

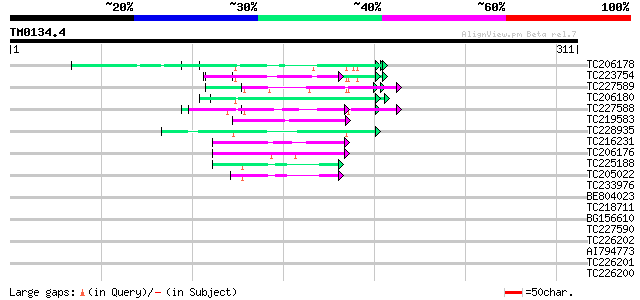
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.4
(311 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 65 5e-11
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 64 1e-10
TC227589 55 3e-08
TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 53 2e-07
TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L1... 53 2e-07
TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich p... 52 4e-07
TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana geno... 50 1e-06
TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (G... 47 1e-05
TC206176 weakly similar to UP|Q8LPA7 (Q8LPA7) Cold shock protein... 44 1e-04
TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like prot... 43 2e-04
TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, pa... 43 2e-04
TC233976 40 0.001
BE804023 33 0.22
TC218711 similar to UP|Q90Z60 (Q90Z60) Rev-Erb beta (Fragment), ... 32 0.28
BG156610 weakly similar to GP|9828614|gb|A F1N21.3 {Arabidopsis ... 32 0.48
TC227590 similar to UP|GRP2_NICSY (P27484) Glycine-rich protein ... 32 0.48
TC226202 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp... 32 0.48
AI794773 similar to GP|11875626|gb XRN4 {Arabidopsis thaliana}, ... 32 0.48
TC226201 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp... 32 0.48
TC226200 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp... 32 0.48
>TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (69%)
Length = 1138
Score = 64.7 bits (156), Expect = 5e-11
Identities = 47/179 (26%), Positives = 71/179 (39%), Gaps = 6/179 (3%)
Frame = +2
Query: 35 RERPQPLIEASKYQQSRLDRAGTGGPVRSGSQEYRQLGPYQHPKGRGFTPRSFKSSTTAT 94
R++ + S+ + R+ +RS YR PY+ RGF+ R
Sbjct: 188 RKKKWKMSSDSRSRSRSRSRSPMDRKIRSDRFSYRD-APYRRDSRRGFS-RDNLCKNCKR 361
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMK 154
PG ++ C CG GH A+ C + C+NC + GH+A +C P++
Sbjct: 362 PGHYARECPNVAICHNCGLPGHIASECTTKS-LCWNCKEPGHMASSC------PNEG--- 511
Query: 155 GKFPAIARGTEKQVSCYRCGEIGHFASRCSATG------PRCFNCHRVGHLAVVCKKPK 207
C+ CG+ GH A CSA C NC++ GH+A C K
Sbjct: 512 --------------ICHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECTNEK 646
Score = 51.6 bits (122), Expect = 4e-07
Identities = 35/103 (33%), Positives = 40/103 (37%), Gaps = 4/103 (3%)
Frame = +2
Query: 105 EVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGT 164
E C C K GH A C P C CN GH+A C D G ARG
Sbjct: 641 EKACNNCRKTGHLARDCPND-PICNLCNVSGHVARQCPKANV-LGDRSGGGGGGGGARGG 814
Query: 165 E----KQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVC 203
+ V C C ++GH + C C NC GHLA C
Sbjct: 815 GGGGYRDVVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 943
Score = 48.9 bits (115), Expect = 3e-06
Identities = 39/143 (27%), Positives = 51/143 (35%), Gaps = 31/143 (21%)
Frame = +2
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANACL------GQGPRCFNCNQIGHLAVNCKTFQAGP 148
PG + S E C CGK GH A C G C NC + GH+A C +A
Sbjct: 476 PGHMASSCPNEGICHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECTNEKACN 655
Query: 149 SDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC---SATGPR---------------- 189
+ K +AR C C GH A +C + G R
Sbjct: 656 NCRKTGH----LARDCPNDPICNLCNVSGHVARQCPKANVLGDRSGGGGGGGGARGGGGG 823
Query: 190 ------CFNCHRVGHLAVVCKKP 206
C NC ++GH++ C P
Sbjct: 824 GYRDVVCRNCQQLGHMSRDCMGP 892
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 63.5 bits (153), Expect = 1e-10
Identities = 38/123 (30%), Positives = 45/123 (35%), Gaps = 27/123 (21%)
Frame = +2
Query: 108 CFRCGKNGHYANACL------------GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKG 155
CF CG GH A C+ G G CF C GH+A +C T + G
Sbjct: 134 CFNCGGFGHLARDCVRGGGGSVGIGGGGGGGSCFRCGGFGHMARDCATGKGNIGGGGSGG 313
Query: 156 KFPAIARGTEKQVSCYRCGEIGHFASRCSATGPR---------------CFNCHRVGHLA 200
C+RCGE+GH A C G R CFNC + GH A
Sbjct: 314 -------------GCFRCGEVGHLARDCGMEGGRFGGGGGSGGGGGKSTCFNCGKPGHFA 454
Query: 201 VVC 203
C
Sbjct: 455 REC 463
Score = 62.0 bits (149), Expect = 3e-10
Identities = 39/115 (33%), Positives = 46/115 (39%), Gaps = 19/115 (16%)
Frame = +2
Query: 108 CFRCGKNGHYANACL----------GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKF 157
C++CG+ GH A C G G CFNC GHLA +C G
Sbjct: 44 CYQCGEFGHLARDCNRSSNSGGGGGGSGGGCFNCGGFGHLARDCVRGGGGSVG------- 202
Query: 158 PAIARGTEKQVSCYRCGEIGHFASRCS---------ATGPRCFNCHRVGHLAVVC 203
G SC+RCG GH A C+ +G CF C VGHLA C
Sbjct: 203 ---IGGGGGGGSCFRCGGFGHMARDCATGKGNIGGGGSGGGCFRCGEVGHLARDC 358
Score = 55.1 bits (131), Expect = 4e-08
Identities = 30/86 (34%), Positives = 40/86 (45%), Gaps = 9/86 (10%)
Frame = +2
Query: 107 TCFRCGKNGHYANACL---------GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKF 157
+CFRCG GH A C G G CF C ++GHLA +C G + G
Sbjct: 227 SCFRCGGFGHMARDCATGKGNIGGGGSGGGCFRCGEVGHLARDC-----GMEGGRFGGGG 391
Query: 158 PAIARGTEKQVSCYRCGEIGHFASRC 183
+ G + +C+ CG+ GHFA C
Sbjct: 392 GSGGGGGKS--TCFNCGKPGHFAREC 463
Score = 44.3 bits (103), Expect = 7e-05
Identities = 29/97 (29%), Positives = 34/97 (34%), Gaps = 12/97 (12%)
Frame = +2
Query: 123 GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASR 182
G G C+ C + GHLA +C S N G G C+ CG GH A
Sbjct: 29 GGGAACYQCGEFGHLARDCNR-----SSNSGGG-------GGGSGGGCFNCGGFGHLARD 172
Query: 183 C------------SATGPRCFNCHRVGHLAVVCKKPK 207
C G CF C GH+A C K
Sbjct: 173 CVRGGGGSVGIGGGGGGGSCFRCGGFGHMARDCATGK 283
Score = 31.2 bits (69), Expect = 0.63
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = +2
Query: 96 GARSQSAHPEVTCFRCGKNGHYANACL 122
G S + TCF CGK GH+A C+
Sbjct: 386 GGGSGGGGGKSTCFNCGKPGHFARECV 466
>TC227589
Length = 547
Score = 55.5 bits (132), Expect = 3e-08
Identities = 37/125 (29%), Positives = 46/125 (36%), Gaps = 26/125 (20%)
Frame = +2
Query: 108 CFRCGKNGHYANACLGQGPR-------CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAI 160
CF C K GH A CL + C C GH +C+ P D K +
Sbjct: 170 CFICKKGGHRAKDCLEKHTSRSKSVAICLKCGNSGHDMFSCRN-DYSPDDLKEIQCYVCK 346
Query: 161 ARG----------TEKQVSCYRCGEIGHFASRCS---------ATGPRCFNCHRVGHLAV 201
G T ++SCY+CG++GH CS AT C C GH A
Sbjct: 347 RVGHLCCVNTDDATPGEISCYKCGQLGHTGLACSKLPDEITSAATPSSCCKCGEAGHFAQ 526
Query: 202 VCKKP 206
C P
Sbjct: 527 ECTSP 541
Score = 48.1 bits (113), Expect = 5e-06
Identities = 27/88 (30%), Positives = 37/88 (41%)
Frame = +2
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG 187
CFNC + GH AVNC +++ CY CG +GH A +C+
Sbjct: 47 CFNCGEDGHAAVNCS--------------------AAKRKKPCYVCGGLGHNARQCT-KA 163
Query: 188 PRCFNCHRVGHLAVVCKKPKVEPSVNTA 215
CF C + GH A C + S + A
Sbjct: 164 QDCFICKKGGHRAKDCLEKHTSRSKSVA 247
Score = 47.0 bits (110), Expect = 1e-05
Identities = 30/108 (27%), Positives = 42/108 (38%), Gaps = 13/108 (12%)
Frame = +2
Query: 108 CFRCGKNGHYANACLGQGPR--CFNCNQIGHLAVNC----KTFQAGPSDNKMKGKFPAIA 161
CF CG++GH A C + C+ C +GH A C F ++ K
Sbjct: 47 CFNCGEDGHAAVNCSAAKRKKPCYVCGGLGHNARQCTKAQDCFICKKGGHRAKDCLEKHT 226
Query: 162 RGTEKQVSCYRCGEIGHFASRC-------SATGPRCFNCHRVGHLAVV 202
++ C +CG GH C +C+ C RVGHL V
Sbjct: 227 SRSKSVAICLKCGNSGHDMFSCRNDYSPDDLKEIQCYVCKRVGHLCCV 370
Score = 37.0 bits (84), Expect = 0.011
Identities = 17/39 (43%), Positives = 22/39 (55%), Gaps = 2/39 (5%)
Frame = +2
Query: 169 SCYRCGEIGHFASRCSATGPR--CFNCHRVGHLAVVCKK 205
+C+ CGE GH A CSA + C+ C +GH A C K
Sbjct: 44 ACFNCGEDGHAAVNCSAAKRKKPCYVCGGLGHNARQCTK 160
Score = 37.0 bits (84), Expect = 0.011
Identities = 24/93 (25%), Positives = 32/93 (33%)
Frame = +2
Query: 93 ATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNK 152
ATPG E++C++CG+ GH AC
Sbjct: 383 ATPG--------EISCYKCGQLGHTGLAC------------------------------- 445
Query: 153 MKGKFPAIARGTEKQVSCYRCGEIGHFASRCSA 185
K P SC +CGE GHFA C++
Sbjct: 446 --SKLPDEITSAATPSSCCKCGEAGHFAQECTS 538
>TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (42%)
Length = 655
Score = 53.1 bits (126), Expect = 2e-07
Identities = 32/99 (32%), Positives = 37/99 (37%)
Frame = +3
Query: 105 EVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGT 164
E C C K GH A C P C CN GH+A C G G
Sbjct: 126 EKACNNCRKTGHLARDCPND-PICNLCNVSGHVARQCPKANVLGDXXGGGGGARGGGGGG 302
Query: 165 EKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVC 203
+ V C C ++GH + C C NC GHLA C
Sbjct: 303 YRDVVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 419
Score = 43.1 bits (100), Expect = 2e-04
Identities = 31/104 (29%), Positives = 37/104 (34%), Gaps = 6/104 (5%)
Frame = +3
Query: 111 CGKNGHYANACL------GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGT 164
CGK GH A C G C NC + GH+A C +A
Sbjct: 9 CGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECTNEKA------------------ 134
Query: 165 EKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKV 208
C C + GH A C P C C+ GH+A C K V
Sbjct: 135 -----CNNCRKTGHLARDC-PNDPICNLCNVSGHVARQCPKANV 248
Score = 33.9 bits (76), Expect = 0.097
Identities = 16/43 (37%), Positives = 21/43 (48%), Gaps = 6/43 (13%)
Frame = +3
Query: 171 YRCGEIGHFASRCSATGPR------CFNCHRVGHLAVVCKKPK 207
+ CG+ GH A CSA C NC++ GH+A C K
Sbjct: 3 HTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECTNEK 131
>TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L16.11 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(10%)
Length = 1300
Score = 52.8 bits (125), Expect = 2e-07
Identities = 33/125 (26%), Positives = 48/125 (38%), Gaps = 16/125 (12%)
Frame = +1
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANACLGQGPR-------CFNCNQIGHLAVNCKTFQAG 147
P + ++ C +CG +GH +C + C+ C ++GHL T A
Sbjct: 226 PEKHTSTSKSIAICLKCGNSGHDIFSCRNDYSQDDLKEIQCYVCKRLGHLCC-VNTDDAT 402
Query: 148 PSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCS---------ATGPRCFNCHRVGH 198
P + +SCY+CG++GH CS AT CF C GH
Sbjct: 403 PGE-----------------ISCYKCGQLGHTGLACSRLRDEITSGATPSSCFKCGEEGH 531
Query: 199 LAVVC 203
A C
Sbjct: 532 FAREC 546
Score = 49.3 bits (116), Expect = 2e-06
Identities = 31/88 (35%), Positives = 39/88 (44%)
Frame = +1
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG 187
CFNC + GH AVNC + +K K P CY CG +GH A +CS
Sbjct: 61 CFNCGEEGHAAVNC---------SAVKRKKP-----------CYVCGCLGHNARQCSKV- 177
Query: 188 PRCFNCHRVGHLAVVCKKPKVEPSVNTA 215
CF C + GH A C + S + A
Sbjct: 178 QDCFICKKGGHRAKDCPEKHTSTSKSIA 261
Score = 47.8 bits (112), Expect = 6e-06
Identities = 28/93 (30%), Positives = 43/93 (46%), Gaps = 5/93 (5%)
Frame = +1
Query: 99 SQSAHPEVTCFRCGKNGHYA-----NACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKM 153
SQ E+ C+ C + GH +A G+ C+ C Q+GH + C + D
Sbjct: 319 SQDDLKEIQCYVCKRLGHLCCVNTDDATPGE-ISCYKCGQLGHTGLACSRLR----DEIT 483
Query: 154 KGKFPAIARGTEKQVSCYRCGEIGHFASRCSAT 186
G P+ SC++CGE GHFA C+++
Sbjct: 484 SGATPS---------SCFKCGEEGHFARECTSS 555
Score = 37.4 bits (85), Expect = 0.009
Identities = 21/55 (38%), Positives = 28/55 (50%), Gaps = 2/55 (3%)
Frame = +1
Query: 169 SCYRCGEIGHFASRCSATGPR--CFNCHRVGHLAVVCKKPKVEPSVNTARGKHPA 221
+C+ CGE GH A CSA + C+ C +GH A C KV+ +G H A
Sbjct: 58 ACFNCGEEGHAAVNCSAVKRKKPCYVCGCLGHNARQCS--KVQDCFICKKGGHRA 216
>TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich protein 2,
partial (45%)
Length = 995
Score = 51.6 bits (122), Expect = 4e-07
Identities = 26/65 (40%), Positives = 32/65 (49%)
Frame = +2
Query: 123 GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASR 182
G GP C+NC +IGHLA +C Q G + G+ G C+ CGE GHFA
Sbjct: 410 GGGPECYNCGRIGHLARDCYHGQGGGGGD--DGRNRRRGGGGGGGGGCFNCGEEGHFARE 583
Query: 183 CSATG 187
C G
Sbjct: 584 CPNVG 598
Score = 37.0 bits (84), Expect = 0.011
Identities = 35/129 (27%), Positives = 44/129 (33%), Gaps = 23/129 (17%)
Frame = +2
Query: 42 IEASKYQQSRLD---RAGTGGPVRSGSQEYRQLGPYQHPKGRGFTPRSFKSSTTATPGAR 98
++ + +SRL R G GG R G G Y +GRG G
Sbjct: 284 VDVTSAVRSRLPGGFRGGGGGRGRGG-------GRYGGGEGRG-------------RGFG 403
Query: 99 SQSAHPEVTCFRCGKNGHYANACL--------------------GQGPRCFNCNQIGHLA 138
+ PE C+ CG+ GH A C G G CFNC + GH A
Sbjct: 404 RRGGGPE--CYNCGRIGHLARDCYHGQGGGGGDDGRNRRRGGGGGGGGGCFNCGEEGHFA 577
Query: 139 VNCKTFQAG 147
C G
Sbjct: 578 RECPNVGKG 604
Score = 33.9 bits (76), Expect = 0.097
Identities = 34/127 (26%), Positives = 52/127 (40%), Gaps = 18/127 (14%)
Frame = +2
Query: 187 GPRCFNCHRVGHLAVVCKKPK----VEPSVNTARGKHPAARGKVYTMDGEEAEGADELVK 242
GP C+NC R+GHLA C + + N RG G + GEE A E
Sbjct: 416 GPECYNCGRIGHLARDCYHGQGGGGGDDGRNRRRGGGGGGGGGCFNC-GEEGHFARECPN 592
Query: 243 GERKNDGNLLTILSHSSIIRSITLVAHV----------THLFYL----YCLSIDFIVITP 288
+ N+ + + SSI S+ L+ V +H +L C S +++
Sbjct: 593 VGKGNE*ECVMV---SSIELSLLLLVGVG*MDSVIFVSSHWHFLPSFQICFSRALLIVLS 763
Query: 289 NNTLFDR 295
N LF++
Sbjct: 764 NALLFNK 784
>TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K24M7, partial (36%)
Length = 1224
Score = 50.4 bits (119), Expect = 1e-06
Identities = 36/132 (27%), Positives = 47/132 (35%), Gaps = 12/132 (9%)
Frame = +1
Query: 84 PRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANAC-----LGQGPRCFNCNQIGHLA 138
P S K PG + P +CF C H A C + C C + GH A
Sbjct: 214 PGSRKRHLLRVPGMK-----PGESCFICKAMDHIAKLCPEKAEWEKNKICLRCRRRGHRA 378
Query: 139 VNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC-------SATGPRCF 191
NC P + G + CY CGE GH ++C CF
Sbjct: 379 KNC----------------PEVLDGAKDAKYCYNCGENGHALTQCLHPLQEGGTKFAECF 510
Query: 192 NCHRVGHLAVVC 203
C++ GHL+ C
Sbjct: 511 VCNQRGHLSKNC 546
Score = 35.4 bits (80), Expect = 0.033
Identities = 25/91 (27%), Positives = 31/91 (33%), Gaps = 6/91 (6%)
Frame = +1
Query: 125 GPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC- 183
G CF C + H+A C K ++ EK C RC GH A C
Sbjct: 262 GESCFICKAMDHIAKLCPE----------KAEW-------EKNKICLRCRRRGHRAKNCP 390
Query: 184 -----SATGPRCFNCHRVGHLAVVCKKPKVE 209
+ C+NC GH C P E
Sbjct: 391 EVLDGAKDAKYCYNCGENGHALTQCLHPLQE 483
>TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (GRP2)
(AT4g38680/F20M13_240), partial (42%)
Length = 891
Score = 46.6 bits (109), Expect = 1e-05
Identities = 26/75 (34%), Positives = 33/75 (43%)
Frame = +3
Query: 112 GKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCY 171
G G Y G G C+ C + GH+A +C Q G + G G SCY
Sbjct: 183 GGGGGYGGGGGGGGGGCYKCGETGHIARDCS--QGGGGGGRYGG-------GGGGGGSCY 335
Query: 172 RCGEIGHFASRCSAT 186
CGE GHFA C ++
Sbjct: 336 NCGESGHFARDCPSS 380
Score = 38.1 bits (87), Expect = 0.005
Identities = 16/47 (34%), Positives = 22/47 (46%), Gaps = 13/47 (27%)
Frame = +3
Query: 108 CFRCGKNGHYANACL-------------GQGPRCFNCNQIGHLAVNC 141
C++CG+ GH A C G G C+NC + GH A +C
Sbjct: 231 CYKCGETGHIARDCSQGGGGGGRYGGGGGGGGSCYNCGESGHFARDC 371
Score = 37.7 bits (86), Expect = 0.007
Identities = 18/47 (38%), Positives = 20/47 (42%), Gaps = 13/47 (27%)
Frame = +3
Query: 170 CYRCGEIGHFASRCS-------------ATGPRCFNCHRVGHLAVVC 203
CY+CGE GH A CS G C+NC GH A C
Sbjct: 231 CYKCGETGHIARDCSQGGGGGGRYGGGGGGGGSCYNCGESGHFARDC 371
>TC206176 weakly similar to UP|Q8LPA7 (Q8LPA7) Cold shock protein-1, partial
(77%)
Length = 883
Score = 43.5 bits (101), Expect = 1e-04
Identities = 26/85 (30%), Positives = 36/85 (41%), Gaps = 10/85 (11%)
Frame = +3
Query: 112 GKNGHYANACLGQGPRCFNCNQIGHLAVNCK------TFQAGPSDNKMKG----KFPAIA 161
G G+ G G C+NC + GHLA +C + G + G ++
Sbjct: 387 GGGGYGGGGRGGGGGACYNCGESGHLARDCSGGGGGDRYGGGGGGGRYGGGGGGRYVGGG 566
Query: 162 RGTEKQVSCYRCGEIGHFASRCSAT 186
G SCY CGE GHFA C ++
Sbjct: 567 GGGGGGGSCYSCGESGHFARDCPSS 641
Score = 30.4 bits (67), Expect = 1.1
Identities = 11/19 (57%), Positives = 12/19 (62%)
Frame = +3
Query: 169 SCYRCGEIGHFASRCSATG 187
+CY CGE GH A CS G
Sbjct: 432 ACYNCGESGHLARDCSGGG 488
>TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like protein, partial
(62%)
Length = 1131
Score = 43.1 bits (100), Expect = 2e-04
Identities = 28/77 (36%), Positives = 31/77 (39%), Gaps = 5/77 (6%)
Frame = +1
Query: 112 GKNGHYANACLGQGP-----RCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEK 166
G G +G+GP RCFNC GH A +CK AG NK
Sbjct: 301 GPRGSRDREYMGRGPPPGSGRCFNCGIDGHWARDCK---AGDWKNK-------------- 429
Query: 167 QVSCYRCGEIGHFASRC 183
CYRCGE GH C
Sbjct: 430 ---CYRCGERGHIEKNC 471
Score = 40.4 bits (93), Expect = 0.001
Identities = 26/87 (29%), Positives = 33/87 (37%), Gaps = 2/87 (2%)
Frame = +1
Query: 58 GGPVRSGSQEYRQLGPYQHPKGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHY 117
GGP S +EY GP P G G CF CG +GH+
Sbjct: 298 GGPRGSRDREYMGRGP---PPGSG-------------------------RCFNCGIDGHW 393
Query: 118 ANACLGQG--PRCFNCNQIGHLAVNCK 142
A C +C+ C + GH+ NCK
Sbjct: 394 ARDCKAGDWKNKCYRCGERGHIEKNCK 474
Score = 35.4 bits (80), Expect = 0.033
Identities = 22/74 (29%), Positives = 31/74 (41%), Gaps = 2/74 (2%)
Frame = +1
Query: 147 GPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG--PRCFNCHRVGHLAVVCK 204
G D + G+ P G C+ CG GH+A C A +C+ C GH+ CK
Sbjct: 310 GSRDREYMGRGPPPGSGR-----CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 474
Query: 205 KPKVEPSVNTARGK 218
P + RG+
Sbjct: 475 N---SPKKLSQRGR 507
>TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (69%)
Length = 1623
Score = 42.7 bits (99), Expect = 2e-04
Identities = 27/67 (40%), Positives = 29/67 (42%), Gaps = 5/67 (7%)
Frame = +2
Query: 122 LGQGP-----RCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEI 176
LG+GP RCFNC GH A +CK AG NK CYRCGE
Sbjct: 281 LGRGPPPGSGRCFNCGLDGHWARDCK---AGDWKNK-----------------CYRCGER 400
Query: 177 GHFASRC 183
GH C
Sbjct: 401 GHIERNC 421
Score = 39.3 bits (90), Expect = 0.002
Identities = 15/37 (40%), Positives = 21/37 (56%), Gaps = 2/37 (5%)
Frame = +2
Query: 108 CFRCGKNGHYANACLGQG--PRCFNCNQIGHLAVNCK 142
CF CG +GH+A C +C+ C + GH+ NCK
Sbjct: 314 CFNCGLDGHWARDCKAGDWKNKCYRCGERGHIERNCK 424
Score = 34.7 bits (78), Expect = 0.057
Identities = 19/52 (36%), Positives = 27/52 (51%), Gaps = 3/52 (5%)
Frame = +2
Query: 170 CYRCGEIGHFASRCSATG--PRCFNCHRVGHLAVVCK-KPKVEPSVNTARGK 218
C+ CG GH+A C A +C+ C GH+ CK PK ++T RG+
Sbjct: 314 CFNCGLDGHWARDCKAGDWKNKCYRCGERGHIERNCKNSPK---KLSTRRGR 460
>TC233976
Length = 763
Score = 40.0 bits (92), Expect = 0.001
Identities = 16/40 (40%), Positives = 21/40 (52%)
Frame = +1
Query: 170 CYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVE 209
C +CG +GH A C+ CFN GHL+ C P+ E
Sbjct: 271 CRKCGRLGHNAHECTYREVTCFNYQGKGHLSTNCPHPRKE 390
Score = 40.0 bits (92), Expect = 0.001
Identities = 19/52 (36%), Positives = 25/52 (47%), Gaps = 5/52 (9%)
Frame = +1
Query: 95 PGARSQSAHPEVT-----CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNC 141
P R S P + C +CG+ GH A+ C + CFN GHL+ NC
Sbjct: 217 PAGRGGSGAPAIVTTPLRCRKCGRLGHNAHECTYREVTCFNYQGKGHLSTNC 372
>BE804023
Length = 407
Score = 32.7 bits (73), Expect = 0.22
Identities = 17/47 (36%), Positives = 25/47 (53%), Gaps = 3/47 (6%)
Frame = -3
Query: 161 ARGTEKQV--SCYRCGEIGHFASRC-SATGPRCFNCHRVGHLAVVCK 204
A+G K+ SC C + GH RC +C C+++GH V+CK
Sbjct: 252 AKGNWKKFYPSC*HCRKKGHPPFRC*KRPDAKCSQCNQMGHEVVICK 112
Score = 32.7 bits (73), Expect = 0.22
Identities = 19/55 (34%), Positives = 26/55 (46%), Gaps = 1/55 (1%)
Frame = -3
Query: 89 SSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQ-GPRCFNCNQIGHLAVNCK 142
SST+ + +P +C C K GH C + +C CNQ+GH V CK
Sbjct: 270 SSTSTNAKGNWKKFYP--SC*HCRKKGHPPFRC*KRPDAKCSQCNQMGHEVVICK 112
>TC218711 similar to UP|Q90Z60 (Q90Z60) Rev-Erb beta (Fragment), partial (6%)
Length = 1204
Score = 32.3 bits (72), Expect = 0.28
Identities = 25/86 (29%), Positives = 29/86 (33%), Gaps = 6/86 (6%)
Frame = -3
Query: 108 CFRCGKNGHYANAC-----LGQGPR-CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIA 161
C C + GH C L CF C +IGH C QAG G+F
Sbjct: 521 CRACRQPGHRFQQCQRLKCLSMDEEVCFFCGEIGHSLGKCDVSQAG------GGRF---- 372
Query: 162 RGTEKQVSCYRCGEIGHFASRCSATG 187
C C GHF+ C G
Sbjct: 371 ------AKCLLCYGHGHFSYNCPQNG 312
Score = 31.2 bits (69), Expect = 0.63
Identities = 20/69 (28%), Positives = 25/69 (35%), Gaps = 13/69 (18%)
Frame = -3
Query: 86 SFKSSTTATPGARSQSAH-------PEVTCFRCGKNGHYANAC------LGQGPRCFNCN 132
S + PG R Q E CF CG+ GH C G+ +C C
Sbjct: 530 SLRCRACRQPGHRFQQCQRLKCLSMDEEVCFFCGEIGHSLGKCDVSQAGGGRFAKCLLCY 351
Query: 133 QIGHLAVNC 141
GH + NC
Sbjct: 350 GHGHFSYNC 324
>BG156610 weakly similar to GP|9828614|gb|A F1N21.3 {Arabidopsis thaliana},
partial (23%)
Length = 293
Score = 31.6 bits (70), Expect = 0.48
Identities = 14/37 (37%), Positives = 19/37 (50%)
Frame = +1
Query: 189 RCFNCHRVGHLAVVCKKPKVEPSVNTARGKHPAARGK 225
RCFNC H C +P+ +VN AR K + R +
Sbjct: 49 RCFNCGSYNHSLRECPRPRDNIAVNNARDKLKSRRNQ 159
>TC227590 similar to UP|GRP2_NICSY (P27484) Glycine-rich protein 2, partial
(7%)
Length = 586
Score = 31.6 bits (70), Expect = 0.48
Identities = 10/18 (55%), Positives = 15/18 (82%)
Frame = +3
Query: 169 SCYRCGEIGHFASRCSAT 186
SC++CGE GHFA C+++
Sbjct: 6 SCFKCGEEGHFARECTSS 59
Score = 28.9 bits (63), Expect = 3.1
Identities = 8/15 (53%), Positives = 12/15 (79%)
Frame = +3
Query: 107 TCFRCGKNGHYANAC 121
+CF+CG+ GH+A C
Sbjct: 6 SCFKCGEEGHFAREC 50
>TC226202 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp22 splicing
factor), partial (72%)
Length = 1003
Score = 31.6 bits (70), Expect = 0.48
Identities = 12/26 (46%), Positives = 13/26 (49%)
Frame = +3
Query: 162 RGTEKQVSCYRCGEIGHFASRCSATG 187
R + CY CGE GHFA C G
Sbjct: 810 RSGGSDLKCYECGEPGHFARECRMRG 887
Score = 29.3 bits (64), Expect = 2.4
Identities = 10/30 (33%), Positives = 17/30 (56%)
Frame = +3
Query: 96 GARSQSAHPEVTCFRCGKNGHYANACLGQG 125
G +S ++ C+ CG+ GH+A C +G
Sbjct: 798 GRGGRSGGSDLKCYECGEPGHFARECRMRG 887
>AI794773 similar to GP|11875626|gb XRN4 {Arabidopsis thaliana}, partial
(14%)
Length = 440
Score = 31.6 bits (70), Expect = 0.48
Identities = 10/20 (50%), Positives = 13/20 (65%)
Frame = +1
Query: 123 GQGPRCFNCNQIGHLAVNCK 142
GQ +CF C Q+GH A C+
Sbjct: 31 GQQEKCFQCGQLGHFAAECR 90
Score = 31.6 bits (70), Expect = 0.48
Identities = 9/18 (50%), Positives = 15/18 (83%)
Frame = +1
Query: 166 KQVSCYRCGEIGHFASRC 183
+Q C++CG++GHFA+ C
Sbjct: 34 QQEKCFQCGQLGHFAAEC 87
Score = 31.2 bits (69), Expect = 0.63
Identities = 13/39 (33%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Frame = +1
Query: 108 CFRCGKNGHYANACLGQ-GPRCFNCNQIGHLAVNCKTFQ 145
CF+CG+ GH+A C G+ G + + N + ++ K +Q
Sbjct: 46 CFQCGQLGHFAAECRGKPGEKAEDWNPVDDTPIHKKKYQ 162
>TC226201 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp22 splicing
factor), partial (72%)
Length = 820
Score = 31.6 bits (70), Expect = 0.48
Identities = 12/26 (46%), Positives = 13/26 (49%)
Frame = +1
Query: 162 RGTEKQVSCYRCGEIGHFASRCSATG 187
R + CY CGE GHFA C G
Sbjct: 223 RSGGSDLKCYECGEPGHFARECRMRG 300
Score = 29.3 bits (64), Expect = 2.4
Identities = 10/30 (33%), Positives = 17/30 (56%)
Frame = +1
Query: 96 GARSQSAHPEVTCFRCGKNGHYANACLGQG 125
G +S ++ C+ CG+ GH+A C +G
Sbjct: 211 GRGGRSGGSDLKCYECGEPGHFARECRMRG 300
>TC226200 similar to UP|O81126 (O81126) 9G8-like SR protein (RSZp22 splicing
factor), partial (89%)
Length = 936
Score = 31.6 bits (70), Expect = 0.48
Identities = 12/26 (46%), Positives = 13/26 (49%)
Frame = +2
Query: 162 RGTEKQVSCYRCGEIGHFASRCSATG 187
R + CY CGE GHFA C G
Sbjct: 314 RSGGSDLKCYECGEPGHFARECRMRG 391
Score = 29.3 bits (64), Expect = 2.4
Identities = 10/30 (33%), Positives = 17/30 (56%)
Frame = +2
Query: 96 GARSQSAHPEVTCFRCGKNGHYANACLGQG 125
G +S ++ C+ CG+ GH+A C +G
Sbjct: 302 GRGGRSGGSDLKCYECGEPGHFARECRMRG 391
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.134 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,099,586
Number of Sequences: 63676
Number of extensions: 238083
Number of successful extensions: 1588
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 1348
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1503
length of query: 311
length of database: 12,639,632
effective HSP length: 97
effective length of query: 214
effective length of database: 6,463,060
effective search space: 1383094840
effective search space used: 1383094840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0134.4


