

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.14
(277 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
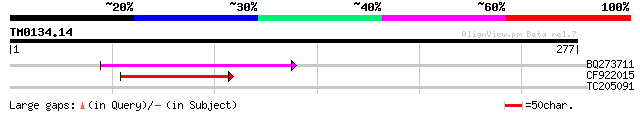
Score E
Sequences producing significant alignments: (bits) Value
BQ273711 77 7e-15
CF922015 55 3e-08
TC205091 weakly similar to UP|Q6TXI7 (Q6TXI7) LRRGT00012, partia... 29 2.0
>BQ273711
Length = 409
Score = 77.4 bits (189), Expect = 7e-15
Identities = 41/96 (42%), Positives = 56/96 (57%)
Frame = -2
Query: 45 QREVALDQNRGLNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYL 104
+R+V + RGL FR+ PPKF+G DP+ A L + E EKIFE + E KV AT++
Sbjct: 288 ERDVEPAEYRGLMAFRKNHPPKFSGDYDPEGARLWLAETEKIFEAMGCLEEHKVLYATFM 109
Query: 105 LLGDAEYWWKGTRGIMRANHEEVN*NSFRTAFLEKY 140
L G+AE WWK + A + N+F+ FLE Y
Sbjct: 108 LQGEAENWWKFVKPSFVAPGGVIPWNAFKDKFLENY 1
>CF922015
Length = 172
Score = 55.1 bits (131), Expect = 3e-08
Identities = 25/55 (45%), Positives = 38/55 (68%)
Frame = -3
Query: 55 GLNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDA 109
GL+ F+R +PP F GG DP+ A+ ++EIEKIF V++ + KV AT++L +A
Sbjct: 167 GLDRFQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADEA 3
>TC205091 weakly similar to UP|Q6TXI7 (Q6TXI7) LRRGT00012, partial (5%)
Length = 1360
Score = 29.3 bits (64), Expect = 2.0
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +1
Query: 82 EIEKIFEVLQTAEGAKVSLATYLLLGDAEYWW 113
++E++F +E KV LAT G A YWW
Sbjct: 1192 KVEQLFACHHISEERKVPLATLSFQGYALYWW 1287
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.138 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,923,192
Number of Sequences: 63676
Number of extensions: 95485
Number of successful extensions: 569
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 568
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 569
length of query: 277
length of database: 12,639,632
effective HSP length: 96
effective length of query: 181
effective length of database: 6,526,736
effective search space: 1181339216
effective search space used: 1181339216
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0134.14


