

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.6
(378 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
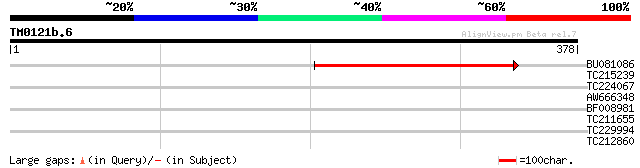
Sequences producing significant alignments: (bits) Value
BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza s... 141 4e-34
TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%) 35 0.073
TC224067 34 0.12
AW666348 similar to GP|20302596|dbj Ser/Thr kinase {Arabidopsis ... 30 2.3
BF008981 28 5.2
TC211655 28 5.2
TC229994 28 5.2
TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and... 28 8.9
>BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 426
Score = 141 bits (356), Expect = 4e-34
Identities = 68/136 (50%), Positives = 91/136 (66%)
Frame = +2
Query: 204 KSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGLCNGTRMIVN 263
KSD +T+EF+N S +PNH+IKLKV IML+RN+DQ GLCN TR+I+
Sbjct: 17 KSDTIESEGFSTITTEFINSLSTSRLPNHKIKLKVDSRIMLLRNLDQNEGLCNSTRLIIT 196
Query: 264 ALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFAMTINKSQGQ 323
+II A +++ G T +IPR++ +PS + PFK RR+FP+I+ +AMTINKSQGQ
Sbjct: 197 MFADHIIEAKIMSGKGQGNTVYIPRLATSPSQSPWPFKLIRRKFPIIVSYAMTINKSQGQ 376
Query: 324 SSTHVGLYLPRPVFTH 339
VGLYLP PVF+H
Sbjct: 377 LLASVGLYLPTPVFSH 424
>TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%)
Length = 340
Score = 34.7 bits (78), Expect = 0.073
Identities = 15/22 (68%), Positives = 19/22 (86%)
Frame = -3
Query: 303 QRRQFPVILCFAMTINKSQGQS 324
QR FP+I+CFAMT NKS+GQ+
Sbjct: 338 QR**FPLIVCFAMTTNKSEGQT 273
>TC224067
Length = 415
Score = 33.9 bits (76), Expect = 0.12
Identities = 22/51 (43%), Positives = 30/51 (58%), Gaps = 1/51 (1%)
Frame = -2
Query: 300 FKFQRRQFPVILCFAMTINKSQGQSSTHV-GLYLPRPVFTHVQLYVALSRV 349
FKFQ ++C+ M INKS GQ+ + V G+ LPRP YV L++V
Sbjct: 387 FKFQ--DVNSLVCYTMKINKSHGQTLSQVGGVLLPRPN**SHGQYVTLNQV 241
>AW666348 similar to GP|20302596|dbj Ser/Thr kinase {Arabidopsis thaliana},
partial (10%)
Length = 420
Score = 29.6 bits (65), Expect = 2.3
Identities = 24/70 (34%), Positives = 35/70 (49%), Gaps = 2/70 (2%)
Frame = -3
Query: 277 RNTMGETTFIP-RMSLTPSNADIPFKFQRRQFPVI-LCFAMTINKSQGQSSTHVGLYLPR 334
R+T+ TFI SL S + PF++Q R FP+ +C++ I S Q H +
Sbjct: 223 RHTIVTETFISITRSLINSWSWSPFRYQGRNFPIYHICYSGNILISHVQLCNHFSCH--- 53
Query: 335 PVFTHVQLYV 344
+ VQLYV
Sbjct: 52 -TYCCVQLYV 26
>BF008981
Length = 419
Score = 28.5 bits (62), Expect = 5.2
Identities = 26/92 (28%), Positives = 44/92 (47%), Gaps = 2/92 (2%)
Frame = +2
Query: 143 PLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEIE-YLNCDT 201
P L V +Y KL L++N Q +I +L VE + F+ +++L + + + + +
Sbjct: 116 PFLISVGGNY-KLWLELDRNGAEQNYSI--SSLLIVEVVLTFL*SVMLVKSLSLHSHAFS 286
Query: 202 PCK-SDEDSGVNAEWVTSEFLNDYKCSGIPNH 232
C S + G N WV + LN CS P +
Sbjct: 287 SCSFSASEIGTNPSWVPLKSLNSLCCSSKPRN 382
>TC211655
Length = 596
Score = 28.5 bits (62), Expect = 5.2
Identities = 16/46 (34%), Positives = 25/46 (53%)
Frame = -2
Query: 121 QPLMKLNHSFRFLLIFIEPCKDPLLEIVNFSYPKLLFNLEKNSFFQ 166
+PL K SF + ++F+ CK PLL F K L ++ + FF+
Sbjct: 526 KPLSKDPSSFMYHVLFLSSCKTPLLLSAKFMVRKYL-HITFSMFFE 392
>TC229994
Length = 459
Score = 28.5 bits (62), Expect = 5.2
Identities = 26/92 (28%), Positives = 44/92 (47%), Gaps = 2/92 (2%)
Frame = +3
Query: 143 PLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEIE-YLNCDT 201
P L V +Y KL L++N Q +I +L VE + F+ +++L + + + + +
Sbjct: 93 PFLISVGGNY-KLWLELDRNGAEQNYSI--SSLLIVEVVLTFL*SVMLVKSLSLHSHAFS 263
Query: 202 PCK-SDEDSGVNAEWVTSEFLNDYKCSGIPNH 232
C S + G N WV + LN CS P +
Sbjct: 264 SCSFSASEIGTNPSWVPLKSLNSLCCSSKPRN 359
>TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and
recombination protein PIF1, mitochondrial precursor,
partial (8%)
Length = 596
Score = 27.7 bits (60), Expect = 8.9
Identities = 18/39 (46%), Positives = 24/39 (61%)
Frame = +2
Query: 313 FAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKS 351
+AM+I+K QG + V L R F +YVALSRV+S
Sbjct: 5 WAMSIHKCQGMTLERVHTDLSR-AFGCGMVYVALSRVRS 118
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.343 0.151 0.491
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,282,406
Number of Sequences: 63676
Number of extensions: 247710
Number of successful extensions: 2156
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 2151
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2156
length of query: 378
length of database: 12,639,632
effective HSP length: 99
effective length of query: 279
effective length of database: 6,335,708
effective search space: 1767662532
effective search space used: 1767662532
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0121b.6


