

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0133.3
(700 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
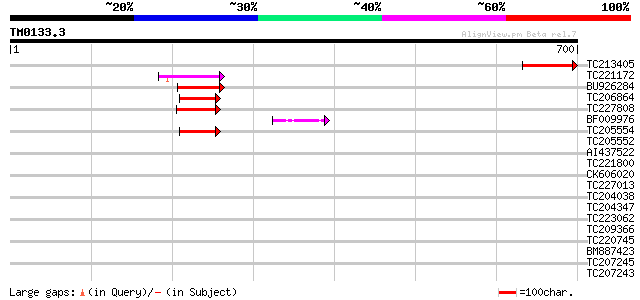
Score E
Sequences producing significant alignments: (bits) Value
TC213405 similar to UP|Q96280 (Q96280) Orf12 protein, partial (8%) 116 3e-26
TC221172 55 8e-08
BU926284 51 2e-06
TC206864 similar to UP|Q9SFU7 (Q9SFU7) T1B9.17 protein (AT3g0717... 49 1e-05
TC227808 similar to UP|Q9SFU7 (Q9SFU7) T1B9.17 protein (AT3g0717... 48 2e-05
BF009976 45 1e-04
TC205554 44 3e-04
TC205552 similar to UP|PMM_ARATH (O80840) Probable phosphomannom... 41 0.002
AI437522 41 0.002
TC221800 PRF|1503276A.0|226263|1503276A chlorophyll a/b binding ... 38 0.013
CK606020 32 0.96
TC227013 similar to UP|GAC1_YEAST (P28006) Protein phosphatase 1... 32 1.2
TC204038 similar to UP|Q8VZC0 (Q8VZC0) DTDP-glucose 4-6-dehydrat... 32 1.2
TC204347 homologue to UP|Q9SP48 (Q9SP48) Homeodomain-leucine zip... 30 2.8
TC223062 similar to UP|Q96280 (Q96280) Orf12 protein, partial (8%) 30 2.8
TC209366 homologue to UP|Q6T7B6 (Q6T7B6) C-type lectin-2, partia... 30 2.8
TC220745 similar to UP|Q9LHD1 (Q9LHD1) Multidrug resistance p-gl... 30 3.6
BM887423 weakly similar to GP|14141201|gb| SRp46 splicing factor... 30 4.7
TC207245 similar to UP|Q75LK9 (Q75LK9) Expressed protein, partia... 29 6.2
TC207243 similar to UP|Q93Z55 (Q93Z55) AT3g58470/F14P22_60, part... 29 6.2
>TC213405 similar to UP|Q96280 (Q96280) Orf12 protein, partial (8%)
Length = 518
Score = 116 bits (291), Expect = 3e-26
Identities = 55/67 (82%), Positives = 61/67 (90%)
Frame = +3
Query: 634 FGKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANA 693
FGKGKKKS GRRWQ GTIIRYEVPYSEH SFTELK+FV+ +SP NIIPSVNNDGPES++A
Sbjct: 3 FGKGKKKSTGRRWQQGTIIRYEVPYSEHSSFTELKEFVRVVSPDNIIPSVNNDGPESSDA 182
Query: 694 MISLMSS 700
MISL+ S
Sbjct: 183 MISLLLS 203
>TC221172
Length = 910
Score = 55.5 bits (132), Expect = 8e-08
Identities = 34/87 (39%), Positives = 50/87 (57%), Gaps = 5/87 (5%)
Frame = +3
Query: 184 LDRGDDVAVR---DDNVDAQFAP--KVSRVVDWLRDLGLGKYEDVFVREEVDWDTLQWLT 238
+DR +D VR +D V+++ K V WL +LGL +Y +F EVD + L LT
Sbjct: 516 VDRNEDPRVRVSENDGVESESRERRKSDGVRSWLYELGLSRYAPMFEIHEVDDELLPMLT 695
Query: 239 EEDLLSMGVTALGPRKKIVHALCEFRK 265
EDL MG+ A+G R+K+ A+ + RK
Sbjct: 696 LEDLKDMGINAVGSRRKMYTAIQKLRK 776
>BU926284
Length = 265
Score = 50.8 bits (120), Expect = 2e-06
Identities = 26/58 (44%), Positives = 36/58 (61%)
Frame = +1
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFRK 265
V WL +LGL +Y +F EVD + L LT EDL MG+ A+G R+K+ A+ + RK
Sbjct: 10 VRSWLYELGLSRYAPMFEIHEVDDELLPMLTLEDLKDMGINAVGSRRKMYTAIQKLRK 183
>TC206864 similar to UP|Q9SFU7 (Q9SFU7) T1B9.17 protein (AT3g07170/T1B9_17),
partial (67%)
Length = 1113
Score = 48.5 bits (114), Expect = 1e-05
Identities = 27/51 (52%), Positives = 35/51 (67%)
Frame = +3
Query: 210 DWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
D+L+ LGL KY F EEVD L +T+EDL +MG+ +GPRKKI+ AL
Sbjct: 639 DFLQSLGLEKYLLSFQAEEVDMTALNHMTDEDLKAMGI-PMGPRKKILLAL 788
>TC227808 similar to UP|Q9SFU7 (Q9SFU7) T1B9.17 protein (AT3g07170/T1B9_17),
partial (53%)
Length = 1017
Score = 47.8 bits (112), Expect = 2e-05
Identities = 28/55 (50%), Positives = 36/55 (64%)
Frame = +3
Query: 206 SRVVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
S V ++LR L L KY F EEVD L +T+EDL +MG+ +GPRKKI+ AL
Sbjct: 657 SSVDEFLRSLCLEKYLITFQAEEVDMTALNHMTDEDLKAMGI-PMGPRKKILLAL 818
>BF009976
Length = 394
Score = 44.7 bits (104), Expect = 1e-04
Identities = 29/73 (39%), Positives = 41/73 (55%), Gaps = 2/73 (2%)
Frame = +1
Query: 325 KVSTAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDVPKWCSIEGTPFRVDAFKY--LRG 382
+ S P+ +E +++ S R KG + NR R P + + GT F VDAF+Y + G
Sbjct: 91 RCSIFPDTGKEFENAKS--SRKRKG-SCGENRVTRSCPFYKKMPGTMFTVDAFRYGCVEG 261
Query: 383 DCSHWFLTHFHLD 395
CS +FLTHFH D
Sbjct: 262 -CSAYFLTHFHCD 297
>TC205554
Length = 635
Score = 43.5 bits (101), Expect = 3e-04
Identities = 20/51 (39%), Positives = 32/51 (62%)
Frame = +3
Query: 210 DWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
+WL LGLG++ +F + L LT + L MG +A+GPR+K++HA+
Sbjct: 279 NWLTSLGLGQFVRIFQGNSLGKYQLVNLTMKKLKDMGASAVGPRRKLIHAM 431
>TC205552 similar to UP|PMM_ARATH (O80840) Probable phosphomannomutase (PMM)
, partial (25%)
Length = 1033
Score = 41.2 bits (95), Expect = 0.002
Identities = 19/50 (38%), Positives = 30/50 (60%)
Frame = +3
Query: 211 WLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
WL LGL ++ +F + L LT + L MG +A+GPR+K++HA+
Sbjct: 414 WLTSLGLAQFVRIFQGNSLSKYQLVNLTMKKLKDMGASAVGPRRKLIHAM 563
>AI437522
Length = 181
Score = 41.2 bits (95), Expect = 0.002
Identities = 21/55 (38%), Positives = 30/55 (54%)
Frame = +2
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCE 262
V WL +LGL +Y +F +VD + L LT EDL+ MG+ A + A+ E
Sbjct: 11 VRSWLYELGLSRYAPMFEIHDVDDELLTMLTSEDLIDMGINAADSSMNMYTAIHE 175
>TC221800 PRF|1503276A.0|226263|1503276A chlorophyll a/b binding protein.
{Glycine max;} , partial (20%)
Length = 864
Score = 38.1 bits (87), Expect = 0.013
Identities = 20/46 (43%), Positives = 29/46 (62%)
Frame = +1
Query: 223 VFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFRKGIA 268
VF EVD + L LT EDL MG++A+G R+K+ A+ + KG +
Sbjct: 22 VFEVHEVDDEVLPLLTLEDLKDMGISAVGSRRKMYTAIQKLGKGFS 159
>CK606020
Length = 340
Score = 32.0 bits (71), Expect = 0.96
Identities = 23/70 (32%), Positives = 39/70 (54%)
Frame = -3
Query: 83 PSTIDSVGTALTDSSSSSSSASVSAFVISSPVKVKMKGTAYFGNSIESKLVVSRANAINN 142
PST S ++ T SSSSSSS+S+S+ SS ++ +S S S ++++
Sbjct: 326 PSTSSSSTSSSTSSSSSSSSSSISSSSSSS-----SSSSSSSSSSSSSSSDGSISSSLAT 162
Query: 143 AEAGSELNLS 152
E+GS++ L+
Sbjct: 161 GESGSDIGLN 132
>TC227013 similar to UP|GAC1_YEAST (P28006) Protein phosphatase 1 regulatory
subunit GAC1, partial (3%)
Length = 1149
Score = 31.6 bits (70), Expect = 1.2
Identities = 31/99 (31%), Positives = 46/99 (46%), Gaps = 2/99 (2%)
Frame = -1
Query: 72 YEIENCSLDFIPSTIDSVGTALTDSSSSSSSASVSAFVISSPVKVKMKGTAYFGNSIESK 131
+ + N S I + V T L SSSSSSS+S S+ + SS + F S S
Sbjct: 348 WPLSNASSAEIGDPSEFVRTKLNSSSSSSSSSSSSSSISSSLC------FSSFSFSTFSS 187
Query: 132 LVVSRA--NAINNAEAGSELNLSDELDGSVRCPLCEVDI 168
L++S A + N SE +L+D+ + C L V +
Sbjct: 186 LILSLAMKSCSNFPANSSEESLNDKGESETACILVSVPL 70
>TC204038 similar to UP|Q8VZC0 (Q8VZC0) DTDP-glucose 4-6-dehydratase-like
protein, partial (37%)
Length = 1102
Score = 31.6 bits (70), Expect = 1.2
Identities = 25/72 (34%), Positives = 34/72 (46%)
Frame = +1
Query: 43 HKLETTAFGGSGKENVPPNASSSTIHDDGYEIENCSLDFIPSTIDSVGTALTDSSSSSSS 102
H+ E ++G + PPN SS+ L PST S +A SSS+SSS
Sbjct: 7 HRREEESWGAARAHPTPPNPSSTP-----------DLSPDPSTTSSASSASCSSSSASSS 153
Query: 103 ASVSAFVISSPV 114
A S+ SSP+
Sbjct: 154 APPSS--SSSPL 183
>TC204347 homologue to UP|Q9SP48 (Q9SP48) Homeodomain-leucine zipper protein
56, partial (71%)
Length = 2071
Score = 30.4 bits (67), Expect = 2.8
Identities = 39/160 (24%), Positives = 59/160 (36%), Gaps = 2/160 (1%)
Frame = +1
Query: 93 LTDSSSSSSSASVSAFVISSPVKVKMKGTAYFGNSIESKLVVSRANAINNAEAGSELNLS 152
LT SSSSSSS+S + + S P+ ++ NS +S + ++ E S N
Sbjct: 535 LTSSSSSSSSSSNNNKIPSFPLMKRL-------NSSDSSTALMTIFPSSSTEEHSPRNSH 693
Query: 153 DELDGSVRCPLCEVDISNLTEEQRHLHTNDCLDRGDDVAVRDDNVDAQFAPKVSRVVDWL 212
R L +D EE H D V + N + + + R V
Sbjct: 694 HMYGREFRSMLDGLDEEGCVEEPGHQSEKKRRLSVDQVKALEKNFEVENKLEPERKVKLA 873
Query: 213 RDLGLGKYEDV--FVREEVDWDTLQWLTEEDLLSMGVTAL 250
++LGL + F W T Q + +L AL
Sbjct: 874 QELGLQPRQVAVWFQNRRARWKTKQLERDYGVLKANYDAL 993
>TC223062 similar to UP|Q96280 (Q96280) Orf12 protein, partial (8%)
Length = 605
Score = 30.4 bits (67), Expect = 2.8
Identities = 13/22 (59%), Positives = 18/22 (81%)
Frame = +3
Query: 663 SFTELKDFVKHISPANIIPSVN 684
SF+EL+DFV+ + P IIP+VN
Sbjct: 6 SFSELQDFVQVLRPDKIIPTVN 71
>TC209366 homologue to UP|Q6T7B6 (Q6T7B6) C-type lectin-2, partial (8%)
Length = 839
Score = 30.4 bits (67), Expect = 2.8
Identities = 23/88 (26%), Positives = 37/88 (41%)
Frame = +2
Query: 96 SSSSSSSASVSAFVISSPVKVKMKGTAYFGNSIESKLVVSRANAINNAEAGSELNLSDEL 155
S+S S + FV +K+KMK ++ + S++V E L
Sbjct: 338 STSGSMITQIPCFVQLKSLKLKMKSSSSISDEGVSRIV--------------EYLLK--- 466
Query: 156 DGSVRCPLCEVDISNLTEEQRHLHTNDC 183
+CPL +VD+ N E HL+ +C
Sbjct: 467 ----KCPLAKVDVINCRE*YEHLNPQNC 538
>TC220745 similar to UP|Q9LHD1 (Q9LHD1) Multidrug resistance p-glycoprotein,
partial (18%)
Length = 1050
Score = 30.0 bits (66), Expect = 3.6
Identities = 25/86 (29%), Positives = 35/86 (40%), Gaps = 9/86 (10%)
Frame = +3
Query: 38 RPLKKH----KLETTAFGGSGKENVPPNASSSTIHDDGYEIENCSL-----DFIPSTIDS 88
R L+KH E T FGG+ +EN+ AS++ D EI + DFI S D
Sbjct: 174 RSLRKHIALVSQEPTLFGGTIRENIAYGASNNNNKVDETEIIEAARAANAHDFIASLKDG 353
Query: 89 VGTALTDSSSSSSSASVSAFVISSPV 114
T+ D S I+ +
Sbjct: 354 YDTSCRDRGVQLSGGQKQRIAIARAI 431
>BM887423 weakly similar to GP|14141201|gb| SRp46 splicing factor {Homo
sapiens}, partial (9%)
Length = 294
Score = 29.6 bits (65), Expect = 4.7
Identities = 20/69 (28%), Positives = 34/69 (48%)
Frame = -3
Query: 44 KLETTAFGGSGKENVPPNASSSTIHDDGYEIENCSLDFIPSTIDSVGTALTDSSSSSSSA 103
K+ F SG++ P ST H NCS D +PS+ + T T SSSS++
Sbjct: 229 KIPKRFFSSSGEKYFLP---FSTFHSS----HNCSTDLVPSSSRPMVT*ATASSSSAAEH 71
Query: 104 SVSAFVISS 112
+ + ++++
Sbjct: 70 ASNTLILAT 44
>TC207245 similar to UP|Q75LK9 (Q75LK9) Expressed protein, partial (59%)
Length = 860
Score = 29.3 bits (64), Expect = 6.2
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = -2
Query: 378 KYLRGDCSHWFLTHFHL 394
+Y +GDCSH THFH+
Sbjct: 205 EYPQGDCSHKMCTHFHI 155
>TC207243 similar to UP|Q93Z55 (Q93Z55) AT3g58470/F14P22_60, partial (60%)
Length = 883
Score = 29.3 bits (64), Expect = 6.2
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = -2
Query: 378 KYLRGDCSHWFLTHFHL 394
+Y +GDCSH THFH+
Sbjct: 447 EYPQGDCSHKMCTHFHI 397
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,838,327
Number of Sequences: 63676
Number of extensions: 452577
Number of successful extensions: 2795
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 2482
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2668
length of query: 700
length of database: 12,639,632
effective HSP length: 104
effective length of query: 596
effective length of database: 6,017,328
effective search space: 3586327488
effective search space used: 3586327488
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0133.3


