

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0126.7
(542 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
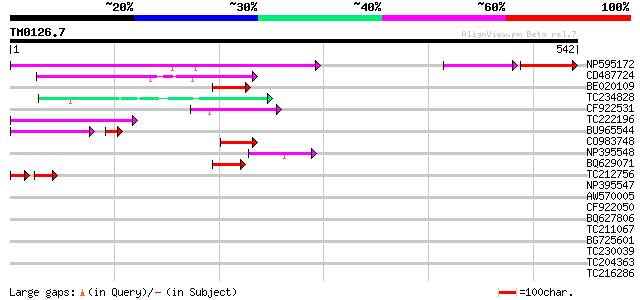
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 177 1e-44
CD487724 103 1e-22
BE020109 60 3e-09
TC234828 59 7e-09
CF922531 54 2e-07
TC222196 weakly similar to UP|Q77MR7 (Q77MR7) UL45 (Envelope tra... 52 5e-07
BU965544 44 6e-06
CO983748 48 1e-05
NP395548 reverse transcriptase [Glycine max] 45 8e-05
BQ629071 44 2e-04
TC212756 31 8e-04
NP395547 reverse transcriptase [Glycine max] 38 0.010
AW570005 38 0.013
CF922050 37 0.029
BQ627806 34 0.19
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 33 0.25
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 32 0.94
TC230039 homologue to UP|Q9M5J0 (Q9M5J0) Pectin methylesterase i... 31 1.6
TC204363 similar to UP|O22403 (O22403) ADL2 protein, partial (15%) 30 2.7
TC216286 similar to UP|Q8GTR0 (Q8GTR0) Sugar transporter, partia... 29 6.1
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 177 bits (448), Expect = 1e-44
Identities = 109/305 (35%), Positives = 171/305 (55%), Gaps = 8/305 (2%)
Frame = +1
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VE 59
LSL ++ +++ G++G + V +LVD G S NFI + PV P + V
Sbjct: 1129 LSLNAMRGSNGVGTIRFTGQVGGIAVKILVDGGSSDNFIQPRVAQVLKLPVEPAPNLRVL 1308
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ +G + +G + + L IQ ++ +L + G DV+LG W A+LG A++ L+
Sbjct: 1309 VGNGQILSAEGIVQQLPLHIQGQEVKVPVYLLQISGADVILGSTWLATLGPHVADYAALT 1488
Query: 120 LRFHVGEK-VLLRGELGLIRKNASLSSLVKALQQQA--EGFWVEFQHLAEVQEELVAVPE 176
L+F +K + L+GE A L + ++ E F ++ ++ L +P
Sbjct: 1489 LKFFQNDKFITLQGEGNSEATQAQLHHFRRLQNTKSIEECFAIQLIQKEVPEDTLKDLPT 1668
Query: 177 ----QLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKL 232
+L L+ Y+++F+ LPP R QDHAI L++G+ +RPYR PH QK +IEK+
Sbjct: 1669 NIDPELAILLHTYAQVFAVPASLPPQREQDHAIPLKQGSGPVKVRPYRYPHTQKDQIEKM 1848
Query: 233 VKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYK 292
++EML GI +PS S FS ++LVKKKDGSWRFC DYRA + IT+ + FP+ +DELL +
Sbjct: 1849 IQEMLVQGIIQPSNSPFSLPILLVKKKDGSWRFCTDYRALNAITVKDSFPMPTVDELLDE 2028
Query: 293 LGEAK 297
L A+
Sbjct: 2029 LHGAQ 2043
Score = 42.4 bits (98), Expect(2) = 2e-09
Identities = 21/54 (38%), Positives = 35/54 (63%)
Frame = +1
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFIIR 542
G VL Q P+A++S ++ + Q++S Y REL+A+ A+ ++ L G+ FIIR
Sbjct: 2710 GAVLGQNGHPIAYFSKKLAPRMQKQSAYTRELLAITEALSKFRHYLLGNKFIIR 2871
Score = 37.7 bits (86), Expect(2) = 2e-09
Identities = 29/71 (40%), Positives = 36/71 (49%)
Frame = +2
Query: 415 MSRG*GDFWGSQATIEGL*KDMGRLLDP*LCY*KRIVLCGVRRHD*LLTSSSMP*LTCQL 474
MS *GDF QA IE L M LLD *L + K+ V G+ + L+SS P* Q
Sbjct: 2468 MSSS*GDF*A*QAIIEDLLNLMPILLDH*LIFYKKTVSYGIMKQKQHLSSSKKP*RRLQY 2647
Query: 475 WQSLIFNNFCY 485
+ IF N +
Sbjct: 2648 YHYQIFLNHLF 2680
>CD487724
Length = 676
Score = 103 bits (258), Expect = 1e-22
Identities = 73/223 (32%), Positives = 117/223 (51%), Gaps = 11/223 (4%)
Frame = +3
Query: 26 VIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VELEDGHKVECQGKCRDVQLSIQHIPI 84
++ LVD G +HNF+ Q L+S P TP + V + +GH ++C C + +SIQ+I
Sbjct: 24 LVYLVDGGSTHNFVQQPLVSQLGLPCRSTPPLRVMVGNGHHLKCTTICEAIPISIQNIEF 203
Query: 85 EQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRFHVGEKVL-LRGE----LGLIRK 139
++ + G ++VLG+ W +LG I ++ LS++F +++ L+GE LGL+
Sbjct: 204 LVHLYVLPIVGANIVLGVQWLKTLGPILVDYNSLSMQFFYQHRLVQLKGESEAQLGLLNH 383
Query: 140 NASLSSLVKALQQQAEGFWVEFQHLAEVQEELVA-----VPEQLQGLMGDYSELFSTIWD 194
+ ++ L Q E V + H+A + E +P+ +Q L+ +S LF
Sbjct: 384 HQ-----LRRLHQTHEP--VTYFHIAILTENTSPTSSPPLPQPIQHLLDQFSALFQ*PQG 542
Query: 195 LPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEML 237
LPP+R DH IHL + N+R Y PHY +EIE V ML
Sbjct: 543 LPPARETDHHIHLLP*SEPVNMRLY*YPHY*NNEIEHQVNLML 671
>BE020109
Length = 422
Score = 59.7 bits (143), Expect = 3e-09
Identities = 27/36 (75%), Positives = 30/36 (83%)
Frame = -1
Query: 195 LPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIE 230
LPPSR QDH+I L+ GA IPNIRPYR P YQK+EIE
Sbjct: 113 LPPSRRQDHSIELKNGAEIPNIRPYRYPDYQKNEIE 6
>TC234828
Length = 857
Score = 58.5 bits (140), Expect = 7e-09
Identities = 56/227 (24%), Positives = 92/227 (39%), Gaps = 3/227 (1%)
Frame = +3
Query: 28 VLVDCGVSHNFISQELMSSAHFPVVETPY--MVELEDGHKVECQGKCRDVQLSIQHIPIE 85
VL D G +H+FIS + E PY +V V C + ++
Sbjct: 219 VLYDSGATHSFISHACVERLGLCATELPYDMVVSTPTSEPVTTSRVCLKCPIIVEGRSFM 398
Query: 86 QEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRFHVGEKVLLRGELGLIRKNASLSS 145
+ L LDV+LG+ W S HI + +E L F G V+ L N
Sbjct: 399 ADLICLPLAHLDVILGMDWL-STNHIFLDCKEKMLVF--GGDVVPSEPLKEDAANEET-- 563
Query: 146 LVKALQQQAEGFWVEFQHLAEVQEELVAVPEQLQGLMGDYSELF-STIWDLPPSRVQDHA 204
+ + V F E E+ +P ++ ++ E+F + +LPP R +
Sbjct: 564 ------EDVRTYMVLFSMYVEEDAEVSCIP-----VVSEFPEVFPDDVCELPPEREVEFI 710
Query: 205 IHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSS 251
I + GA +I PYR + +E++ V+++L RPS S + +
Sbjct: 711 IDVVPGANPVSIAPYRMSPVELAEVKAQVQDLLSKQFVRPSASPWGA 851
>CF922531
Length = 602
Score = 53.5 bits (127), Expect = 2e-07
Identities = 33/89 (37%), Positives = 50/89 (56%), Gaps = 2/89 (2%)
Frame = -2
Query: 174 VPEQLQGLMGDYSELF--STIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEK 231
+P ++Q L+ ++ ++F LPP R +H I L A++PN YR + EIE
Sbjct: 268 LPPKVQELLHEFGDIFPKEIPLGLPPLRGIEHQIDLVPRASLPNRPTYRTNPQETKEIES 89
Query: 232 LVKEMLDVGIFRPSISQFSSLVILVKKKD 260
VKE+L+ G + S+S LV+LV KKD
Sbjct: 88 QVKELLEKGWVQESLSLCVVLVLLVPKKD 2
>TC222196 weakly similar to UP|Q77MR7 (Q77MR7) UL45 (Envelope transmembrane
protein), partial (11%)
Length = 880
Score = 52.4 bits (124), Expect = 5e-07
Identities = 33/123 (26%), Positives = 63/123 (50%), Gaps = 1/123 (0%)
Frame = +3
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VE 59
+SL +++ + ++L+V G + +V+ L+D G HNFI + L ++ P TP + V
Sbjct: 165 ISLHALSSHMAPKTLRVLGRVANQQVLTLIDGGSIHNFIQELLATTLGLPTQPTPPLWVL 344
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ + H++ C V + +Q + + L D VL +*W SLG ++ L+
Sbjct: 345 VGNEHEI*CHLLYMKVLVCVQGQEFMVDLHVLRLCDEDHVL*V*WLKSLGPFVIDYNNLT 524
Query: 120 LRF 122
++F
Sbjct: 525 MKF 533
>BU965544
Length = 434
Score = 43.9 bits (102), Expect(2) = 6e-06
Identities = 21/82 (25%), Positives = 45/82 (54%), Gaps = 1/82 (1%)
Frame = -2
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVET-PYMVE 59
+SL +++ + + +V+G + R+ +L+D G ++NFI + + P +T P
Sbjct: 328 ISLNALSGLLALETFRVYGFINHARLTILIDIGSTNNFIQPCVAKFLNLPTKDTSPLREM 149
Query: 60 LEDGHKVECQGKCRDVQLSIQH 81
+ +G+ ++C+ C + L IQH
Sbjct: 148 VGNGYFLDCRQLCHVIALQIQH 83
Score = 24.3 bits (51), Expect(2) = 6e-06
Identities = 8/17 (47%), Positives = 13/17 (76%)
Frame = -3
Query: 92 NLGGLDVVLGL*WFASL 108
++ G+DV+LG+ WF L
Sbjct: 54 SISGIDVILGIEWFKHL 4
>CO983748
Length = 802
Score = 47.8 bits (112), Expect = 1e-05
Identities = 18/36 (50%), Positives = 26/36 (72%)
Frame = +1
Query: 202 DHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEML 237
D H +GA IPN+RPY+ PHYQ +++EK + +ML
Sbjct: 694 DFIYHSPQGATIPNLRPYKLPHYQNNKMEKTINDML 801
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 45.1 bits (105), Expect = 8e-05
Identities = 27/84 (32%), Positives = 45/84 (53%), Gaps = 19/84 (22%)
Frame = +1
Query: 229 IEKLVKEMLDVGIFRP-SISQFSSLVILVKKKDG------------------SWRFCIDY 269
+ K V ++L+VG+ P S S + S V++V KK+G SW+ CIDY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 270 RAFHKITIPNKFPILIIDELLYKL 293
R ++ T + FP+ +D++L +L
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERL 252
>BQ629071
Length = 395
Score = 43.5 bits (101), Expect = 2e-04
Identities = 19/31 (61%), Positives = 22/31 (70%)
Frame = -1
Query: 195 LPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQ 225
LPP R D AI L+EG I N+RPYR PHY+
Sbjct: 239 LPPKRRHDRAIILKEGV*ITNVRPYRYPHYE 147
Score = 28.9 bits (63), Expect = 6.1
Identities = 18/47 (38%), Positives = 26/47 (55%), Gaps = 9/47 (19%)
Frame = -2
Query: 222 PHYQKSEIEKLVKE---------MLDVGIFRPSISQFSSLVILVKKK 259
P Y K E E L+ + L+ I +PSI+ +SS VIL++KK
Sbjct: 211 PLY*KRESELLMSDPTGTHIMNVTLNAKIIKPSINHYSSPVILIRKK 71
>TC212756
Length = 493
Score = 30.8 bits (68), Expect(2) = 8e-04
Identities = 14/19 (73%), Positives = 16/19 (83%)
Frame = +3
Query: 1 LSLKSIAEFTSRRSLKVWG 19
LSL S+A FTSRRSL +WG
Sbjct: 51 LSLHSVAGFTSRRSLILWG 107
Score = 30.0 bits (66), Expect(2) = 8e-04
Identities = 12/22 (54%), Positives = 17/22 (76%)
Frame = +2
Query: 24 LRVIVLVDCGVSHNFISQELMS 45
L IVL+DCG +HN I+ +L+S
Sbjct: 113 LEEIVLIDCGATHNCIAYQLVS 178
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 38.1 bits (87), Expect = 0.010
Identities = 25/84 (29%), Positives = 41/84 (48%), Gaps = 19/84 (22%)
Frame = +1
Query: 229 IEKLVKEMLDVGIFRP-SISQFSSLVILVKKKDGS------------------WRFCIDY 269
+ K V ++L+ G+ P S S + S V +V KK G WR CIDY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 270 RAFHKITIPNKFPILIIDELLYKL 293
R ++ T + +P+ +D++L +L
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRL 252
>AW570005
Length = 413
Score = 37.7 bits (86), Expect = 0.013
Identities = 18/54 (33%), Positives = 28/54 (51%)
Frame = -2
Query: 487 ETGVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFI 540
+ G VL Q P+AF+S + R S Y EL A+ +++W + L G H +
Sbjct: 163 DMGAVLSQEGHPIAFFSKEFCPKLVRSSTYVHELAAITNVVKKWRQYLLGHHLV 2
>CF922050
Length = 303
Score = 36.6 bits (83), Expect = 0.029
Identities = 19/43 (44%), Positives = 26/43 (60%)
Frame = -2
Query: 195 LPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEML 237
LPP R H+I LQ A ++ PY+ PH+ K +IE+ V ML
Sbjct: 287 LPPHRNLVHSITLQPKAGSVSVLPYQYPHHHKIDIEQ*V*SML 159
>BQ627806
Length = 435
Score = 33.9 bits (76), Expect = 0.19
Identities = 17/44 (38%), Positives = 24/44 (53%)
Frame = +1
Query: 498 PLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFII 541
PL F + R S Y REL A+ A+++W + L G HF+I
Sbjct: 181 PLPFSFMPFCSKLLRASTYVRELAAITVAVKKWRQYLLGHHFVI 312
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 33.5 bits (75), Expect = 0.25
Identities = 18/51 (35%), Positives = 27/51 (52%)
Frame = +1
Query: 491 VLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFII 541
VLLQ P+A++S + Y++EL A++RA Q W L F+I
Sbjct: 361 VLLQGGHPIAYFSEKLHSATLNYPTYDKELYALIRAPQTWEHFLVCKEFVI 513
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 31.6 bits (70), Expect = 0.94
Identities = 13/37 (35%), Positives = 21/37 (56%)
Frame = -3
Query: 253 VILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDEL 289
V++VKK +G WR C DY + + +P+ ID +
Sbjct: 124 VVMVKKPNGKWRICTDYIDLN*ACPKDAYPLPNIDHM 14
>TC230039 homologue to UP|Q9M5J0 (Q9M5J0) Pectin methylesterase isoform alpha
(Fragment) , partial (70%)
Length = 913
Score = 30.8 bits (68), Expect = 1.6
Identities = 31/125 (24%), Positives = 54/125 (42%), Gaps = 11/125 (8%)
Frame = +1
Query: 107 SLGHIRANFEE--LSLRFHVGEKVLLRGELGL-IRKNASLSSLVKALQQQAEGF--WVEF 161
S GH+R FEE L + HV + + +L ++K AS + + + +GF W+
Sbjct: 1 SKGHVRDRFEEGLLEISHHVSNSLAMLKKLPAGVKKLASKNEVFPGYGKIKDGFPTWLST 180
Query: 162 QHLAEVQEELVAVPEQLQGLM------GDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPN 215
+ +Q AV E L+ G+++ + + P S IH++ GA N
Sbjct: 181 KDRKLLQ---AAVNETNFNLLVAKDGTGNFTTIAEAVAVAPNSSATRFVIHIKAGAYFEN 351
Query: 216 IRPYR 220
+ R
Sbjct: 352 VEVIR 366
>TC204363 similar to UP|O22403 (O22403) ADL2 protein, partial (15%)
Length = 1368
Score = 30.0 bits (66), Expect = 2.7
Identities = 22/105 (20%), Positives = 48/105 (44%), Gaps = 15/105 (14%)
Frame = +3
Query: 127 KVLLRGELGLIRKN-------ASLSSLVKALQQQAEGFWVE-------FQHLAEVQEELV 172
K+LLR G++RKN A + LV +++ +++ F+ + + +E+
Sbjct: 372 KLLLRSYYGIVRKNVEDLIPKAIMHFLVNNTKRELHNVFIKKLYRDNLFEEMLQEPDEIA 551
Query: 173 AVPEQLQGLMGDYSELFSTIWDLP-PSRVQDHAIHLQEGAAIPNI 216
++ + L+ Y + F + +LP + + L E + +P I
Sbjct: 552 VKRKRCRELLRAYQQAFKDLEELPLEAETVERGYSLPETSGLPKI 686
>TC216286 similar to UP|Q8GTR0 (Q8GTR0) Sugar transporter, partial (45%)
Length = 1342
Score = 28.9 bits (63), Expect = 6.1
Identities = 21/64 (32%), Positives = 29/64 (44%)
Frame = -3
Query: 20 ELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYMVELEDGHKVECQGKCRDVQLSI 79
ELGQLR +L G H S EL +++ ++ LE + CQ R V+ S
Sbjct: 461 ELGQLREQMLRQTGTCHQQKSTELPC-----LIQVKLLLWLEGSLLLPCQRNSRKVEPSF 297
Query: 80 QHIP 83
Q P
Sbjct: 296 QAYP 285
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.349 0.158 0.543
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,284,380
Number of Sequences: 63676
Number of extensions: 353432
Number of successful extensions: 2618
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 1942
Number of HSP's successfully gapped in prelim test: 69
Number of HSP's that attempted gapping in prelim test: 658
Number of HSP's gapped (non-prelim): 2029
length of query: 542
length of database: 12,639,632
effective HSP length: 102
effective length of query: 440
effective length of database: 6,144,680
effective search space: 2703659200
effective search space used: 2703659200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0126.7


