

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.2
(253 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
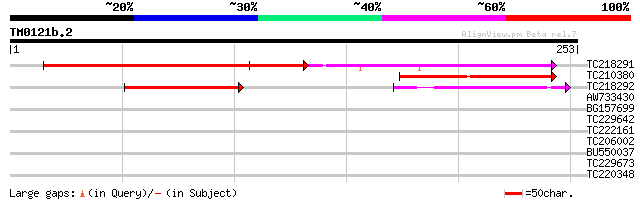
Score E
Sequences producing significant alignments: (bits) Value
TC218291 similar to GB|AAM91387.1|22137084|AY133557 At3g13350/MD... 124 5e-29
TC210380 similar to PIR|B96789|B96789 protein T23E18.4 [imported... 95 3e-20
TC218292 similar to GB|AAM91387.1|22137084|AY133557 At3g13350/MD... 64 7e-11
AW733430 28 5.2
BG157699 27 8.9
TC229642 homologue to UP|Q9ZQT9 (Q9ZQT9) WERBP-1 protein, partia... 27 8.9
TC222161 weakly similar to UP|Q9SGD7 (Q9SGD7) T23G18.9, partial ... 27 8.9
TC206002 similar to GB|AAL49941.1|17979249|AY070475 AT3g04610/F7... 27 8.9
BU550037 27 8.9
TC229673 similar to GB|AAM26654.1|20856224|AY101533 At1g14810/F1... 27 8.9
TC220348 homologue to UP|SECA_PEA (Q41062) Preprotein translocas... 27 8.9
>TC218291 similar to GB|AAM91387.1|22137084|AY133557 At3g13350/MDC11_14
{Arabidopsis thaliana;} , partial (22%)
Length = 697
Score = 124 bits (310), Expect = 5e-29
Identities = 60/118 (50%), Positives = 80/118 (66%)
Frame = +3
Query: 16 KFYPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGY 75
K YP P A ++ IV DA LF TL+ FH +G+KY + +GG LDLH L+VEVT R G
Sbjct: 195 KPYPAPTARYQDIVRDANLFWGTLQAFHKILGTKYKVATVGGTSLDLHRLFVEVTSRGGI 374
Query: 76 EKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGT 133
EKV+ ++KW+EV FNF T TSAS V++K Y ++LY++EQV++F QG P GT
Sbjct: 375 EKVIVDRKWKEVILTFNFKDTITSASXVVRKSYLSMLYHYEQVYYFGRQGIPPPTPGT 548
Score = 43.1 bits (100), Expect = 1e-04
Identities = 41/149 (27%), Positives = 63/149 (41%), Gaps = 12/149 (8%)
Frame = +3
Query: 108 YWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTLCFMEMDSLQGMYPNI------HAVTEF 161
+W L F ++ K + + GT L+ L F+E+ S G+ I + F
Sbjct: 252 FWGTLQAFHKILGTKYK--VATVGGTSLDLHRL-FVEVTSRGGIEKVIVDRKWKEVILTF 422
Query: 162 ESICDVLSGSSSSGKPDLAL------VEYAPKHANNGPESNVEAKGYPRYGEGRIEEKFD 215
+ S S K L++ V Y + P + + G I+ KFD
Sbjct: 423 NFKDTITSASXVVRKSYLSMLYHYEQVYYFGRQGIPPPTPGTPVQAHDDMVSGTIDAKFD 602
Query: 216 CGYLVSVKLGSEVLRGVIFHPEETVPPPS 244
GY+V+V LGSE L+GV+FH + V S
Sbjct: 603 GGYVVTVILGSEQLKGVLFHVPDNVSQSS 689
>TC210380 similar to PIR|B96789|B96789 protein T23E18.4 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(32%)
Length = 807
Score = 95.1 bits (235), Expect = 3e-20
Identities = 46/70 (65%), Positives = 54/70 (76%)
Frame = +2
Query: 175 GKPDLALVEYAPKHANNGPESNVEAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIF 234
G+P+LA+VEY+PK +N PES E G G IE KFDCGYLVSVKLGSEVLRGV++
Sbjct: 8 GRPELAMVEYSPKPMDNSPESRAEDTSCLS-GNGTIEGKFDCGYLVSVKLGSEVLRGVLY 184
Query: 235 HPEETVPPPS 244
HPE+ VPPPS
Sbjct: 185 HPEQLVPPPS 214
>TC218292 similar to GB|AAM91387.1|22137084|AY133557 At3g13350/MDC11_14
{Arabidopsis thaliana;} , partial (37%)
Length = 972
Score = 63.9 bits (154), Expect = 7e-11
Identities = 28/53 (52%), Positives = 38/53 (70%)
Frame = +3
Query: 52 IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVL 104
I +GG LDLH L+VEVT R G EKV+ ++KW+EV FNF T T+AS+++
Sbjct: 48 ISTVGGTPLDLHRLFVEVTSRGGIEKVIVDRKWKEVILTFNFKDTITNASFMI 206
Score = 48.5 bits (114), Expect = 3e-06
Identities = 30/79 (37%), Positives = 43/79 (53%)
Frame = +2
Query: 172 SSSGKPDLALVEYAPKHANNGPESNVEAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRG 231
SS+ P++A V N+ P + + + G I+ KFD GY+V+V LGSE L+G
Sbjct: 239 SSTTIPEVAAV-------NDSPVQSTPVQAHDDMVSGTIDAKFDVGYVVTVTLGSEQLKG 397
Query: 232 VIFHPEETVPPPSSRVVGT 250
V+FH + V SS GT
Sbjct: 398 VLFHVPDNV-SQSSHAEGT 451
>AW733430
Length = 184
Score = 27.7 bits (60), Expect = 5.2
Identities = 14/36 (38%), Positives = 22/36 (60%), Gaps = 2/36 (5%)
Frame = +2
Query: 208 GRIEEKFDCGYLVSVKLG--SEVLRGVIFHPEETVP 241
G I+ F+ GYL+SVK+ LRG++F ++ P
Sbjct: 29 GVIDGTFNAGYLLSVKVADTDAFLRGLVFVEDQVSP 136
>BG157699
Length = 421
Score = 26.9 bits (58), Expect = 8.9
Identities = 12/31 (38%), Positives = 16/31 (50%)
Frame = +3
Query: 101 SYVLKKHYWNLLYNFEQVHFFKVQGPITPPA 131
SY L H+ L + Q HF +Q P PP+
Sbjct: 153 SYFLHLHHPPLYPSSSQTHFLSIQNPNFPPS 245
>TC229642 homologue to UP|Q9ZQT9 (Q9ZQT9) WERBP-1 protein, partial (18%)
Length = 934
Score = 26.9 bits (58), Expect = 8.9
Identities = 16/48 (33%), Positives = 23/48 (47%)
Frame = +2
Query: 170 GSSSSGKPDLALVEYAPKHANNGPESNVEAKGYPRYGEGRIEEKFDCG 217
GSS++ K LA E N+G + KG + EG +E+ CG
Sbjct: 242 GSSNATKDSLAKNEMEASQVNHGRSGPDQVKGSTTFEEGSLEK---CG 376
>TC222161 weakly similar to UP|Q9SGD7 (Q9SGD7) T23G18.9, partial (52%)
Length = 978
Score = 26.9 bits (58), Expect = 8.9
Identities = 12/43 (27%), Positives = 22/43 (50%), Gaps = 1/43 (2%)
Frame = +1
Query: 102 YVLKKHYWNLLYNFEQVHFFKVQGPITPPA-GTVLNSFTLCFM 143
Y + ++ + LL +HF +Q + PP G +L S C++
Sbjct: 157 YAMPEYLFQLLRKST*LHFLWIQATLAPPVKGKMLRSLCACYV 285
>TC206002 similar to GB|AAL49941.1|17979249|AY070475 AT3g04610/F7O18_9
{Arabidopsis thaliana;} , partial (53%)
Length = 1624
Score = 26.9 bits (58), Expect = 8.9
Identities = 13/29 (44%), Positives = 15/29 (50%)
Frame = +3
Query: 56 GGRKLDLHTLYVEVTRRSGYEKVVAEKKW 84
GG T+ +E G E VVAEKKW
Sbjct: 96 GGEHEAEETIEIEAEVHGGEEAVVAEKKW 182
>BU550037
Length = 728
Score = 26.9 bits (58), Expect = 8.9
Identities = 14/38 (36%), Positives = 18/38 (46%)
Frame = +2
Query: 84 WREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFF 121
W S+ NFS TT + + L WN L Q +FF
Sbjct: 158 WLSSSSLENFSFTTNTVAIGLHLSMWNFL-KISQAYFF 268
>TC229673 similar to GB|AAM26654.1|20856224|AY101533 At1g14810/F10B6_6
{Arabidopsis thaliana;} , partial (51%)
Length = 1442
Score = 26.9 bits (58), Expect = 8.9
Identities = 17/45 (37%), Positives = 21/45 (45%)
Frame = +1
Query: 109 WNLLYNFEQVHFFKVQGPITPPAGTVLNSFTLCFMEMDSLQGMYP 153
WN ++ V FK+Q I+ A N FTLCF L G P
Sbjct: 703 WNWKTSWSVVVCFKIQHLISLFAA*NPNGFTLCFAPQPPLLGPPP 837
>TC220348 homologue to UP|SECA_PEA (Q41062) Preprotein translocase secA
subunit, chloroplast precursor, partial (25%)
Length = 776
Score = 26.9 bits (58), Expect = 8.9
Identities = 12/33 (36%), Positives = 20/33 (60%)
Frame = -3
Query: 162 ESICDVLSGSSSSGKPDLALVEYAPKHANNGPE 194
ES+C L +S+G+P L ++E + A + PE
Sbjct: 450 ESLCSTLVVPTSTGRPVLCILEISTTTARHFPE 352
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.137 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,415,944
Number of Sequences: 63676
Number of extensions: 172552
Number of successful extensions: 826
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 821
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 824
length of query: 253
length of database: 12,639,632
effective HSP length: 95
effective length of query: 158
effective length of database: 6,590,412
effective search space: 1041285096
effective search space used: 1041285096
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0121b.2


