

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
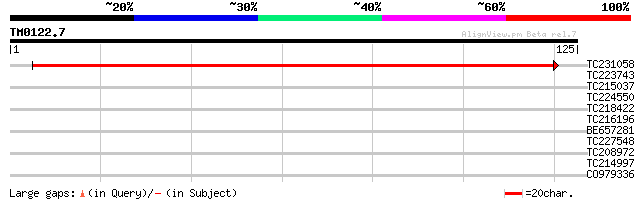
Query= TM0122.7
(125 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC231058 207 8e-55
TC223743 weakly similar to UP|Q9LV57 (Q9LV57) Aldehyde dehydroge... 28 1.1
TC215037 similar to UP|Q8RWP9 (Q8RWP9) Deetiolated 1-like protei... 27 1.8
TC224550 GB|AAN18113.1|23308287|BT000544 At1g78870/F9K20_8 {Arab... 27 2.4
TC218422 similar to UP|Q9FUY4 (Q9FUY4) Protein kinase AKINbetaga... 27 3.1
TC216196 weakly similar to UP|BRAT_DROME (Q8MQJ9) Brain tumor pr... 27 3.1
BE657281 26 4.1
TC227548 similar to UP|SYS_ARATH (Q39230) Seryl-tRNA synthetase ... 26 4.1
TC208972 25 7.0
TC214997 similar to UP|Q9SS31 (Q9SS31) Calmodulin-like protein (... 25 7.0
CO979336 25 9.1
>TC231058
Length = 573
Score = 207 bits (528), Expect = 8e-55
Identities = 100/116 (86%), Positives = 109/116 (93%)
Frame = +2
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
VYRISTAKEWEELQ+NGS+FGG+LDKSSAFIHLSKLDQVRSTLE F++N KEELYLLQID
Sbjct: 35 VYRISTAKEWEELQSNGSSFGGDLDKSSAFIHLSKLDQVRSTLENFFLNSKEELYLLQID 214
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSDGRFTCCLLD 121
AKKLGDGLVYE+VDG NSFPHFYGPSRSF PL LD VTKAEKL+L+DGRF+C LLD
Sbjct: 215 AKKLGDGLVYEIVDGKNSFPHFYGPSRSFGPLPLDAVTKAEKLTLTDGRFSCSLLD 382
>TC223743 weakly similar to UP|Q9LV57 (Q9LV57) Aldehyde dehydrogenase ALDH1a,
partial (5%)
Length = 733
Score = 28.1 bits (61), Expect = 1.1
Identities = 17/47 (36%), Positives = 24/47 (50%), Gaps = 4/47 (8%)
Frame = +1
Query: 66 AKKLGDGLVYEMVDGSNSFPH----FYGPSRSFSPLSLDVVTKAEKL 108
AK +G + V GSN FPH FY S+ F PL + + + K+
Sbjct: 148 AKLFINGEFLDSVSGSNYFPHHLPLFYNISKCFFPLYVKQIVYSLKV 288
>TC215037 similar to UP|Q8RWP9 (Q8RWP9) Deetiolated 1-like protein, partial
(44%)
Length = 785
Score = 27.3 bits (59), Expect = 1.8
Identities = 17/44 (38%), Positives = 22/44 (49%), Gaps = 3/44 (6%)
Frame = +3
Query: 22 GSTFGG---NLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLL 62
GS GG N D AF+ + +D + FY N +ELYLL
Sbjct: 99 GSVDGGVSRNADHHPAFVAVYNMDTTE--IVSFYQNSADELYLL 224
>TC224550 GB|AAN18113.1|23308287|BT000544 At1g78870/F9K20_8 {Arabidopsis
thaliana;} , complete
Length = 831
Score = 26.9 bits (58), Expect = 2.4
Identities = 12/21 (57%), Positives = 15/21 (71%), Gaps = 1/21 (4%)
Frame = +2
Query: 9 ISTAKEWEELQANGS-TFGGN 28
+ TAKEW L A+G+ FGGN
Sbjct: 491 VETAKEWTRLYASGA*CFGGN 553
>TC218422 similar to UP|Q9FUY4 (Q9FUY4) Protein kinase AKINbetagamma-2, partial
(81%)
Length = 1968
Score = 26.6 bits (57), Expect = 3.1
Identities = 13/50 (26%), Positives = 23/50 (46%)
Frame = +1
Query: 36 IHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGLVYEMVDGSNSFP 85
+HL+ L + + R++ NC L +LQ+ + G + SN P
Sbjct: 967 LHLASLSGILKCICRYFRNCSSSLPILQLPICAIPVGTWVPKIGESNRRP 1116
>TC216196 weakly similar to UP|BRAT_DROME (Q8MQJ9) Brain tumor protein,
partial (3%)
Length = 1679
Score = 26.6 bits (57), Expect = 3.1
Identities = 19/82 (23%), Positives = 32/82 (38%)
Frame = +3
Query: 38 LSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPL 97
L+K+ Q L + + E+L ++ K+G Y+ SFPH + P
Sbjct: 156 LAKIQQEDERLAKAQLEEDEQLARAIQESLKIGSPPQYDNGSSILSFPHLFPPGYR---- 323
Query: 98 SLDVVTKAEKLSLSDGRFTCCL 119
+ K + GRF C+
Sbjct: 324 ----ICAGCKTEIGQGRFLSCM 377
>BE657281
Length = 719
Score = 26.2 bits (56), Expect = 4.1
Identities = 12/38 (31%), Positives = 21/38 (54%)
Frame = +2
Query: 73 LVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSL 110
LV + +DG+ + P +GP R S L + + +K +L
Sbjct: 41 LVVQSMDGNYTTPMIHGPPRKISDLYISPTQEKKKKTL 154
>TC227548 similar to UP|SYS_ARATH (Q39230) Seryl-tRNA synthetase
(Serine--tRNA ligase) (SerRS) , partial (48%)
Length = 1594
Score = 26.2 bits (56), Expect = 4.1
Identities = 11/24 (45%), Positives = 14/24 (57%)
Frame = -2
Query: 14 EWEELQANGSTFGGNLDKSSAFIH 37
+W++L G T NL SSAF H
Sbjct: 471 QWKQLDREGLTLCRNLQSSSAFPH 400
>TC208972
Length = 1190
Score = 25.4 bits (54), Expect = 7.0
Identities = 19/46 (41%), Positives = 25/46 (54%)
Frame = -3
Query: 58 ELYLLQIDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVT 103
E YLLQ + + L LV V S + + GPS +FS L L +VT
Sbjct: 219 EKYLLQQNKENLCLTLVVHQVHLSPTKSYQEGPSSAFSLLYLLLVT 82
>TC214997 similar to UP|Q9SS31 (Q9SS31) Calmodulin-like protein (At3g10190),
partial (45%)
Length = 1107
Score = 25.4 bits (54), Expect = 7.0
Identities = 12/34 (35%), Positives = 16/34 (46%)
Frame = -1
Query: 85 PHFYGPSRSFSPLSLDVVTKAEKLSLSDGRFTCC 118
PH G PL+ DV A + ++ GRF C
Sbjct: 336 PHVNGGGGDLGPLAGDVRENAGRSLVTGGRFVIC 235
>CO979336
Length = 413
Score = 25.0 bits (53), Expect = 9.1
Identities = 13/30 (43%), Positives = 17/30 (56%)
Frame = +3
Query: 82 NSFPHFYGPSRSFSPLSLDVVTKAEKLSLS 111
N H + S SFSP+S +EK+SLS
Sbjct: 18 NLHKHLHPSSSSFSPISTLGFKHSEKISLS 107
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.136 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,556,618
Number of Sequences: 63676
Number of extensions: 65341
Number of successful extensions: 270
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 270
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 270
length of query: 125
length of database: 12,639,632
effective HSP length: 86
effective length of query: 39
effective length of database: 7,163,496
effective search space: 279376344
effective search space used: 279376344
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0122.7


