

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
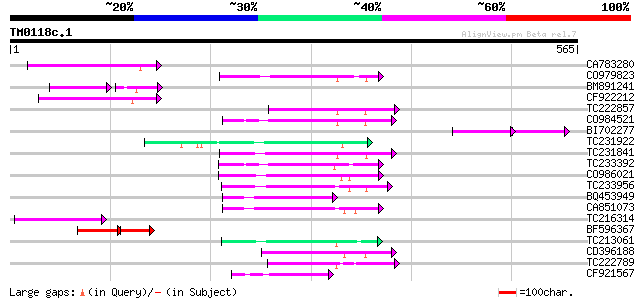
Query= TM0118c.1
(565 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 84 2e-16
CO979823 70 3e-12
BM891241 59 1e-11
CF922212 65 8e-11
TC222857 63 3e-10
CO984521 63 4e-10
BI702277 49 5e-10
TC231922 62 7e-10
TC231841 62 7e-10
TC233392 62 9e-10
CO986021 62 9e-10
TC233956 61 1e-09
BQ453949 61 2e-09
CA851073 60 2e-09
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 59 6e-09
BF596367 42 6e-09
TC213061 57 2e-08
CD396188 weakly similar to GP|6175788|gb|A ORF144 {Xestia c-nigr... 57 2e-08
TC222789 57 3e-08
CF921567 55 1e-07
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 84.0 bits (206), Expect = 2e-16
Identities = 44/139 (31%), Positives = 69/139 (48%), Gaps = 5/139 (3%)
Frame = +2
Query: 18 SCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDFKAFNTAL 77
S F L + +++E + RF WGG+ R + W+ W T+C PK GG+G +D + FNT L
Sbjct: 5 SFFSLP*KIIAKLESLQRRFLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTTL 184
Query: 78 LAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAWI-----FQE 132
L K W + + AK+ ++ Y + E G + S W L K+ + +
Sbjct: 185 LGKWRWDLFYIQQEPWAKVLQSKYGGWRALEEGSSGSKDSAWWKDLIKTQQLQRNIPLKR 364
Query: 133 GGLWRVGNGKNIRA*EDKW 151
+W+VG G I+ ED W
Sbjct: 365 ETIWKVGGGDRIKFWEDLW 421
>CO979823
Length = 853
Score = 70.1 bits (170), Expect = 3e-12
Identities = 50/175 (28%), Positives = 83/175 (46%), Gaps = 12/175 (6%)
Frame = -2
Query: 210 QRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQCC 269
QR+D + W G YST++GY + + +E + A + WK +
Sbjct: 711 QREDKWLWKPEPGGHYSTKSGYHVLWGELTEEIQDADFAEI---------WKLKIPTKAA 559
Query: 270 ETAWRACLDILPVRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPLVRMIWFASP--L 326
AWR D LP +S L RR V V + CP C+ +EE H F+C +W+ S +
Sbjct: 558 VFAWRLVRDRLPTKSNLRRRQVMVQDMVCPLCNNIEEGAAHLFFNCTKTLPLWWESMSWV 379
Query: 327 NLRFDEEGSVHDFFLSFLATADTNIA-GI--------FLAVLYSVWQARNELLFK 372
NL+ + FL + T+IA G+ ++A+ +++WQ RN+++F+
Sbjct: 378 NLKTAMPQTPRQHFLQY----GTDIADGLKSKRWKCWWIALTWTIWQHRNKVVFQ 226
>BM891241
Length = 407
Score = 58.9 bits (141), Expect(2) = 1e-11
Identities = 24/62 (38%), Positives = 37/62 (58%)
Frame = -2
Query: 40 GGDVSHRTMHWLKWDTLCRPKGCGGIGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKA 99
GGD H+ + W+KW+ +C PK GG+G +D FN ALL K W + + + + A+I +
Sbjct: 406 GGDHDHKKIPWVKWEVVCLPKTKGGLGIKDISKFNIALLGKWIWALASDQQQLWARIINS 227
Query: 100 VY 101
Y
Sbjct: 226 KY 221
Score = 28.9 bits (63), Expect(2) = 1e-11
Identities = 19/52 (36%), Positives = 25/52 (47%), Gaps = 5/52 (9%)
Frame = -3
Query: 106 SFLEAKKGYRPSYVWLSLQK-----SAWIFQEGGLWRVGNGKNIRA*EDKWV 152
S + +K GY SY W L+K IF + W+VG G I DKW+
Sbjct: 201 SIIVSKGGY--SYWWRDLRKFYHQSDHSIFHQYMSWKVGCGDKINFWTDKWL 52
>CF922212
Length = 445
Score = 65.1 bits (157), Expect = 8e-11
Identities = 35/130 (26%), Positives = 60/130 (45%), Gaps = 7/130 (5%)
Frame = -2
Query: 29 QIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDFKAFNTALLAKNWWRIMTK 88
++ R+ F WGG + + + W+ WD++C PK G+G RD + FN ALL K W +
Sbjct: 420 KLVRLQQWFLWGGGLGQKKIAWVNWDSVCLPKEKEGLGNRDLRKFNYALLGKRRWNLFHH 241
Query: 89 LKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWL-------SLQKSAWIFQEGGLWRVGNG 141
+ A++ + Y + E ++ S+ W S + +W + R+G
Sbjct: 240 QGELGARVLDSKYKRWRNLDEERRVKSESFWWQEISFITHSTEDGSWFEKRTKTGRLGCR 61
Query: 142 KNIRA*EDKW 151
+R ED W
Sbjct: 60 AKVRFWEDGW 31
>TC222857
Length = 657
Score = 63.2 bits (152), Expect = 3e-10
Identities = 41/138 (29%), Positives = 66/138 (47%), Gaps = 8/138 (5%)
Frame = -3
Query: 259 LWKADAVPQCCETAWRACLDILPVRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPLV 317
LWK + AWR D LP ++ L R V + + CP C VEET H C
Sbjct: 613 LWKIKIPSKFLMFAWRLLWDRLPTKANLRARQVQISDLTCPFCRRVEETASHMFIHCIKT 434
Query: 318 RMIWFASP--LNLRFDEEGSVHDFFLSFLATADTNIAG-----IFLAVLYSVWQARNELL 370
+ IW+ + +N++ S+ D F+ F + + I ++AV +S+WQ RN ++
Sbjct: 433 QPIWWETMNWINMQGPLPWSITDHFMQFSSLKEAGIRSRRWQWWWMAVTWSIWQLRNSIV 254
Query: 371 FKWKDSSIDQLLQRASSL 388
F ++L++ AS L
Sbjct: 253 FSNATFDGNKLVEDASFL 200
>CO984521
Length = 716
Score = 62.8 bits (151), Expect = 4e-10
Identities = 47/181 (25%), Positives = 75/181 (40%), Gaps = 8/181 (4%)
Frame = -2
Query: 213 DCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQCCETA 272
D + W GCYST++ Y + + GE + +LWK + A
Sbjct: 646 DQWKWAAEPSGCYSTKSAYKAL-HHVTVGEEQDGKFK--------ELWKLRVPLKVAIFA 494
Query: 273 WRACLDILPVRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPLVRMIWFASP--LNLR 329
WR D LP ++ L ++ V + CP C VEET H F C V +W+ S +N+
Sbjct: 493 WRLIQDKLPTKANLRKKRVELQEYLCPLCRSVEETASHLFFHCSKVSPLWWESQSWVNMM 314
Query: 330 FDEEGSVHDFFLSFLATADTNIAG-----IFLAVLYSVWQARNELLFKWKDSSIDQLLQR 384
F + A + G + A+ YS+W+ RN ++F + +L+
Sbjct: 313 GVFPYQPDQHFSQHIFGASVGLQGKRWQWWWFALTYSIWKHRNSIIFSNANFDAHKLMDD 134
Query: 385 A 385
A
Sbjct: 133 A 131
>BI702277
Length = 420
Score = 49.3 bits (116), Expect(2) = 5e-10
Identities = 22/63 (34%), Positives = 36/63 (56%)
Frame = +2
Query: 442 GEVLAAATTELGPVLSPVLAEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEF 501
GEVLA+AT SP +AE L +W++++A +L + ++ ETDC E+ + W
Sbjct: 2 GEVLASATRLEDQCYSPQIAEYLAFKWSIQMANQLLLQNVFFETDCREIASAWRNASQRI 181
Query: 502 SHL 504
H+
Sbjct: 182 VHI 190
Score = 33.1 bits (74), Expect(2) = 5e-10
Identities = 16/57 (28%), Positives = 30/57 (52%)
Frame = +1
Query: 502 SHLATVVFDCRSLLPNFDVFNFTFVRRTSNGAADALARLAFHYGCMIWIEEAPSEIT 558
++L ++ DCR + + D ++ N AD+LA LAF +E+AP +++
Sbjct: 184 TYLDAILQDCRDMTTHIDTVALVHTKKIGNIVADSLA*LAFVLVDQSGVEDAPPQLS 354
>TC231922
Length = 803
Score = 62.0 bits (149), Expect = 7e-10
Identities = 57/245 (23%), Positives = 92/245 (37%), Gaps = 18/245 (7%)
Frame = -3
Query: 135 LWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLG-----VVHVSDLMVTNGGCWD---R 186
+WRVG G I+ +D W+ G L + + ++ + N W+ R
Sbjct: 708 VWRVGCGDKIKFWQDSWLSEGCNLQQKYNQLYTISRQ*NLTISKMGKFSQNARSWEFKWR 529
Query: 187 NKV---EFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEA 243
++ E+ I I + Q D FW G YST++ Y + S +
Sbjct: 528 RRLFDYEYAMAVDFMNEISGISIQ-NQAHDSMFWKADSSGVYSTKSAYRLLMPSISPAPS 352
Query: 244 SSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGV-VGNPCCPRCSE 302
++ LW P+ +WR LD LP R L RR + + + CP C
Sbjct: 351 R---------RIFQILWHLKIPPRAAVFSWRLFLDRLPTRGNLSRRSIPIQDIMCPLCGC 199
Query: 303 VEETVHHALFSCPLVRMIWFASPLNLRF------DEEGSVHDFFLSFLATADTNIAGIFL 356
E H F C + + +W+ S +R D F F AT++ + G++
Sbjct: 198 QHEEAGHLFFHCKMTKGLWWESMRWIRVIGALPADPASHFIQFCDGFAATSNHSRRGMWW 19
Query: 357 AVLYS 361
L S
Sbjct: 18 IALSS 4
>TC231841
Length = 791
Score = 62.0 bits (149), Expect = 7e-10
Identities = 44/184 (23%), Positives = 78/184 (41%), Gaps = 8/184 (4%)
Frame = +3
Query: 210 QRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQCC 269
Q+ D + W G Y+ ++ Y + + E + ++ +LWK +
Sbjct: 54 QQGDQWVWKADPSGQYTAKSAYGVLWGEMFEEQQDG---------VFEELWKLKLPSKIT 206
Query: 270 ETAWRACLDILPVRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPLVRMIWFASP--L 326
AWR D LP RS L R+ + V +P CP C EE+ H F C + +W+ S +
Sbjct: 207 IFAWRLIRDRLPTRSNLRRKQIEVDDPRCPFCRSAEESAAHLFFHCSRIAPVWWESLSWV 386
Query: 327 NLRFDEEGSVHDFFLSFLATADTNIAGI-----FLAVLYSVWQARNELLFKWKDSSIDQL 381
NL FL + + +LA+ +++W+ RN ++F + +++
Sbjct: 387 NLLGVFPNHPRQHFLQHIYGVTAGMRASRWKWWWLALTWTIWKQRNNMIFSNGTFNANKI 566
Query: 382 LQRA 385
L A
Sbjct: 567 LDEA 578
>TC233392
Length = 1145
Score = 61.6 bits (148), Expect = 9e-10
Identities = 49/175 (28%), Positives = 75/175 (42%), Gaps = 11/175 (6%)
Frame = -1
Query: 209 QQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQC 268
QQ D + W + YSTR+ Y + + EGE + + LWK +
Sbjct: 668 QQVPDTWVWKHEPNELYSTRSAYKLL-QGHLEGEDQDGALQ--------DLWKLKIPAKA 516
Query: 269 CETAWRACLDILPVRSILHRRGVV-GNPCCPRCSEVEETVHHALFSCPLVRMIWF----- 322
AWR D LP +S LHRR VV + CP C EE H C +R +W+
Sbjct: 515 SIFAWRLIRDRLPTKSNLHRRQVVLEDSLCPFCRIREEDASHIFLECNKIRPLWWESQTW 336
Query: 323 -----ASPLNLRFDEEGSVHDFFLSFLATADTNIAGIFLAVLYSVWQARNELLFK 372
SP+N R VH + + ++A+ +S+W+ RN+++F+
Sbjct: 335 VRSLGVSPINKRQHFLQHVHG---TPGSKRYNRWKTWWIALTWSLWKHRNQVIFQ 180
>CO986021
Length = 900
Score = 61.6 bits (148), Expect = 9e-10
Identities = 50/172 (29%), Positives = 76/172 (44%), Gaps = 8/172 (4%)
Frame = -1
Query: 209 QQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQC 268
QQ D W G YST++ Y + G+ SS+ + LWK P+
Sbjct: 795 QQLQDSMLWKADPTGIYSTKSAYRLLLPNNRPGQHSSN---------FKILWKLKIPPRA 643
Query: 269 CETAWRACLDILPVRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPLVRMIWFASPLN 327
+WR D L +R+ L RR V + + CP C +E H F C + +W+ S
Sbjct: 642 ELFSWRLFRDRLSIRANLLRRHVALQDIMCPLCGNHQEEAGHLFFHCRMTIGLWWESMNW 463
Query: 328 LR----FDEEGSVH--DFFLSFLATADTNIAGI-FLAVLYSVWQARNELLFK 372
R F ++ + H F F A + N + ++A+ ++WQ RN LLFK
Sbjct: 462 TRTLGAFLDDPAAHFIQFSDGFGAQRNHNRRCLWWIALTSTIWQHRNSLLFK 307
>TC233956
Length = 757
Score = 61.2 bits (147), Expect = 1e-09
Identities = 46/182 (25%), Positives = 80/182 (43%), Gaps = 12/182 (6%)
Frame = +2
Query: 212 DDCFFWPETRDGCYSTRTGYSFI*KKFS-EGEASSSSARVMPPALWMKLWKADAVPQCCE 270
+D + W G ST++ Y I + EG+ + KLW+ P+
Sbjct: 56 NDTWVWRAESTGIISTKSAYQVIKSELDDEGQHLG----------FKKLWEIKVPPKALS 205
Query: 271 TAWRACLDILPVRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPLVRMIWFASPLNLR 329
WR D LP + L +R + V + CP C ET H F+C + +W+ L
Sbjct: 206 FVWRLLWDRLPTKDNLIKRQIQVEDDLCPFCHSQSETASHLFFTCGKIMPLWWEF---LS 376
Query: 330 FDEEGSVH-----DFFLSFLATADTNIAGI-----FLAVLYSVWQARNELLFKWKDSSID 379
+ +E V D FL ++A + ++ ++AV S+W+ RN+++F+ + I
Sbjct: 377 WVKEDKVFHCRPMDNFLQHYSSAASKVSNTRRTMWWIAVTNSIWRLRNDIIFQNQTVDIT 556
Query: 380 QL 381
+L
Sbjct: 557 RL 562
>BQ453949
Length = 450
Score = 60.8 bits (146), Expect = 2e-09
Identities = 32/115 (27%), Positives = 55/115 (47%), Gaps = 1/115 (0%)
Frame = +3
Query: 213 DCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQCCETA 272
D W DG YST++ Y+F+ + ++ + + + +W P+ C +
Sbjct: 30 DLLLWGSNSDGSYSTKSAYNFL---------KNEDSQTIEDSAFKNIWNLKLPPRACAFS 182
Query: 273 WRACLDILPVRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPLVRMIWFASPL 326
R + +P + L RR V + + CP C E EETV H ++SC R +W+ + L
Sbjct: 183 *RIF*NRIPTKVNLRRRHVKLPSSNCPLCDEEEETVGHVMYSCIKTRHLWWETEL 347
>CA851073
Length = 633
Score = 60.5 bits (145), Expect = 2e-09
Identities = 44/171 (25%), Positives = 72/171 (41%), Gaps = 11/171 (6%)
Frame = +1
Query: 213 DCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQCCETA 272
D W G YST++ Y + + + V ++ +WK P+ +
Sbjct: 28 DTMVWKADPCGVYSTKSAYRIL---------MTCNRHVSEANIFKTIWKLKIPPRAAVFS 180
Query: 273 WRACLDILPVRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPLVRMIWFASPLNLRFD 331
WR D LP R L RR V + CP C +E H F+C + R +W+ S +R++
Sbjct: 181 WRLIKDRLPTRHNLLRRNVSIQENECPLCGYEQEEADHLFFNCKMTRGLWWES---MRWN 351
Query: 332 E-----EGSVHDFFLSF-----LATADTNIAGIFLAVLYSVWQARNELLFK 372
+ S F+ F T G ++A+ S+W+ RN L+F+
Sbjct: 352 QIVGPLSVSPASHFVQFCDGFGAGRNHTRWCGWWIALTSSIWKHRNLLIFQ 504
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 58.9 bits (141), Expect = 6e-09
Identities = 26/92 (28%), Positives = 46/92 (49%)
Frame = +2
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I + A+P +S F + + ++ + +F WGG H + W+KW +C PK GG
Sbjct: 23 IKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADICNPKIDGG 202
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKI 96
+G +D FN AL + W + + + A++
Sbjct: 203 LGIKDLSNFNAALRGRWIWGLASNHNQLWARL 298
>BF596367
Length = 419
Score = 42.4 bits (98), Expect(2) = 6e-09
Identities = 19/45 (42%), Positives = 28/45 (62%)
Frame = +1
Query: 68 RDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKK 112
R KAF+ A+LAK WR++ +++ + A YFP S L+AKK
Sbjct: 58 RRIKAFHKAMLAKQMWRLIDNKDALVT*VLTAKYFPNSSLLDAKK 192
Score = 36.2 bits (82), Expect(2) = 6e-09
Identities = 13/34 (38%), Positives = 23/34 (67%)
Frame = +3
Query: 111 KKGYRPSYVWLSLQKSAWIFQEGGLWRVGNGKNI 144
KK Y P+Y W S+ S + + G +WR+G+G+++
Sbjct: 189 KKAYMPNYSWSSV**SKDLAEHGAIWRIGDGRSV 290
>TC213061
Length = 823
Score = 57.4 bits (137), Expect = 2e-08
Identities = 42/171 (24%), Positives = 68/171 (39%), Gaps = 11/171 (6%)
Frame = +2
Query: 212 DDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQCCET 271
+D + W G Y R+ Y I + E E + +LWK +
Sbjct: 125 EDQWVWAAESSGSYLARSAYRVIREGIPEEEQDRE---------FKELWKLKVPMKVTMF 277
Query: 272 AWRACLDILPVRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPLVRMIWFAS------ 324
AWR DILP R L R+ V + CP C ++E+ H F C + IW+ S
Sbjct: 278 AWRLLKDILPTRDNLRRKRVELHEYVCPLCRSMDESASHLFFHCSKILPIWWESLSWVKL 457
Query: 325 ----PLNLRFDEEGSVHDFFLSFLATADTNIAGIFLAVLYSVWQARNELLF 371
P + R H+ + +LA+ +++W+ RN+++F
Sbjct: 458 VGAFPHHPRHHFHQHSHEVYQGLQGN---RWKWWWLALTWTIWKHRNDIIF 601
>CD396188 weakly similar to GP|6175788|gb|A ORF144 {Xestia c-nigrum
granulovirus}, partial (5%)
Length = 701
Score = 57.4 bits (137), Expect = 2e-08
Identities = 39/142 (27%), Positives = 63/142 (43%), Gaps = 8/142 (5%)
Frame = -1
Query: 252 PPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVV-GNPCCPRCSEVEETVHHA 310
P + + +LW+ P AWR D LP + L RR +V N CP C E+ H
Sbjct: 647 PNSGFRQLWEIKIPPTALSFAWRLLWDRLPSKENLIRRQIVLQNDLCPFCQSQVESASHL 468
Query: 311 LFSCPLVRMIWFASPLNLRFDE--EGSVHDFFLSFLATADTNIAG-----IFLAVLYSVW 363
F+C V +W+ +R D D FL + A + + ++A S+W
Sbjct: 467 FFTCHKVMPLWWEFNTWVREDRVLHSKPMDNFLQHCSLAGSRNSNRRRKIWWIAATRSIW 288
Query: 364 QARNELLFKWKDSSIDQLLQRA 385
RN+++F + I +L+ +A
Sbjct: 287 NLRNDMIFNNQPFDISKLVDKA 222
>TC222789
Length = 954
Score = 56.6 bits (135), Expect = 3e-08
Identities = 43/142 (30%), Positives = 64/142 (44%), Gaps = 11/142 (7%)
Frame = +2
Query: 258 KLWKADAVPQCCETAWRACLDILPVRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPL 316
KLWK + AWR D LP + LHRR + V + CP C VEE H F C
Sbjct: 359 KLWKLKVPIKYEVFAWRLLRDRLPTKVNLHRRQIQVMDRSCPFCRNVEEDAGHLFFHCSK 538
Query: 317 VRMIWFAS----------PLNLRFDEEGSVHDFFLSFLATADTNIAGIFLAVLYSVWQAR 366
+ IW+ S P + R + HD ++ T +LAV +++WQ R
Sbjct: 539 IIPIWWESLSWVNISGALPKDPR--QHFLQHDLIMAG-GIRTTRWKCWWLAVTWTIWQQR 709
Query: 367 NELLFKWKDSSIDQLLQRASSL 388
N+++F ++L+ A+ L
Sbjct: 710 NKIIFFNDSFDANKLIDEAAFL 775
>CF921567
Length = 686
Score = 54.7 bits (130), Expect = 1e-07
Identities = 34/102 (33%), Positives = 50/102 (48%), Gaps = 1/102 (0%)
Frame = -1
Query: 222 DGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILP 281
+G YST++ Y +* EG AS+ RV+ +W P+ +WR + LP
Sbjct: 677 NGLYSTKSAYKVL*----EGHASAIEDRVLNI-----MWSLKIPPRASAFSWRLFKNRLP 525
Query: 282 VRSILHRRGV-VGNPCCPRCSEVEETVHHALFSCPLVRMIWF 322
R L RR V + CP C EE+V+H F+C R +W+
Sbjct: 524 TRENLKRRQVSLHTYSCPLCDLEEESVNHLFFNCSKTRSLWW 399
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.139 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,217,878
Number of Sequences: 63676
Number of extensions: 602409
Number of successful extensions: 3650
Number of sequences better than 10.0: 110
Number of HSP's better than 10.0 without gapping: 3569
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3607
length of query: 565
length of database: 12,639,632
effective HSP length: 102
effective length of query: 463
effective length of database: 6,144,680
effective search space: 2844986840
effective search space used: 2844986840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0118c.1


