

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
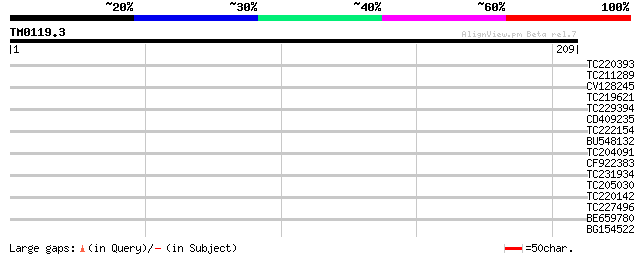
Query= TM0119.3
(209 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC220393 similar to PIR|S67495|S67495 huntingtin-associated prot... 31 0.35
TC211289 29 1.8
CV128245 29 1.8
TC219621 similar to UP|Q94AC8 (Q94AC8) AT4g28030/T13J8_140, part... 28 2.3
TC229394 similar to GB|BAA98102.1|8978074|AB025609 protein serin... 28 3.0
CD409235 28 3.0
TC222154 28 3.9
BU548132 28 3.9
TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 28 3.9
CF922383 28 3.9
TC231934 similar to UP|NFL1_HUMAN (Q14494) Nuclear factor erythr... 28 3.9
TC205030 UP|ACCD_SOYBN (P49158) Acetyl-coenzyme A carboxylase ca... 27 5.1
TC220142 similar to UP|Q9FKR4 (Q9FKR4) Emb|CAB41546.1|, partial ... 27 5.1
TC227496 similar to UP|Q7XA59 (Q7XA59) M355, partial (28%) 27 6.7
BE659780 similar to GP|4768916|gb| CLC-Nt2 protein {Nicotiana ta... 27 8.7
BG154522 27 8.7
>TC220393 similar to PIR|S67495|S67495 huntingtin-associated protein HAP1-A -
rat {Rattus norvegicus;} , partial (5%)
Length = 872
Score = 31.2 bits (69), Expect = 0.35
Identities = 29/112 (25%), Positives = 50/112 (43%), Gaps = 6/112 (5%)
Frame = +1
Query: 55 RGKLKNHFDLNKFPKIEDSFEIESETIEHSLDVIKTVEDSLNATKIVEESDIESEEEGRP 114
+GKL + + +K E + +IES ++ ++ K V+ N + E+SD++ EEEG
Sbjct: 148 KGKLISREEFDKIMAKEKTEQIESTEMDEAM--AKAVDIGENDDE--EDSDVDGEEEGEE 315
Query: 115 TIIRKKQIWHEKDTELDSEECAP------STSRSFDSIENLHELNKDAEEED 160
+ + EL + P S + ENL + D EEE+
Sbjct: 316 EKLSDYWSVLKSTPELRQSKPKPKKEGPLSLDEVIEESENLTDFLMDFEEEE 471
>TC211289
Length = 823
Score = 28.9 bits (63), Expect = 1.8
Identities = 26/81 (32%), Positives = 38/81 (46%), Gaps = 2/81 (2%)
Frame = +1
Query: 81 IEHSLDVIKTVEDSLNATKIVEESDIESEEEGRPTIIRKKQIWHEKDTE--LDSEECAPS 138
+EH +D+ +DSL K+ D + EE T + + H KD + + C
Sbjct: 154 LEHEIDM----KDSLALAKLYLNFDGKEEEA---TDVYSELWKHVKDFSNCVQNGLCVEK 312
Query: 139 TSRSFDSIENLHELNKDAEEE 159
+RS S N+ ELN AEEE
Sbjct: 313 CNRSGASDSNMPELNDLAEEE 375
>CV128245
Length = 755
Score = 28.9 bits (63), Expect = 1.8
Identities = 15/32 (46%), Positives = 20/32 (61%)
Frame = +2
Query: 15 VAPNGIYTFTSFLFPALLFSSSHYMGMKSHTS 46
+A ++ + SFLF L FSSSH+ SHTS
Sbjct: 503 LAQKSLFLYFSFLFFLLFFSSSHH-SFYSHTS 595
>TC219621 similar to UP|Q94AC8 (Q94AC8) AT4g28030/T13J8_140, partial (61%)
Length = 849
Score = 28.5 bits (62), Expect = 2.3
Identities = 14/38 (36%), Positives = 21/38 (54%)
Frame = -3
Query: 137 PSTSRSFDSIENLHELNKDAEEEDHFSHEREAMKNEVM 174
P+T RS +ENLHE +K +E + + EA E +
Sbjct: 259 PTTIRSSPLVENLHEEHKSSERVAIWGNGSEAFMQETL 146
>TC229394 similar to GB|BAA98102.1|8978074|AB025609 protein serine/threonine
kinase-like {Arabidopsis thaliana;} , partial (63%)
Length = 1170
Score = 28.1 bits (61), Expect = 3.0
Identities = 17/71 (23%), Positives = 35/71 (48%), Gaps = 4/71 (5%)
Frame = +3
Query: 57 KLKNHFDL---NKFPKIEDSFEIESETIEHSLD-VIKTVEDSLNATKIVEESDIESEEEG 112
+L+N + L K K+ DS ++ S+ ++++++ +L + + ES
Sbjct: 711 RLRNQYSLPAARKIAKLADSCLKKNPEDRPSMSQIVESLKQALQYSDTTSQDIAESSSSS 890
Query: 113 RPTIIRKKQIW 123
R ++RKK IW
Sbjct: 891 RSKLVRKK*IW 923
>CD409235
Length = 595
Score = 28.1 bits (61), Expect = 3.0
Identities = 20/85 (23%), Positives = 38/85 (44%), Gaps = 2/85 (2%)
Frame = +1
Query: 89 KTVEDSLNATKIVEESDIESEEEGRPTIIRKKQIWHEKDTE--LDSEECAPSTSRSFDSI 146
+TV + A +I ++++ + + + + I TE + S+ P +R F+ +
Sbjct: 64 RTVHTNQMAKEIKKDTEHKGKRRTKNLNSHRFSICQASSTEKHISSKHFHPLRNRGFNDL 243
Query: 147 ENLHELNKDAEEEDHFSHEREAMKN 171
+ LH HFSH R A +N
Sbjct: 244 QYLH----------HFSHSRPARRN 288
>TC222154
Length = 600
Score = 27.7 bits (60), Expect = 3.9
Identities = 24/69 (34%), Positives = 33/69 (47%), Gaps = 5/69 (7%)
Frame = +1
Query: 68 PKIED----SFEIESETIEHSLDVIKTVED-SLNATKIVEESDIESEEEGRPTIIRKKQI 122
P +ED S IES + +DV K + N K+ EE+ IES + G T + K
Sbjct: 358 PLVEDLKVVSHCIESIANKACVDVSKVDWSYTYNRKKLPEENGIESNQNGLRTRLVPKDW 537
Query: 123 WHEKDTELD 131
W E EL+
Sbjct: 538 WVEDLCELE 564
>BU548132
Length = 613
Score = 27.7 bits (60), Expect = 3.9
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = -3
Query: 132 SEECAPSTSRSFDSIENLHELNKDAEEEDH 161
SE+ A + R +E+ H + DAE+EDH
Sbjct: 170 SEDVAFTLERPMTLVEDFHTKDVDAEDEDH 81
>TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(70%)
Length = 2075
Score = 27.7 bits (60), Expect = 3.9
Identities = 22/86 (25%), Positives = 42/86 (48%)
Frame = +2
Query: 42 KSHTSKEDDLKSHRGKLKNHFDLNKFPKIEDSFEIESETIEHSLDVIKTVEDSLNATKIV 101
KS S E LK + K +N ++ ++E+ E+E E +E ++ + E+
Sbjct: 56 KSSLSTEPPLKQQQPKPEN--PPQEYEEVEE--EVEEEEVEEEVEEEEEEEEEDEE---- 211
Query: 102 EESDIESEEEGRPTIIRKKQIWHEKD 127
EE + E EEE + +++Q ++D
Sbjct: 212 EEEEEEEEEEENDVVKQQQQQQQDED 289
>CF922383
Length = 691
Score = 27.7 bits (60), Expect = 3.9
Identities = 29/121 (23%), Positives = 52/121 (42%), Gaps = 25/121 (20%)
Frame = +1
Query: 78 SETIEHSLDVIKTVEDSLNATKIVEESD-----IESEEEGRPTIIRKKQIWHEKDTEL-- 130
S+ E +++ K V ++ TK+VEE+D I+ P+ ++K++ + D E
Sbjct: 172 SQYKEANMEEEKKVISEVSVTKVVEEADHKNESIKETNGDLPSEVKKEEEENAFDGEFIK 351
Query: 131 -------------DSEECAPSTSRSF----DSIENLH-ELNKDAEEEDHFSHEREAMKNE 172
+E + S SR F + I+ L EL + E HE + +K E
Sbjct: 352 VEKEENSIDDKSHKTERSSDSPSREFLEAQEKIQELEVELQRLTESLKTSEHENDQLKGE 531
Query: 173 V 173
+
Sbjct: 532 I 534
>TC231934 similar to UP|NFL1_HUMAN (Q14494) Nuclear factor erythroid 2
related factor 1 (NF-E2 related factor 1) (NFE2-related
factor 1) (Nuclear factor, erythroid derived 2, like 1)
(Transcription factor 11) (Transcription factor HBZ17)
(Transcription factor LCR-F1) (Locus con, partial (3%)
Length = 696
Score = 27.7 bits (60), Expect = 3.9
Identities = 25/91 (27%), Positives = 42/91 (45%)
Frame = +2
Query: 82 EHSLDVIKTVEDSLNATKIVEESDIESEEEGRPTIIRKKQIWHEKDTELDSEECAPSTSR 141
E +++ ++ D+LN V ES ++ E++ R + KQ+ S+SR
Sbjct: 140 EPEVEICASMLDALNECIQVSESHLD-EKQVRSIVDEIKQV------------LTASSSR 280
Query: 142 SFDSIENLHELNKDAEEEDHFSHEREAMKNE 172
HE + A+EED + ERE +K E
Sbjct: 281 K-------HERAERAKEEDFDAEERELLKEE 352
>TC205030 UP|ACCD_SOYBN (P49158) Acetyl-coenzyme A carboxylase carboxyl
transferase subunit beta (ACCASE beta chain) , complete
Length = 4503
Score = 27.3 bits (59), Expect = 5.1
Identities = 14/27 (51%), Positives = 18/27 (65%), Gaps = 2/27 (7%)
Frame = -1
Query: 42 KSHTSKE--DDLKSHRGKLKNHFDLNK 66
K T+KE D +KSHRG L N + +NK
Sbjct: 4167 KIKTTKETRDSIKSHRGPLWNKWKINK 4087
>TC220142 similar to UP|Q9FKR4 (Q9FKR4) Emb|CAB41546.1|, partial (37%)
Length = 755
Score = 27.3 bits (59), Expect = 5.1
Identities = 21/97 (21%), Positives = 51/97 (51%), Gaps = 5/97 (5%)
Frame = +1
Query: 85 LDVIKT-VEDSLNATKIVEESDIESEE----EGRPTIIRKKQIWHEKDTELDSEECAPST 139
LDV++ ++D + A ++ E + + +E EG T++ + +K T+ +S S
Sbjct: 298 LDVLQNQIDDFITAQEVKLEEERKEKEALAAEGGWTVVVHHK-GRKKTTDSESGIAVGSV 474
Query: 140 SRSFDSIENLHELNKDAEEEDHFSHEREAMKNEVMSM 176
+++ + + +K+ ++ + REA +NE+M++
Sbjct: 475 AQAAVENKMTKKKHKEVGQDFYRCQRREAQRNEIMTL 585
>TC227496 similar to UP|Q7XA59 (Q7XA59) M355, partial (28%)
Length = 688
Score = 26.9 bits (58), Expect = 6.7
Identities = 13/55 (23%), Positives = 23/55 (41%)
Frame = +2
Query: 105 DIESEEEGRPTIIRKKQIWHEKDTELDSEECAPSTSRSFDSIENLHELNKDAEEE 159
D E +++ T + +W L+ A + S +E L + + D EEE
Sbjct: 317 DDEDDDDYYATTAPPESVWAASPVTLNDAAAAAAAEESESELEGLDDADDDGEEE 481
>BE659780 similar to GP|4768916|gb| CLC-Nt2 protein {Nicotiana tabacum},
partial (20%)
Length = 644
Score = 26.6 bits (57), Expect = 8.7
Identities = 16/47 (34%), Positives = 25/47 (53%), Gaps = 1/47 (2%)
Frame = +3
Query: 1 MENCKHTNMQYIYRVAPNGIYTFTSFLFPALLFSSSHY-MGMKSHTS 46
+ CK TN ++ + T TS +FP+L SS+H + SH+S
Sbjct: 363 VRGCKSTN------ISISSFVTATSSIFPSLSASSTHVNFSLTSHSS 485
>BG154522
Length = 220
Score = 26.6 bits (57), Expect = 8.7
Identities = 12/41 (29%), Positives = 21/41 (50%)
Frame = +2
Query: 111 EGRPTIIRKKQIWHEKDTELDSEECAPSTSRSFDSIENLHE 151
+G + +RKK+ H +TE +S +CA ++ HE
Sbjct: 53 KGESSCMRKKRCCHS*ETEKESVDCASVFGEPMKCMQARHE 175
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.132 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,428,028
Number of Sequences: 63676
Number of extensions: 105100
Number of successful extensions: 474
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 471
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 474
length of query: 209
length of database: 12,639,632
effective HSP length: 93
effective length of query: 116
effective length of database: 6,717,764
effective search space: 779260624
effective search space used: 779260624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0119.3


