

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0131.12
(78 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
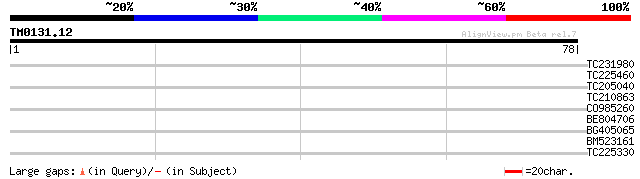
Score E
Sequences producing significant alignments: (bits) Value
TC231980 similar to UP|Q8S562 (Q8S562) KAP-2, partial (19%) 36 0.003
TC225460 homologue to UP|ATP2_MAIZE (P19023) ATP synthase beta c... 27 2.5
TC205040 similar to UP|Q9FFF5 (Q9FFF5) Nucleic acid binding prot... 27 2.5
TC210863 similar to UP|O49654 (O49654) Leucine rich repeat recep... 26 3.2
CO985260 25 5.5
BE804706 25 7.2
BG405065 25 7.2
BM523161 25 7.2
TC225330 similar to UP|ATPX_SPIOL (P31853) ATP synthase B' chain... 25 9.4
>TC231980 similar to UP|Q8S562 (Q8S562) KAP-2, partial (19%)
Length = 590
Score = 36.2 bits (82), Expect = 0.003
Identities = 16/43 (37%), Positives = 25/43 (57%)
Frame = -3
Query: 14 LLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFS 56
LL+T + + VW+ TW +A+DI + Y+ + T P I FS
Sbjct: 582 LLLTATXSKSEQVWDQTWHWMADDIVYNYKKSSTSPSQIFYFS 454
>TC225460 homologue to UP|ATP2_MAIZE (P19023) ATP synthase beta chain,
mitochondrial precursor , partial (26%)
Length = 859
Score = 26.6 bits (57), Expect = 2.5
Identities = 9/26 (34%), Positives = 16/26 (60%)
Frame = -3
Query: 15 LITNIMNNPDHVWNITWKLLANDIQH 40
L +++N +HV + WK++ IQH
Sbjct: 407 LCNDLLNTANHVKRLLWKVIVFSIQH 330
>TC205040 similar to UP|Q9FFF5 (Q9FFF5) Nucleic acid binding protein-like,
partial (84%)
Length = 1092
Score = 26.6 bits (57), Expect = 2.5
Identities = 13/45 (28%), Positives = 24/45 (52%)
Frame = +2
Query: 27 WNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILLKIDNHLQ 71
++++WK L Q + T P +P+F+LS K + D+ +Q
Sbjct: 173 FSLSWK*LRLPAQSKRSSRTTPPAELPSFALSLKMWMNSTDSAIQ 307
>TC210863 similar to UP|O49654 (O49654) Leucine rich repeat receptor
kinase-like protein (At4g22730), partial (23%)
Length = 843
Score = 26.2 bits (56), Expect = 3.2
Identities = 10/28 (35%), Positives = 18/28 (63%)
Frame = -1
Query: 8 RRLFASLLITNIMNNPDHVWNITWKLLA 35
R L+ S T ++N+ H W++ WKL++
Sbjct: 699 R*LYMSR*HTQLLNHIHHGWSLVWKLMS 616
>CO985260
Length = 771
Score = 25.4 bits (54), Expect = 5.5
Identities = 14/53 (26%), Positives = 26/53 (48%)
Frame = -1
Query: 7 LRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSS 59
LRR A + T++ + +HV + +ND+ H T ++ G++ F S
Sbjct: 579 LRRWSAGMGETSVQDQAEHVPEDPVPVTSNDVVHAEAPTNSEVGVVSDFITES 421
>BE804706
Length = 190
Score = 25.0 bits (53), Expect = 7.2
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = -3
Query: 38 IQHEYRGTLTQPGMIPTFSLSSKPILLK 65
++H++R + +P IP SLS LLK
Sbjct: 86 LRHDFRSSARRPRAIPDISLSLDSPLLK 3
>BG405065
Length = 394
Score = 25.0 bits (53), Expect = 7.2
Identities = 11/25 (44%), Positives = 16/25 (64%), Gaps = 1/25 (4%)
Frame = +2
Query: 2 HSGNELRRLFASLLITNIMN-NPDH 25
H GN++ R++ SL +MN NP H
Sbjct: 272 HPGNKIERVYWSLSAHEVMNSNPGH 346
>BM523161
Length = 360
Score = 25.0 bits (53), Expect = 7.2
Identities = 13/49 (26%), Positives = 24/49 (48%)
Frame = -1
Query: 25 HVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILLKIDNHLQSN 73
H T K+ + + RG T+ +P LSS + +I+ H+Q++
Sbjct: 174 HQIKFTLKIAVHSLS--IRGNFTESEPLPCVKLSSVAVAFEIEVHMQNS 34
>TC225330 similar to UP|ATPX_SPIOL (P31853) ATP synthase B' chain,
chloroplast precursor (Subunit II) , partial (67%)
Length = 982
Score = 24.6 bits (52), Expect = 9.4
Identities = 12/43 (27%), Positives = 19/43 (43%)
Frame = +1
Query: 35 ANDIQHEYRGTLTQPGMIPTFSLSSKPILLKIDNHLQSNGKSL 77
A QH ++ + + +L S P+ L D H N K+L
Sbjct: 1 ATHTQHNTTSSICVAKNLDSLALKSNPLRLSRDTHTHPNHKAL 129
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,181,085
Number of Sequences: 63676
Number of extensions: 56602
Number of successful extensions: 275
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 275
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 275
length of query: 78
length of database: 12,639,632
effective HSP length: 54
effective length of query: 24
effective length of database: 9,201,128
effective search space: 220827072
effective search space used: 220827072
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0131.12


