDTLSNPVRTQAGENPDRLALNITSSGGNQVNMTTGVVNVTLGGT 666
Query: 164 VYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAGD-DSAAVS 222
VY+D+QLAVYQVDKVLLPRDFF AKPP PAPAPEKAK SKKKSAD T A D +S A+S
Sbjct: 667 VYTDHQLAVYQVDKVLLPRDFFEAKPPVPAPAPEKAKGSKKKSADSTATTADDAESDAIS 846
Query: 223 VKQR-LVWGPLAVAMVVVAAFS 243
VKQR ++W +AV + +AAF+
Sbjct: 847 VKQRHVMW--VAVLALFIAAFT 906
>TC217373 weakly similar to UP|Q7Y250 (Q7Y250) Arabinogalactan protein,
partial (66%)
Length = 982
Score = 338 bits (868), Expect = 1e-93
Identities = 177/243 (72%), Positives = 204/243 (83%), Gaps = 1/243 (0%)
Frame = +2
Query: 2 KHQLFTLPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQV 61
KH LF+L LL ++ F T A ++PAP+PSS+PTDIIRILKKAGG+TTLIRLL TTQV
Sbjct: 71 KHLLFSLSLLLMVLF-----TSAQTTPAPSPSSAPTDIIRILKKAGGFTTLIRLLTTTQV 235
Query: 62 ATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFD 121
+TQINAQL+NSN GLT FAPNDNAF +LKPGFLNSLNDQQKNELIQFH+LPTFVS+SNFD
Sbjct: 236 STQINAQLLNSNNGLTVFAPNDNAFQSLKPGFLNSLNDQQKNELIQFHVLPTFVSISNFD 415
Query: 122 TLSNPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLP 181
TLSNPVRTQAG++PDRLALN+TSSGN VN+TTG+VN TVGGSVYSD+QLA+YQVDKVLLP
Sbjct: 416 TLSNPVRTQAGDDPDRLALNITSSGNQVNLTTGVVNTTVGGSVYSDHQLAIYQVDKVLLP 595
Query: 182 RDFFVAKPPAPAPAPEKAKASK-KKSADDTEGPAGDDSAAVSVKQRLVWGPLAVAMVVVA 240
RDFFV KPP PAPAP KAKAS KKS + A +DSAA+S+ +W L+VA + A
Sbjct: 596 RDFFVPKPPPPAPAPAKAKASSAKKSTEGPSSAADNDSAAISLNG--IWLSLSVA-TIAA 766
Query: 241 AFS 243
FS
Sbjct: 767 VFS 775
>TC217020 similar to UP|Q6J193 (Q6J193) Fasciclin-like AGP 11, partial (66%)
Length = 815
Score = 221 bits (563), Expect = 2e-58
Identities = 129/245 (52%), Positives = 161/245 (65%), Gaps = 9/245 (3%)
Frame = +3
Query: 8 LPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINA 67
LP L L F T A +PAPA PT+I ++L+KAG +TT I+LL+ +Q+A +IN+
Sbjct: 69 LPFLFLNIFIQ--TISAQVAPAPA---GPTNITQVLEKAGQFTTFIKLLKASQIADRINS 233
Query: 68 QLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPV 127
QL NSN GLT FAP DNAFS+LK G LNS+N Q + +LIQFH+LPT ++S F T SNP+
Sbjct: 234 QLNNSNQGLTVFAPTDNAFSSLKAGTLNSINSQDQMQLIQFHILPTLYTISQFQTASNPL 413
Query: 128 RTQAGENPD-RLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFV 186
TQAG + D LNVT+SGN VN+TTG+V+ TV ++YSDNQLAVYQVDKVLLP F
Sbjct: 414 HTQAGNSDDGEYPLNVTTSGNQVNVTTGVVDTTVSNTIYSDNQLAVYQVDKVLLPMKLFG 593
Query: 187 AKPPA--PAPAPEKAKASKKKSADDTEGPAG------DDSAAVSVKQRLVWGPLAVAMVV 238
A PA PA AP K K A + P+G D S+AVS+K+ V G VA VV
Sbjct: 594 AMAPAASPAEAPAPTKPEKNVRAGAADSPSGSSDTSADASSAVSLKRHTVEGVTFVAAVV 773
Query: 239 VAAFS 243
V S
Sbjct: 774 VMIVS 788
>TC208218 weakly similar to UP|Q6J189 (Q6J189) Fasciclin-like AGP 15, partial
(46%)
Length = 1043
Score = 210 bits (535), Expect = 4e-55
Identities = 119/221 (53%), Positives = 153/221 (68%), Gaps = 17/221 (7%)
Frame = +2
Query: 20 TTTLAASSPAP----------APSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQL 69
T A SP P +P S P DI RILKKA ++ LIRLL+TT++ IN+QL
Sbjct: 167 TPKATAPSPKPLVPTLPQSPDSPDSVPDDITRILKKAKMFSVLIRLLKTTEIMNNINSQL 346
Query: 70 INSNAG-LTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVR 128
I + +G +T AP+D+AFSNLK GFLNSLN+ QK EL+QFH+LP FVS SNFD+LSNPV+
Sbjct: 347 ITAKSGGITILAPDDSAFSNLKAGFLNSLNEGQKIELVQFHILPEFVSSSNFDSLSNPVQ 526
Query: 129 TQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVA- 187
T AG++P RL LNV + GN+VN++TG+VN TV G VYSDN+L +Y VDKVLLP DFF+
Sbjct: 527 TVAGKDPARLPLNVNALGNSVNISTGVVNATVLGVVYSDNKLGIYHVDKVLLPLDFFITN 706
Query: 188 KPPAPAPA--PEKAKASKKKSADDTEGPAG---DDSAAVSV 223
K PA AP+ + KA+K S++D + + S AVSV
Sbjct: 707 KAPASAPSTLAKSPKAAKDNSSEDDQEETNQHQNKSGAVSV 829
>TC217372 weakly similar to UP|Q6J193 (Q6J193) Fasciclin-like AGP 11, partial
(42%)
Length = 451
Score = 204 bits (520), Expect = 2e-53
Identities = 104/134 (77%), Positives = 118/134 (87%)
Frame = +2
Query: 2 KHQLFTLPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQV 61
K+ LF+L LL ++ F S TT PAP+PSS+PTDIIRILKKAGG+TTLIRLL TTQV
Sbjct: 65 KNLLFSLSLLLMVLFTSSQTT-----PAPSPSSTPTDIIRILKKAGGFTTLIRLLTTTQV 229
Query: 62 ATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFD 121
+TQINAQL+NSN GLT FAPNDNAF +LKPGFLNSLNDQQKNELIQFH+LPTFVS+SNFD
Sbjct: 230 STQINAQLLNSNNGLTVFAPNDNAFQSLKPGFLNSLNDQQKNELIQFHVLPTFVSISNFD 409
Query: 122 TLSNPVRTQAGENP 135
TLSNPVRTQAG++P
Sbjct: 410 TLSNPVRTQAGDDP 451
>TC219315 weakly similar to UP|Q6J190 (Q6J190) Fasciclin-like AGP 14, partial
(32%)
Length = 1223
Score = 200 bits (509), Expect = 4e-52
Identities = 121/242 (50%), Positives = 160/242 (66%), Gaps = 21/242 (8%)
Frame = +3
Query: 19 STTTLAASSPAPAPSSS------PT-DIIRILKKAGGYTTLIRLLQTTQVATQINAQLIN 71
+T L +P+ AP+ + PT DI++IL+KA ++ L RLL+TTQ+ Q+N+QL+
Sbjct: 267 NTVPLVPVTPSGAPTPTTIIPKGPTIDIVQILRKAKRFSVLTRLLKTTQLINQLNSQLVT 446
Query: 72 SNAG-LTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQ 130
S++G LT FAP D+AFS LK GFLNSL D+QK EL+QFH L + +S+SNFDTL+NPV+TQ
Sbjct: 447 SSSGGLTLFAPEDSAFSKLKAGFLNSLTDRQKVELLQFHTLSSVISISNFDTLTNPVQTQ 626
Query: 131 AGENPDRLALNVTS-SGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFV-AK 188
AG++P RL LNVT+ G+ V+M TG VN +V G+VYSDN+LA+YQVDKVLLP D + +K
Sbjct: 627 AGDDPQRLQLNVTTYGGSQVSMATGAVNASVTGTVYSDNKLAIYQVDKVLLPLDLVLPSK 806
Query: 189 PPAPAPA-----PEKAKASKKKSADDTEGPAGDDS------AAVSVKQRLVWGPLAVAMV 237
PAPAPA KA +ADD DDS A VS + W V +
Sbjct: 807 APAPAPALARKGLPKADKGNSTAADDGTVDGNDDSDGKALPAGVSAGCTMKWVNNVVVVG 986
Query: 238 VV 239
VV
Sbjct: 987 VV 992
>TC234339 similar to UP|Q6J196 (Q6J196) Fasciclin-like AGP 8, partial (22%)
Length = 895
Score = 195 bits (495), Expect = 2e-50
Identities = 109/238 (45%), Positives = 146/238 (60%), Gaps = 23/238 (9%)
Frame = +3
Query: 30 PAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLIN-SNAGLTFFAPNDNAFSN 88
P PS TDI+++L++A + +RL++TTQ+ Q+N+QL+ + GLT AP+D AFS
Sbjct: 18 PQPSGGTTDIVQVLQQANSFNIFLRLMKTTQLINQLNSQLLTIKSGGLTILAPDDGAFSE 197
Query: 89 LKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNT 148
LKPGFLNSL+D +K EL+QFH+LP FVS SNFDTL+NPVRT AG P ++ LNV S G +
Sbjct: 198 LKPGFLNSLSDGKKLELVQFHVLPDFVSASNFDTLTNPVRTLAGNKPGKVELNVISYGGS 377
Query: 149 VNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVA--KPPAPAPA----------- 195
VN++TG VN T+ G +Y+D LAVY+V KVLLP +F KP APAPA
Sbjct: 378 VNISTGAVNATMNGVIYTDKHLAVYRVGKVLLPSEFVATTKKPLAPAPALKPAPVTSAND 557
Query: 196 ------PEKAKASKK---KSADDTEGPAGDDSAAVSVKQRLVWGPLAVAMVVVAAFSS 244
PEK K S K S T+ S V + +W L + +V V F++
Sbjct: 558 VAEAPKPEKEKPSPKPEQSSESSTQVVPTVTSGGVRIGVSGMWVSLVLGLVFVGVFAA 731
>BQ094318 weakly similar to GP|21553523|gb arabinogalactan protein-like
{Arabidopsis thaliana}, partial (45%)
Length = 421
Score = 169 bits (427), Expect = 1e-42
Identities = 88/140 (62%), Positives = 112/140 (79%), Gaps = 2/140 (1%)
Frame = +1
Query: 65 INAQLINSNAG-LTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTL 123
+N+QL+ S +G LT FAP D+AFS LK GFLNSL+D+QK EL+QFH L +F+S+SNFDTL
Sbjct: 1 LNSQLLTSGSGGLTLFAPEDSAFSKLKAGFLNSLSDRQKVELLQFHTLSSFISISNFDTL 180
Query: 124 SNPVRTQAGENPDRLALNVTS-SGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPR 182
+NPV+TQAG++P RL LNVT+ G+ V+M TG VN +V G+VY+DN+LA+YQVDKVLLP
Sbjct: 181 TNPVQTQAGDDPKRLQLNVTTFGGSQVSMATGAVNASVTGTVYTDNKLAIYQVDKVLLPL 360
Query: 183 DFFVAKPPAPAPAPEKAKAS 202
D V APAPAP KAK +
Sbjct: 361 D-LVLPSEAPAPAPGKAKGA 417
>TC204370 weakly similar to GB|AAM19981.1|20466119|AY098971 AT5g44130/MLN1_5
{Arabidopsis thaliana;} , partial (59%)
Length = 1002
Score = 163 bits (413), Expect = 6e-41
Identities = 103/247 (41%), Positives = 144/247 (57%), Gaps = 8/247 (3%)
Frame = +3
Query: 1 MKHQLFTLPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQ 60
M + FTL LL L F + T A +PAPAPS + ++ IL+K G YTTLI+LL+ TQ
Sbjct: 21 MASKSFTLILLILTTFLMAQTQ--AQAPAPAPSGA-VNLTAILEKGGQYTTLIKLLKDTQ 191
Query: 61 VATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNF 120
TQI +QL +++ G T FAP DNAF +LKPG LN L+D +K +LI FH+ P + ++S+
Sbjct: 192 QLTQIESQLKSNSQGFTLFAPTDNAFQSLKPGALNDLSDDKKVKLILFHVTPKYYTISDL 371
Query: 121 DTLSNPVRTQAGENPDRLALNVT-SSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVL 179
T+SNPVRTQA E LN T GN VN++TG+V + ++ LAVYQVDKVL
Sbjct: 372 LTVSNPVRTQATEEEGTWGLNFTGQGGNQVNISTGVVQTQLNNALREKFPLAVYQVDKVL 551
Query: 180 LPRDFF---VAKPPAPAPAPEKAKASKK----KSADDTEGPAGDDSAAVSVKQRLVWGPL 232
LP + F + AP+P+ +K++ + A P GD + R V L
Sbjct: 552 LPLELFGTTKTTHSSEAPSPKGSKSTPEIPSVGKAGGAPSPHGDKKDTNAANGRNVAFGL 731
Query: 233 AVAMVVV 239
V + ++
Sbjct: 732 VVGLALI 752
>TC204373 weakly similar to GB|AAM19981.1|20466119|AY098971 AT5g44130/MLN1_5
{Arabidopsis thaliana;} , partial (60%)
Length = 1283
Score = 159 bits (402), Expect = 1e-39
Identities = 95/216 (43%), Positives = 133/216 (60%), Gaps = 4/216 (1%)
Frame = +3
Query: 7 TLPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQIN 66
TL LL L F + A +PAP+PS + ++ IL+K G YTTL++LL+ TQ TQI
Sbjct: 69 TLILLTLTTFLIAQAQ--AQAPAPSPSGA-VNLTAILEKGGQYTTLMKLLKDTQQLTQIE 239
Query: 67 AQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNP 126
+QL +++ G T FAP DNAF +LKPG LN L+D QK +LI FH+ P + ++S+ T+SNP
Sbjct: 240 SQLKSNSQGFTLFAPTDNAFQSLKPGALNKLSDDQKVKLILFHVTPKYYTISDLLTVSNP 419
Query: 127 VRTQAGENPDRLALNVT-SSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFF 185
VRTQA E LN T GN VN++TG+V + + LAVYQVDKVLLP + F
Sbjct: 420 VRTQATEKEGTWGLNFTGQGGNQVNISTGVVQTQLNNPLREKFPLAVYQVDKVLLPLELF 599
Query: 186 ---VAKPPAPAPAPEKAKASKKKSADDTEGPAGDDS 218
+ + AP+P+ +K++ + + G A DS
Sbjct: 600 GTTKTRASSAAPSPKGSKSTPEIPSVGKAGSAPSDS 707
>TC217021 similar to UP|Q6J193 (Q6J193) Fasciclin-like AGP 11, partial (48%)
Length = 447
Score = 136 bits (342), Expect = 1e-32
Identities = 75/137 (54%), Positives = 96/137 (69%), Gaps = 1/137 (0%)
Frame = +1
Query: 8 LPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINA 67
LPLL L F T A +PAPA PT+I ++L+KAG +TT I+LL+ +Q+A +IN+
Sbjct: 52 LPLLFLNIFIQ--TISAQVAPAPA---GPTNITQVLEKAGQFTTFIKLLKASQIADRINS 216
Query: 68 QLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPV 127
QL NSN GLT FAP DNAFS+LK G LNS+N Q + +LIQFH+LPT ++S F T SNP+
Sbjct: 217 QLNNSNQGLTVFAPTDNAFSSLKAGTLNSINSQDQMQLIQFHILPTLYTISQFQTASNPL 396
Query: 128 RTQAGENPD-RLALNVT 143
TQAG + D LNVT
Sbjct: 397 HTQAGNSDDGEYPLNVT 447
>TC215415 similar to UP|Q6J192 (Q6J192) Fasciclin-like AGP 12, partial (79%)
Length = 1205
Score = 120 bits (300), Expect = 7e-28
Identities = 78/254 (30%), Positives = 125/254 (48%), Gaps = 13/254 (5%)
Frame = +2
Query: 5 LFTLPLLALMFF--FHSTTTLAASSPAPAPSSSP--TDIIRILKKAGGYTTLIRLLQTTQ 60
+F + + L+F F T + + SP PAP+ +P ++ +L AG + T + L++T+
Sbjct: 239 IFIVSNIMLLFSSAFAKTASPPSLSPTPAPAPAPDFVNLTELLSVAGPFHTFLGYLESTK 418
Query: 61 VATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNF 120
V Q N+ G+T F P D+AF+ +K L++L Q ++I FH LP F S++ F
Sbjct: 419 VIDTFQNQANNTEEGITIFVPKDSAFNAIKKTTLSNLTSNQLKQVILFHALPHFYSLAEF 598
Query: 121 DTLSNPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLL 180
+LS T D LN T TV++ +G V +V++ + +A+YQVDKVLL
Sbjct: 599 TSLSQTSSTPTFAGGD-YTLNFTDDSGTVHINSGWSKTKVSSAVHATDPVAIYQVDKVLL 775
Query: 181 PRDFF---VAKPPAPAPAPEKAKASKKKSADDTEGPAG------DDSAAVSVKQRLVWGP 231
P + PAPAP P+ A A+ + + A DDS++ + +W
Sbjct: 776 PEAILGTNIPPAPAPAPTPDIAPAADSPTEHSADSKASSPSSTHDDSSSHKLISYGIWAN 955
Query: 232 LAVAMVVVAAFSSW 245
L +A V W
Sbjct: 956 LVLATFGVLVLF*W 997
>BE804797
Length = 370
Score = 101 bits (252), Expect = 3e-22
Identities = 57/122 (46%), Positives = 77/122 (62%), Gaps = 1/122 (0%)
Frame = +2
Query: 52 LIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLL 111
L L + Q++ +I QL S+ GLT FA D+AFS+LK LN +N Q + +LIQ H+L
Sbjct: 5 LSELRKA*QISDRIRYQLNISDQGLTGFALTDHAFSSLKARTLNYINSQDQMQLIQIHIL 184
Query: 112 PTFVSMSNFDTLSNPVRTQAGENPD-RLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQL 170
PT ++S T SNP+ TQAG + D LNV + N VN+TTG+V+ TV S+YSDN
Sbjct: 185 PTLYTISQLLTASNPLHTQAGNSDDGEYPLNVNTLWNQVNVTTGVVDTTVSNSIYSDNPT 364
Query: 171 AV 172
V
Sbjct: 365 CV 370
>BI471377
Length = 421
Score = 98.6 bits (244), Expect = 2e-21
Identities = 59/127 (46%), Positives = 77/127 (60%), Gaps = 6/127 (4%)
Frame = +2
Query: 124 SNPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRD 183
SNPV T A ++P LNVT+ GN+VN++TG+VN T+ G VY+D LA+Y VDKVL+P D
Sbjct: 2 SNPVLTLASDSPTGYQLNVTAYGNSVNISTGVVNATLTGIVYTDKTLAIYHVDKVLIPLD 181
Query: 184 FFVAKPPAPAPAPEKAKASKKKSA----DDTEGPAGDDSAAVSV--KQRLVWGPLAVAMV 237
F KP APAPA KA + K++A DD A D S A+S+ Q VA++
Sbjct: 182 FSKPKPIAPAPAVAKAPKADKENASAEDDDQAQAAKDSSGAMSLVSTQGTTLVSFGVALL 361
Query: 238 VVAAFSS 244
AA S
Sbjct: 362 AAAATMS 382
>TC216483 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (31%)
Length = 1084
Score = 91.3 bits (225), Expect = 4e-19
Identities = 63/215 (29%), Positives = 102/215 (47%), Gaps = 16/215 (7%)
Frame = +2
Query: 24 AASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPND 83
+A++ APAP+ + ++ I+ K G L N L + GLT F P D
Sbjct: 107 SAAAEAPAPAPTQQNLTNIMSKHGCKVFADTLSAQPDALNTFNDNL---DGGLTVFCPLD 277
Query: 84 NAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVT 143
+AF P F N L K L++FH +P + S + + + T A + ++ V
Sbjct: 278 DAFKAFLPKFKN-LTKSGKAALLEFHAVPVYQSKATLKSNNGLQNTLATDGANKFDFTVQ 454
Query: 144 SSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPE------ 197
+ G V + T + + ++ ++ LA++ ++KVL P++ F + APAPAPE
Sbjct: 455 NDGEDVTLKTKLTTAKITDTLIDEHPLAIFAINKVLXPKELFKGEAEAPAPAPEPSAADA 634
Query: 198 --KAKASKKKS------ADDTEGPAG--DDSAAVS 222
AK KKK ADD++ PA DD+A V+
Sbjct: 635 PAPAKKGKKKKKAADAPADDSDAPADSPDDAADVT 739
>TC225828 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (41%)
Length = 1108
Score = 90.1 bits (222), Expect = 8e-19
Identities = 58/212 (27%), Positives = 102/212 (47%), Gaps = 14/212 (6%)
Frame = +3
Query: 19 STTTLAASSPAPAPSSSPTDIIRILKKAG--GYTTLIRLLQTTQVATQINAQLINSNAGL 76
S +A + AP + S D+I I+ K G + L+R + + N + GL
Sbjct: 45 SAAISSADAEAPTAAPSAIDLISIMSKQGCKAFADLLRGSKALPAFKE------NVDGGL 206
Query: 77 TFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPD 136
T F P D+A S P + N L + QK L+ +H P + S+ + + + T A E +
Sbjct: 207 TVFCPTDSAISGFAPKYKN-LTEAQKVSLLLYHATPVYESLQMLKSSNGIMNTLATEGAN 383
Query: 137 RLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVA-----KPPA 191
+ V S G V++ T + ++ G++ + Y++++VL+PR+ F A + PA
Sbjct: 384 KYDFTVKSEGEDVSLKTKVNTASIVGTLIDQDPFVAYKINRVLMPRELFKASDALDQAPA 563
Query: 192 PAPAPEKAKASKKKSADDT-------EGPAGD 216
+P P K K + KK ++D+ +GP+ D
Sbjct: 564 ESPKPAKKKKNAKKGSEDSDAADAPADGPSDD 659
>TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (57%)
Length = 1652
Score = 87.8 bits (216), Expect = 4e-18
Identities = 59/208 (28%), Positives = 98/208 (46%), Gaps = 8/208 (3%)
Frame = +1
Query: 19 STTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTF 78
S +A + AP + S D+I I+ K G LL+ ++ N + GLT
Sbjct: 511 SAAISSADAEAPTAAPSAIDLISIMSKQG-CKAFADLLRGSKALPSFKE---NVDGGLTV 678
Query: 79 FAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRL 138
F P D+A S P + N L + QK L+ +H P + S+ + + + T A E ++
Sbjct: 679 FCPTDSAVSGFAPKYKN-LTEAQKVSLLLYHATPVYESLQMLKSSNGIMNTLATEGANKY 855
Query: 139 ALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEK 198
V S G V++ T + ++ G++ + Y++++VL+PR+ F A A APA
Sbjct: 856 DFTVQSEGEDVSLKTKVNTASIVGTLIDQDPFVAYKINRVLMPRELFKASDAADAPAQSP 1035
Query: 199 AKASKKK-------SAD-DTEGPAGDDS 218
A KK +AD +GP+ DD+
Sbjct: 1036KPAKKKNKNKNNSHAADAPADGPSDDDA 1119
>TC227271 similar to GB|AAG24276.1|10880493|AF195889 fasciclin-like
arabinogalactan protein FLA8 {Arabidopsis thaliana;} ,
partial (75%)
Length = 1213
Score = 84.7 bits (208), Expect = 3e-17
Identities = 53/187 (28%), Positives = 94/187 (49%), Gaps = 1/187 (0%)
Frame = +2
Query: 29 APAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSN 88
A P S+ +I +++KAG T L+ + + + ++ GLT FAPND AF
Sbjct: 200 AAPPPSADVNITALIEKAG-CKTFASLISSNGLIKTFQS---TADKGLTIFAPNDEAFKA 367
Query: 89 LKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNT 148
L+ L + + L+Q+H ++ + + T + + T A + L V+ +G++
Sbjct: 368 KGVPDLSKLTNAEVVSLLQYHAAAKYLPVGSLKTTKDSINTLASNGAGKFDLTVSVAGDS 547
Query: 149 VNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKA-SKKKSA 207
+ + TG+ + + ++ L++Y VD VLLP + F AK P+PAPAP A S +
Sbjct: 548 LTLHTGVDSSRIAETILDSTPLSIYSVDSVLLPPELF-AKSPSPAPAPGPVTAPSPSPVS 724
Query: 208 DDTEGPA 214
+E P+
Sbjct: 725 TPSEAPS 745
>TC216484 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (58%)
Length = 1213
Score = 67.0 bits (162), Expect(2) = 3e-14
Identities = 43/169 (25%), Positives = 75/169 (43%)
Frame = +3
Query: 24 AASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPND 83
+A++ APAP+ + ++ I+ K G L N L + GLT F P D
Sbjct: 558 SAAAEAPAPAPTQQNLTNIMSKHGCKVFADALSAQPDALNTFNDNL---DGGLTVFCPLD 728
Query: 84 NAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVT 143
+AF P F N L K L++FH +P + S + + + T A + ++ V
Sbjct: 729 DAFKAFLPKFKN-LTKSGKVALLEFHGVPVYQSKATLKSNNGLQNTLATDGANKFDFTVQ 905
Query: 144 SSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAP 192
+ G V + T + + ++ + LA++ + KVL P++ F P
Sbjct: 906 NDGEDVTLKTKLTTAKITDTLIDEQPLAIFAISKVLQPKELFKGGESXP 1052
Score = 28.1 bits (61), Expect(2) = 3e-14
Identities = 18/38 (47%), Positives = 23/38 (60%), Gaps = 6/38 (15%)
Frame = +2
Query: 191 APAPAPEKAKASKKKS----ADDTEGPAG--DDSAAVS 222
A APAP K KKK+ ADD++ PA DD+A V+
Sbjct: 1076 ADAPAPAKNGKKKKKAADAPADDSDAPADSPDDAADVT 1189
>TC227272 similar to GB|AAK55685.1|14190409|AF378882 AT3g60900/T4C21_310
{Arabidopsis thaliana;} , partial (30%)
Length = 678
Score = 69.3 bits (168), Expect = 1e-12
Identities = 33/121 (27%), Positives = 65/121 (53%)
Frame = +1
Query: 94 LNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNTVNMTT 153
L+ L + + L+Q+H ++ + + T + + T A + L V+ +G+++ + T
Sbjct: 28 LSKLTNAEVVSLLQYHAAAKYLPVGSLKTTKDSINTLASNGAGKFDLTVSVAGDSLTLHT 207
Query: 154 GIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKASKKKSADDTEGP 213
G+ + + ++ + L++Y VD VLLP++ F AK P+PAPAPE + + E P
Sbjct: 208 GVDSSRIADTILDSSPLSIYSVDSVLLPQELF-AKSPSPAPAPESVSSPTPAPSPVAEAP 384
Query: 214 A 214
+
Sbjct: 385 S 387
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.130 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,491,318
Number of Sequences: 63676
Number of extensions: 132649
Number of successful extensions: 2282
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 1867
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2192
length of query: 245
length of database: 12,639,632
effective HSP length: 95
effective length of query: 150
effective length of database: 6,590,412
effective search space: 988561800
effective search space used: 988561800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0136.12


