

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0132.6
(162 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
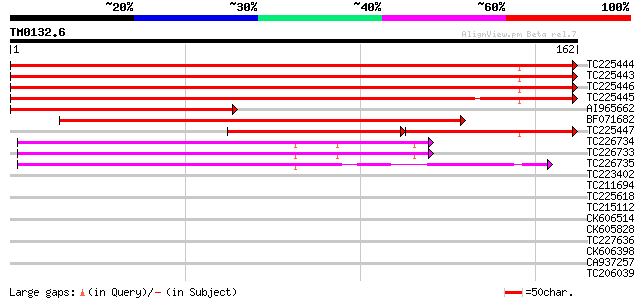
Sequences producing significant alignments: (bits) Value
TC225444 homologue to UP|RL24_CICAR (O65743) 60S ribosomal prote... 288 8e-79
TC225443 homologue to UP|RL24_CICAR (O65743) 60S ribosomal prote... 285 7e-78
TC225446 similar to UP|RL24_HORVU (P50888) 60S ribosomal protein... 284 1e-77
TC225445 homologue to UP|RL24_CICAR (O65743) 60S ribosomal prote... 276 2e-75
AI965662 homologue to GP|10180025|gb| 60S ribosomal protein L24 ... 129 8e-31
BF071682 weakly similar to SP|P38666|RL24 60S ribosomal protein ... 122 7e-29
TC225447 homologue to UP|RL24_CICAR (O65743) 60S ribosomal prote... 89 9e-19
TC226734 similar to UP|O22165 (O22165) 60S ribosomal protein L30... 77 4e-15
TC226733 similar to UP|O22165 (O22165) 60S ribosomal protein L30... 76 6e-15
TC226735 similar to UP|O22165 (O22165) 60S ribosomal protein L30... 75 2e-14
TC223402 weakly similar to UP|Q9SFV8 (Q9SFV8) T1B9.5 protein, pa... 33 0.059
TC211694 weakly similar to UP|WAIP_HUMAN (O43516) Wiskott-Aldric... 33 0.059
TC225618 similar to UP|Q9LSQ5 (Q9LSQ5) 1,4-benzoquinone reductas... 32 0.10
TC215112 32 0.13
CK606514 31 0.30
CK605828 30 0.39
TC227636 weakly similar to UP|Q9SYP0 (Q9SYP0) F9H16.4 protein, p... 30 0.39
CK606398 30 0.39
CA937257 29 0.86
TC206039 29 0.86
>TC225444 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 750
Score = 288 bits (737), Expect = 8e-79
Identities = 142/164 (86%), Positives = 153/164 (92%), Gaps = 2/164 (1%)
Frame = +2
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPG+GIRF+R DSQVFLF NSKCKRYFHNRLKPSKLTWTAM+RKQ
Sbjct: 44 MVLKTELCRFSGAKIYPGKGIRFVRGDSQVFLFANSKCKRYFHNRLKPSKLTWTAMYRKQ 223
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKDIAQEA +K+RR+TKKPYSRSIVGATLEVIQK+RTEKPEVRDAAREA LREIKER+K
Sbjct: 224 HKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDAAREAQLREIKERIK 403
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKG--GKVAAPKGPKLGGGGGKR 162
KTKD+K+AKKAEV K+QKSQGKG K A PKGPKLGGGGGKR
Sbjct: 404 KTKDDKKAKKAEVAAKSQKSQGKGSISKGAMPKGPKLGGGGGKR 535
>TC225443 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 767
Score = 285 bits (729), Expect = 7e-78
Identities = 141/164 (85%), Positives = 152/164 (91%), Gaps = 2/164 (1%)
Frame = +2
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPG+GIRF+R DSQVFLF NSKCKRYFHNRLKPSKLTWTAM+RKQ
Sbjct: 89 MVLKTELCRFSGAKIYPGKGIRFVRGDSQVFLFANSKCKRYFHNRLKPSKLTWTAMYRKQ 268
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKDIAQEA RK+RR+ KKPYSRSIVGATLEVIQK+R EKPEVRDAAREAALREIKER+K
Sbjct: 269 HKKDIAQEAVRKRRRAAKKPYSRSIVGATLEVIQKKRAEKPEVRDAAREAALREIKERIK 448
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKG--GKVAAPKGPKLGGGGGKR 162
KTKDEK+AKKAEV +K+QK+ GKG K A PKGPKLGGGGGKR
Sbjct: 449 KTKDEKKAKKAEVASKSQKAGGKGNVSKGAMPKGPKLGGGGGKR 580
>TC225446 similar to UP|RL24_HORVU (P50888) 60S ribosomal protein L24,
partial (96%)
Length = 865
Score = 284 bits (726), Expect = 1e-77
Identities = 140/163 (85%), Positives = 151/163 (91%), Gaps = 1/163 (0%)
Frame = +2
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPG+GIRF+R DSQVFLF NSKCKRYFHNRLKPSKLTWTAM+RKQ
Sbjct: 95 MVLKTELCRFSGAKIYPGKGIRFVRGDSQVFLFANSKCKRYFHNRLKPSKLTWTAMYRKQ 274
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKDIAQEA +K+RR+ KKPYSRSIVGATLEVIQK+R EKPEVRDAAREAALREIKER+K
Sbjct: 275 HKKDIAQEAVKKRRRAAKKPYSRSIVGATLEVIQKKRAEKPEVRDAAREAALREIKERIK 454
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKG-GKVAAPKGPKLGGGGGKR 162
KTKDEK+AKKAEV K+QK+ GKG K A PKGPKLGGGGGKR
Sbjct: 455 KTKDEKKAKKAEVTAKSQKAGGKGISKGAMPKGPKLGGGGGKR 583
>TC225445 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 723
Score = 276 bits (707), Expect = 2e-75
Identities = 140/164 (85%), Positives = 149/164 (90%), Gaps = 2/164 (1%)
Frame = +2
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPG+GIRF+R DSQVFLF NSKCKRYFHNRLKPSKLTWTAM+RKQ
Sbjct: 56 MVLKTELCRFSGAKIYPGKGIRFVRGDSQVFLFANSKCKRYFHNRLKPSKLTWTAMYRKQ 235
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKDIAQEA +K+RR+ KKPYSRSIVGATLEVIQK+R EKPEVRDA REAALREIKER K
Sbjct: 236 HKKDIAQEAVKKRRRAAKKPYSRSIVGATLEVIQKKRAEKPEVRDANREAALREIKERNK 415
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKG--GKVAAPKGPKLGGGGGKR 162
KTKDEK+AKKAE K+QKSQGKG K A PKGPKLGGGGGKR
Sbjct: 416 KTKDEKKAKKAEF-AKSQKSQGKGNVSKGAMPKGPKLGGGGGKR 544
>AI965662 homologue to GP|10180025|gb| 60S ribosomal protein L24 {Prunus
avium}, partial (34%)
Length = 402
Score = 129 bits (323), Expect = 8e-31
Identities = 59/65 (90%), Positives = 62/65 (94%)
Frame = +3
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPG+GIRF+R DSQVFLF NSKCKRYFHNRLKPSKLTWTAM+RKQ
Sbjct: 63 MVLKTELCRFSGAKIYPGKGIRFVRGDSQVFLFANSKCKRYFHNRLKPSKLTWTAMYRKQ 242
Query: 61 HKKDI 65
HKK I
Sbjct: 243 HKKVI 257
>BF071682 weakly similar to SP|P38666|RL24 60S ribosomal protein L24.
[Mouse-ear cress] {Arabidopsis thaliana}, partial (63%)
Length = 352
Score = 122 bits (306), Expect = 7e-29
Identities = 65/116 (56%), Positives = 77/116 (66%)
Frame = +2
Query: 15 IYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHKKDIAQEAARKKR 74
IYPG I + SQV L NS R HNRLK +KLT TAM++KQHKKDIAQ RK R
Sbjct: 2 IYPGNAITYFHRHSQVILSANS*FNRSSHNRLKHTKLTCTAMYKKQHKKDIAQ*DMRKMR 181
Query: 75 RSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKTKDEKRAKK 130
+ KKPYS SIVGATL++I + + P++ DA EAAL EIKE +K T DE A K
Sbjct: 182 LAAKKPYSMSIVGATLQLIHIKTADNPKLLDATNEAALFEIKETIKNTNDENNANK 349
>TC225447 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
partial (61%)
Length = 687
Score = 89.0 bits (219), Expect = 9e-19
Identities = 44/51 (86%), Positives = 48/51 (93%)
Frame = +1
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALR 113
+DIAQEA RK+RR+ KKPYSRSIVGATLEVIQK+R EKPEVRDAAREAALR
Sbjct: 4 RDIAQEAVRKRRRAAKKPYSRSIVGATLEVIQKKRAEKPEVRDAAREAALR 156
Score = 77.8 bits (190), Expect = 2e-15
Identities = 39/51 (76%), Positives = 44/51 (85%), Gaps = 2/51 (3%)
Frame = +3
Query: 114 EIKERVKKTKDEKRAKKAEVQTKAQKSQGKG--GKVAAPKGPKLGGGGGKR 162
EIKER+KKTKDEK+AKKAEV +K+QK+ GKG K A PKGPKLGGGGGKR
Sbjct: 459 EIKERIKKTKDEKKAKKAEVASKSQKAGGKGNVSKGAMPKGPKLGGGGGKR 611
>TC226734 similar to UP|O22165 (O22165) 60S ribosomal protein L30
(At2g44860/T13E15.13), partial (86%)
Length = 811
Score = 77.0 bits (188), Expect = 4e-15
Identities = 42/126 (33%), Positives = 68/126 (53%), Gaps = 7/126 (5%)
Frame = +3
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
++ E C F + IYPG GI+F+R+D+++F F SKC + F + P K+ WT +R+ H
Sbjct: 87 MRLEKCWFCSSTIYPGHGIQFVRNDAKIFRFCRSKCHKNFKMKRNPRKVKWTKAYRRVHG 266
Query: 63 KDIAQEAARKKRRSTKKP--YSRSIVGATLEV---IQKRRTEKPEVRDAAREAALRE--I 115
KD+ Q++ + R +P Y R++ L+ I K R + E R +E +
Sbjct: 267 KDMTQDSTFEFERKRNRPERYDRNLAENVLKAIPKIDKIRVSREERHHKNRMKGKKEKLL 446
Query: 116 KERVKK 121
KE VK+
Sbjct: 447 KEAVKE 464
>TC226733 similar to UP|O22165 (O22165) 60S ribosomal protein L30
(At2g44860/T13E15.13), partial (86%)
Length = 822
Score = 76.3 bits (186), Expect = 6e-15
Identities = 41/126 (32%), Positives = 68/126 (53%), Gaps = 7/126 (5%)
Frame = +3
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
++ E C F + +YPG GI+F+R+D+++F F SKC + F + P K+ WT +R+ H
Sbjct: 108 MRLEKCWFCSSTVYPGHGIQFVRNDAKIFRFCRSKCHKNFKMKRNPRKVKWTKAYRRVHG 287
Query: 63 KDIAQEAARKKRRSTKKP--YSRSIVGATLEV---IQKRRTEKPEVRDAAREAALRE--I 115
KD+ Q++ + R +P Y R++ L+ I K R + E R +E +
Sbjct: 288 KDMTQDSTFEFERKRNRPERYDRNLAENVLKAIPKIDKIRVTREERHHKNRMKGKKEKLL 467
Query: 116 KERVKK 121
KE VK+
Sbjct: 468 KEAVKE 485
>TC226735 similar to UP|O22165 (O22165) 60S ribosomal protein L30
(At2g44860/T13E15.13), partial (74%)
Length = 936
Score = 74.7 bits (182), Expect = 2e-14
Identities = 44/155 (28%), Positives = 78/155 (49%), Gaps = 2/155 (1%)
Frame = +3
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
++ E C F + YPG GI+F+R+D+++F F SKC + F + P K+ WT +R+ H
Sbjct: 105 MRLEKCWFCSSTTYPGHGIQFVRNDAKIFRFCRSKCHKNFKMKRNPRKVKWTKAYRRVHG 284
Query: 63 KDIAQEAARKKRRSTKKP--YSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
KD+ Q++ + R +P Y R++ L+ I K ++R A E K
Sbjct: 285 KDMTQDSTFEFERKRNRPERYDRNLAENVLKAIPK----IDKIRVAREE----------K 422
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKL 155
K+ + KK ++ +A + +G ++ K P +
Sbjct: 423 HHKNRMKGKKQKLHMEAVRELEQG--ISLVKAPSV 521
>TC223402 weakly similar to UP|Q9SFV8 (Q9SFV8) T1B9.5 protein, partial (11%)
Length = 699
Score = 33.1 bits (74), Expect = 0.059
Identities = 17/49 (34%), Positives = 25/49 (50%)
Frame = +3
Query: 72 KKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
KK R TK+ R +E ++KRRT+K + + EIK+R K
Sbjct: 27 KKGRGTKRRRKRETKKEGMESVRKRRTKKKNIEKKRKTRTRTEIKKRAK 173
>TC211694 weakly similar to UP|WAIP_HUMAN (O43516) Wiskott-Aldrich syndrome
protein interacting protein (WASP interacting protein)
(PRPL-2 protein), partial (5%)
Length = 782
Score = 33.1 bits (74), Expect = 0.059
Identities = 24/67 (35%), Positives = 30/67 (43%), Gaps = 2/67 (2%)
Frame = -1
Query: 96 RRTEKPEVRDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGK--GGKVAAPKGP 153
RR E P+V A LR+ E V A+ E +K G GG V P+G
Sbjct: 644 RRAECPQVHGEA----LRDGVEGVVTVGPAPDAEDGEAAAGEEKGAGAEVGGGVGEPRGE 477
Query: 154 KLGGGGG 160
+ GGGGG
Sbjct: 476 EAGGGGG 456
>TC225618 similar to UP|Q9LSQ5 (Q9LSQ5) 1,4-benzoquinone reductase-like; Trp
repressor binding protein-like (1,4-benzoquinone
reductase-like protein), partial (75%)
Length = 1864
Score = 32.3 bits (72), Expect = 0.10
Identities = 19/49 (38%), Positives = 27/49 (54%)
Frame = -2
Query: 114 EIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGGKR 162
E ERVK+ K A+ V A + +G+GG +A +G K G GG +R
Sbjct: 426 EESERVKRKKSRGMARAVAVAVAACEREGEGGGGSA-EGGKSGAGGSQR 283
>TC215112
Length = 1736
Score = 32.0 bits (71), Expect = 0.13
Identities = 16/42 (38%), Positives = 25/42 (59%)
Frame = -2
Query: 116 KERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGG 157
K++ KK K +K+ KK + + K +K + G AAP P +GG
Sbjct: 235 KKKKKKKKKKKKKKKKKKKKKKKKKKNFLGGXAAPNFPFMGG 110
>CK606514
Length = 280
Score = 30.8 bits (68), Expect = 0.30
Identities = 18/47 (38%), Positives = 26/47 (55%)
Frame = +1
Query: 116 KERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGGKR 162
K+ + K++K +K K K + KGGK GPK GGGGG++
Sbjct: 1 KKPPPQKKNKKFPQKKTPNFKI*KKEKKGGKFF---GPKKGGGGGEK 132
>CK605828
Length = 553
Score = 30.4 bits (67), Expect = 0.39
Identities = 29/103 (28%), Positives = 41/103 (39%), Gaps = 13/103 (12%)
Frame = +3
Query: 71 RKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKTKDEKRAKK 130
+KK R KK G + QK +K + ++ A + K D + AKK
Sbjct: 39 KKKNRGGKKKKGGGKKGKS----QKNPPKKKAPQTPPKKGAPPKNPFNTKTPPDPRGAKK 206
Query: 131 AEVQTKA-------------QKSQGKGGKVAAPKGPKLGGGGG 160
K +K + +GGK P+GP LGGGGG
Sbjct: 207 TRGVHKTPPKGARRKKGGPPKKPKKRGGK--EPQGPPLGGGGG 329
>TC227636 weakly similar to UP|Q9SYP0 (Q9SYP0) F9H16.4 protein, partial (8%)
Length = 1200
Score = 30.4 bits (67), Expect = 0.39
Identities = 24/79 (30%), Positives = 38/79 (47%), Gaps = 4/79 (5%)
Frame = +3
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSI--VGATLEVIQ--KRRTEKPEVRDAAREAALR 113
+++ + + E ARK+ K R + + E +Q KR EK + R A +
Sbjct: 420 KEEEELILKAEKARKEEEEAKLKEKRRLEEIEKAKEALQRKKRNAEKAQQRAALKAQKEA 599
Query: 114 EIKERVKKTKDEKRAKKAE 132
E+KE+ + EKRAKK E
Sbjct: 600 ELKEKER----EKRAKKKE 644
>CK606398
Length = 505
Score = 30.4 bits (67), Expect = 0.39
Identities = 26/94 (27%), Positives = 40/94 (41%), Gaps = 2/94 (2%)
Frame = +2
Query: 71 RKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKTKDEKRAKK 130
RKK+ TKK ++ + QK+ +P + V+K K+ K +
Sbjct: 80 RKKKPKTKKGWAEKNKKMSPPNPQKKENPEP-------------LNSGVEKGKN*KPPQT 220
Query: 131 AEVQTKAQKSQG--KGGKVAAPKGPKLGGGGGKR 162
+ + QK G K G A + LGGGGGK+
Sbjct: 221 PKPGPQTQKGSGNPKIGPPAGARKKNLGGGGGKK 322
>CA937257
Length = 427
Score = 29.3 bits (64), Expect = 0.86
Identities = 14/39 (35%), Positives = 21/39 (52%)
Frame = +1
Query: 123 KDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGGK 161
K +K+ KK + + K +K + G K K GGGGG+
Sbjct: 169 KKKKKKKKKKKKKKKKKKRXXGKKXXEEGEKKKGGGGGR 285
>TC206039
Length = 834
Score = 29.3 bits (64), Expect = 0.86
Identities = 16/46 (34%), Positives = 24/46 (51%)
Frame = -2
Query: 116 KERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGGK 161
K++ KK K +K+ KK + + K +K K K K GGGG+
Sbjct: 257 KKKKKKKKKKKKKKKKKKKKKKKKKIKKKKKKKKKKKKXXXGGGGE 120
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,590,020
Number of Sequences: 63676
Number of extensions: 66355
Number of successful extensions: 694
Number of sequences better than 10.0: 86
Number of HSP's better than 10.0 without gapping: 661
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 680
length of query: 162
length of database: 12,639,632
effective HSP length: 90
effective length of query: 72
effective length of database: 6,908,792
effective search space: 497433024
effective search space used: 497433024
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0132.6


