

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
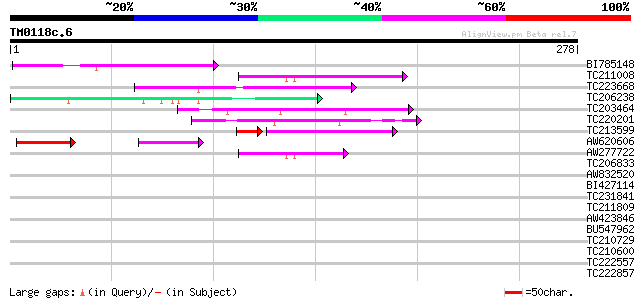
Query= TM0118c.6
(278 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI785148 homologue to GP|25166165|dbj murF {Wigglesworthia brevi... 78 5e-15
TC211008 68 5e-12
TC223668 54 8e-08
TC206238 weakly similar to UP|Q7PX96 (Q7PX96) AgCP12237 (Fragmen... 50 1e-06
TC203464 similar to GB|AAF35920.1|7025820|AC005892 L505.8 {Leish... 50 1e-06
TC220201 similar to UP|Q9ERI5 (Q9ERI5) Apoptotic cell clearance ... 47 1e-05
TC213599 homologue to UP|EXBB_PASHA (P72202) Biopolymer transpor... 43 6e-05
AW620606 34 1e-04
AW277722 similar to SP|O65719|HS73_ Heat shock cognate 70 kDa pr... 43 2e-04
TC206833 homologue to UP|Q6PDS7 (Q6PDS7) Hist1h4h protein (Fragm... 40 0.001
AW832520 40 0.001
BI427114 40 0.002
TC231841 40 0.002
TC211809 40 0.002
AW423846 38 0.004
BU547962 38 0.006
TC210729 similar to UP|Q7QHL7 (Q7QHL7) AgCP7056 (Fragment), part... 37 0.008
TC210600 similar to UP|Q51534 (Q51534) PilW protein, partial (7%) 37 0.008
TC222557 37 0.008
TC222857 37 0.010
>BI785148 homologue to GP|25166165|dbj murF {Wigglesworthia brevipalpis},
partial (2%)
Length = 422
Score = 77.8 bits (190), Expect = 5e-15
Identities = 40/108 (37%), Positives = 53/108 (49%), Gaps = 7/108 (6%)
Frame = -2
Query: 2 PRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDH-------GSWIRSQARSRNS 54
P CS P ED LHCLRD PH++EIWL FG H S ++SQ
Sbjct: 319 PHCSNPREDSLHCLRDFPHAKEIWL--------GFGFAPHLSSLPLM*SGLKSQLNGSKV 164
Query: 55 VRFLAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFITFSNGVA 102
F VW V KW NN +F+D W + + IC HD+ ++ ++ A
Sbjct: 163 FLFTTIVWWV*KWGNNCIFDDEKWDIQRVVQSICFYHDDLVSTNSDCA 20
>TC211008
Length = 516
Score = 67.8 bits (164), Expect = 5e-12
Identities = 36/87 (41%), Positives = 51/87 (58%), Gaps = 4/87 (4%)
Frame = +1
Query: 113 WQPPKQGVVKLNMDESFWEQDR--CMGA--GGLIRDDQGIWLGGFYAHRSGGNALIAEAT 168
W+ P G V LN D S +E R C G+ GGL+RD G +LGGF + + +AE
Sbjct: 64 WRCPXNGWVCLNTDGSVFENHRNGCSGSACGGLVRDSSGCYLGGFTVNLGNTSVTLAELW 243
Query: 169 ALLLGLELVWDLGYRQVMVEVDCGELL 195
++ GL+L WDLG ++V V++D G L
Sbjct: 244 GVVHGLKLAWDLGCKKVKVDIDSGNAL 324
>TC223668
Length = 866
Score = 53.9 bits (128), Expect = 8e-08
Identities = 37/111 (33%), Positives = 49/111 (43%), Gaps = 2/111 (1%)
Frame = -3
Query: 62 WGVWKWRNNMVFEDSPWLVDEACRRICHEH--DEFITFSNGVAGDHGSWLSSRWQPPKQG 119
W WKW N ++F W + C + + + V+ +W S PP
Sbjct: 729 WWAWKWCN*VIFSPLIWSFQFIKKIHC*GYCLQSSLVSKDLVSSKRLTWTSP---PPLLN 559
Query: 120 VVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATAL 170
VKLN+D S + M GGLIRD G WL F H GN L++E AL
Sbjct: 558 EVKLNVDGSGHYEPHVMEVGGLIRDGVGRWLFRFARHCGLGNPLLSELFAL 406
>TC206238 weakly similar to UP|Q7PX96 (Q7PX96) AgCP12237 (Fragment), partial
(9%)
Length = 1259
Score = 50.1 bits (118), Expect = 1e-06
Identities = 45/177 (25%), Positives = 67/177 (37%), Gaps = 24/177 (13%)
Frame = +3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLR-LGALAWTNFGVGDHGSWIRSQARSRNSVRFLA 59
CPRC EE LH LRDCP W L F GD W+ + + F
Sbjct: 225 CPRCPELEETCLHALRDCPKVAAFWRSVLPKKLAPKFFNGDVAVWLETNLSFSEAAFFWP 404
Query: 60 GVWGV-----WKWRNNMVF-EDSPW----LVD---------EACRRICHEH----DEFIT 96
+G+ W+ RN++VF +D W L D + C R H +
Sbjct: 405 TFFGIAVELLWESRNDLVFYKDGTWDYLDLSDITDIVFNRYKDCMRAHASHILMPRNLLK 584
Query: 97 FSNGVAGDHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGF 153
+ + HG WL ++LN+ ++ GG+ RD+ ++ GF
Sbjct: 585 WRRPLPLSHGHWL-----------LRLNVSGAYDRSSDTAACGGIFRDNNDRFVLGF 722
>TC203464 similar to GB|AAF35920.1|7025820|AC005892 L505.8 {Leishmania
major;} , partial (8%)
Length = 1386
Score = 49.7 bits (117), Expect = 1e-06
Identities = 37/121 (30%), Positives = 62/121 (50%), Gaps = 5/121 (4%)
Frame = +2
Query: 83 ACRRICHEHDEFITFSNGVAGDH---GSWLSSRWQPPKQGVVKLNMDESFWE-QDRCMGA 138
+ RR E D+ G +G H G+ SRW+ P+ G VKLN+D S + G
Sbjct: 407 SARRESKERDD------GGSGSHPREGATTVSRWKKPEIGWVKLNVDGSRDPYKSSSAGC 568
Query: 139 GGLIRDDQGIWLGGFYAHRSGGNAL-IAEATALLLGLELVWDLGYRQVMVEVDCGELLQV 197
GG++RD WL GF + A+ E A+L GL++ ++ ++++VE D ++ +
Sbjct: 569 GGVLRDASAKWLRGFAKKLNPTYAVHQTELEAILTGLKVASEMNVKKLIVESDSDSVVSM 748
Query: 198 M 198
+
Sbjct: 749 V 751
>TC220201 similar to UP|Q9ERI5 (Q9ERI5) Apoptotic cell clearance receptor
PtdSerR (Phosphatidylserine receptor), partial (5%)
Length = 1185
Score = 47.0 bits (110), Expect = 1e-05
Identities = 39/117 (33%), Positives = 54/117 (45%), Gaps = 4/117 (3%)
Frame = +1
Query: 90 EHDEFITFSNGVAGDHGSWLSSRWQPPKQGVVKLNMDES-FWEQDRCMGAGGLIRDDQGI 148
EH+E D+G +W+ P+ G VKLN+D S E+ G GG+IRD+ G
Sbjct: 283 EHEETEDDEESEEADNG-----KWKKPESGWVKLNVDGSRIHEEPASAGCGGVIRDEWGT 447
Query: 149 WLGGFYAHRSGG---NALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVMADEE 202
W GF A E A+L GL++ R+ M+ V E L V +D E
Sbjct: 448 WCVGFDQKLDPNICRQAHYTELQAILTGLKVA-----REDMINV---EKLVVESDSE 594
>TC213599 homologue to UP|EXBB_PASHA (P72202) Biopolymer transport exbB
protein, partial (8%)
Length = 825
Score = 42.7 bits (99), Expect(2) = 6e-05
Identities = 22/64 (34%), Positives = 29/64 (44%)
Frame = +3
Query: 127 ESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALLLGLELVWDLGYRQVM 186
ES + GGL+RD G W F +A +AE LGL L WD G + +
Sbjct: 324 ESMHSDNDMAACGGLLRDCHGRWAASFV*KLGNTSAFVAEL*GAYLGLSLAWDKGDKNLE 503
Query: 187 VEVD 190
V +D
Sbjct: 504 VNID 515
Score = 20.8 bits (42), Expect(2) = 6e-05
Identities = 7/13 (53%), Positives = 11/13 (83%)
Frame = +2
Query: 112 RWQPPKQGVVKLN 124
R +PP +GV++LN
Sbjct: 281 R*KPPDEGVIRLN 319
>AW620606
Length = 420
Score = 33.9 bits (76), Expect(2) = 1e-04
Identities = 14/29 (48%), Positives = 21/29 (72%)
Frame = +1
Query: 4 CSAPEEDILHCLRDCPHSQEIWLRLGALA 32
CS ++DI+HCL DCP ++E+ LG L+
Sbjct: 91 CS*NDQDIIHCLWDCPKAREV*SCLGFLS 177
Score = 28.9 bits (63), Expect(2) = 1e-04
Identities = 10/32 (31%), Positives = 16/32 (49%)
Frame = +2
Query: 64 VWKWRNNMVFEDSPWLVDEACRRICHEHDEFI 95
+WKWRNN F W+ + I H++ +
Sbjct: 269 IWKWRNNKFFYVDKWVTQQVICWIYALHNDIV 364
>AW277722 similar to SP|O65719|HS73_ Heat shock cognate 70 kDa protein 3
(Hsc70.3). [Mouse-ear cress] {Arabidopsis thaliana},
partial (8%)
Length = 432
Score = 42.7 bits (99), Expect = 2e-04
Identities = 24/58 (41%), Positives = 32/58 (54%), Gaps = 4/58 (6%)
Frame = +1
Query: 113 WQPPKQGVVKLNMDESFWEQDR--CMGA--GGLIRDDQGIWLGGFYAHRSGGNALIAE 166
W+ P G V LN D S +E R C G+ GGL+RD G +LGGF + + +AE
Sbjct: 247 WRCPPNGWVCLNTDGSVFENHRNGCSGSACGGLVRDSSGCYLGGFTVNLGNTSVTLAE 420
>TC206833 homologue to UP|Q6PDS7 (Q6PDS7) Hist1h4h protein (Fragment),
complete
Length = 1119
Score = 40.0 bits (92), Expect = 0.001
Identities = 25/90 (27%), Positives = 38/90 (41%), Gaps = 14/90 (15%)
Frame = -3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTN--------------FGVGDHGSWIR 46
CP C EE H C +W +L WTN + +G +R
Sbjct: 466 CPLCRNMEESACHLFFQCSRILPVWWE--SLNWTNIVGAFPLNSKQHFLYHIGGLEGGVR 293
Query: 47 SQARSRNSVRFLAGVWGVWKWRNNMVFEDS 76
+ +R V +LA W +WK RNN++F ++
Sbjct: 292 A---NRWKVWWLALTWTIWKHRNNVIFSNA 212
>AW832520
Length = 423
Score = 40.0 bits (92), Expect = 0.001
Identities = 27/89 (30%), Positives = 39/89 (43%)
Frame = -3
Query: 81 DEACRRICHEHDEFITFSNGVAGDHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGG 140
D A I +E ++ NG+ RW PP +G+ KLN D + + AGG
Sbjct: 316 DGALMVILRLMEEITSYPNGI----------RWSPPPEGLFKLNYDGAVNCNSLKVSAGG 167
Query: 141 LIRDDQGIWLGGFYAHRSGGNALIAEATA 169
++RD G ++ GF IAE A
Sbjct: 166 VVRDSHGNFVIGFTTTPDSCTINIAELWA 80
>BI427114
Length = 425
Score = 39.7 bits (91), Expect = 0.002
Identities = 24/79 (30%), Positives = 33/79 (41%), Gaps = 6/79 (7%)
Frame = +3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLR-LGALAWTNFGVGDHGSWIRSQARSRNSVRFLA 59
C RC EE LH LRDCP E W + F GD W+ + F
Sbjct: 15 CARCDFLEETCLHALRDCPKVAEFWRSVIPKKLAPKFFNGDVTVWLERNLSFSEAAFFWP 194
Query: 60 GVWGV-----WKWRNNMVF 73
+G+ W++RN ++F
Sbjct: 195TFFGIAVELLWEFRNAVMF 251
>TC231841
Length = 791
Score = 39.7 bits (91), Expect = 0.002
Identities = 30/98 (30%), Positives = 43/98 (43%), Gaps = 15/98 (15%)
Frame = +3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNF-GVG---------DHGSWIRSQAR 50
CP C + EE H C +W +L+W N GV H + + R
Sbjct: 291 CPFCRSAEESAAHLFFHCSRIAPVWWE--SLSWVNLLGVFPNHPRQHFLQHIYGVTAGMR 464
Query: 51 -SRNSVRFLAGVWGVWKWRNNMVFEDSPW----LVDEA 83
SR +LA W +WK RNNM+F + + ++DEA
Sbjct: 465 ASRWKWWWLALTWTIWKQRNNMIFSNGTFNANKILDEA 578
>TC211809
Length = 895
Score = 39.7 bits (91), Expect = 0.002
Identities = 24/86 (27%), Positives = 38/86 (43%), Gaps = 11/86 (12%)
Frame = +2
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTN-FGVGDHG---SWIRSQARSRNSVR 56
CP C++ EED H C ++ +W +W N G H +++ RS V
Sbjct: 398 CPLCNSSEEDAAHLFFHCTKTEXLWWE--TQSWVNSLGAFPHNPKDHFLQHDNRSAGGVX 571
Query: 57 -------FLAGVWGVWKWRNNMVFED 75
++A W +WK RN +VF +
Sbjct: 572 ARRWKCLWVALTWVIWKHRNGVVFHN 649
>AW423846
Length = 385
Score = 38.1 bits (87), Expect = 0.004
Identities = 22/86 (25%), Positives = 35/86 (40%), Gaps = 11/86 (12%)
Frame = +3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTN----FGVGDHGSWIRSQARSRNSVR 56
CP C EED H C +W +L+WTN F H +++ R+
Sbjct: 114 CPFCKNKEEDAAHLFFTCSKIMPLWW--ASLSWTNTTAPFPENPHMHFLQQVLRNGKDTS 287
Query: 57 F-------LAGVWGVWKWRNNMVFED 75
+ + W +W+ RN +V E+
Sbjct: 288 YQKWQCWWTSLTWSIWQHRNKIVXEE 365
>BU547962
Length = 591
Score = 37.7 bits (86), Expect = 0.006
Identities = 20/66 (30%), Positives = 34/66 (51%)
Frame = -2
Query: 140 GLIRDDQGIWLGGFYAHRSGGNALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVMA 199
G++R+ +L + N+ IAE TAL GLELV + G+ + +E D L++++
Sbjct: 590 GVVRNHNAEFLXXYAESIGQANSTIAELTALRKGLELVLENGWNDIWLEGDAKTLVEIIV 411
Query: 200 DEESCR 205
R
Sbjct: 410 KRRKVR 393
>TC210729 similar to UP|Q7QHL7 (Q7QHL7) AgCP7056 (Fragment), partial (5%)
Length = 727
Score = 37.4 bits (85), Expect = 0.008
Identities = 24/90 (26%), Positives = 38/90 (41%), Gaps = 14/90 (15%)
Frame = +2
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFG--------------VGDHGSWIR 46
CP C EE+ H C Q +W A++W N +G +R
Sbjct: 173 CPFCRVKEENAGHLFFHCTKIQPLWWE--AMSWINLKGAAPLTPKMNFNQHMGLQADGVR 346
Query: 47 SQARSRNSVRFLAGVWGVWKWRNNMVFEDS 76
S +R ++A W +WK RN++VF ++
Sbjct: 347 S---NRWQCWWMALTWSIWKTRNSIVFSNA 427
>TC210600 similar to UP|Q51534 (Q51534) PilW protein, partial (7%)
Length = 1093
Score = 37.4 bits (85), Expect = 0.008
Identities = 24/86 (27%), Positives = 38/86 (43%), Gaps = 11/86 (12%)
Frame = +3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTN-FGVGDHG---SWIRSQARSRNSVR 56
CP C++ EED H C + +W +W N G H +++ S + VR
Sbjct: 579 CPLCNSLEEDAAHLFFPCTKTLPLWWE--TQSWINSLGAFPHNPKDHFLQHDNSSADGVR 752
Query: 57 -------FLAGVWGVWKWRNNMVFED 75
++A W +WK RN +VF +
Sbjct: 753 ARRWKCWWVALTWVIWKHRNGVVFHN 830
>TC222557
Length = 1002
Score = 37.4 bits (85), Expect = 0.008
Identities = 22/84 (26%), Positives = 38/84 (45%), Gaps = 11/84 (13%)
Frame = +1
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTN----FGVGDHGSWIR-------SQA 49
CP C+ EE+ H +CP +W +LAWT F ++++
Sbjct: 556 CPFCNRQEEEASHLFFNCPRILPLWWE--SLAWTKTVGVFSPIPRQNYMQHTLISKGKHL 729
Query: 50 RSRNSVRFLAGVWGVWKWRNNMVF 73
++R ++A W +W+ RN +VF
Sbjct: 730 QARWKCWWVALTWSIWRHRNRIVF 801
>TC222857
Length = 657
Score = 37.0 bits (84), Expect = 0.010
Identities = 28/98 (28%), Positives = 44/98 (44%), Gaps = 15/98 (15%)
Frame = -3
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTN------FGVGDH----GSWIRSQAR 50
CP C EE H C +Q IW + W N + + DH S + R
Sbjct: 496 CPFCRRVEETASHMFIHCIKTQPIWWE--TMNWINMQGPLPWSITDHFMQFSSLKEAGIR 323
Query: 51 SRN-SVRFLAGVWGVWKWRNNMVFEDSPW----LVDEA 83
SR ++A W +W+ RN++VF ++ + LV++A
Sbjct: 322 SRRWQWWWMAVTWSIWQLRNSIVFSNATFDGNKLVEDA 209
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.333 0.143 0.509
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,810,016
Number of Sequences: 63676
Number of extensions: 305078
Number of successful extensions: 1937
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 1906
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1924
length of query: 278
length of database: 12,639,632
effective HSP length: 96
effective length of query: 182
effective length of database: 6,526,736
effective search space: 1187865952
effective search space used: 1187865952
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0118c.6


