

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
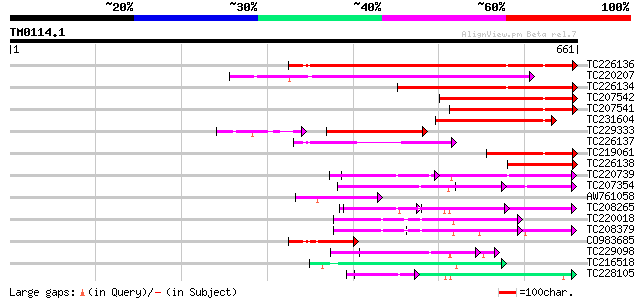
Query= TM0114.1
(661 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC226136 similar to UP|O48847 (O48847) Expressed protein (At2g32... 362 e-100
TC220207 similar to UP|Q6H975 (Q6H975) STY-L protein, partial (43%) 282 3e-76
TC226134 similar to UP|Q6H975 (Q6H975) STY-L protein, partial (20%) 223 2e-58
TC207542 similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial ... 188 6e-48
TC207541 similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial ... 154 1e-37
TC231604 weakly similar to UP|Q6H976 (Q6H976) STYLOSA protein, p... 142 6e-34
TC229333 similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial ... 128 2e-31
TC226137 similar to UP|O48847 (O48847) Expressed protein (At2g32... 114 1e-25
TC219061 similar to UP|Q6H975 (Q6H975) STY-L protein, partial (10%) 112 7e-25
TC226138 similar to UP|O48847 (O48847) Expressed protein (At2g32... 101 9e-22
TC220739 weakly similar to UP|PRP4_HUMAN (O43172) U4/U6 small nu... 90 3e-18
TC207354 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat prot... 85 1e-16
AW761058 76 5e-14
TC208265 similar to UP|Q6S7B0 (Q6S7B0) TAF5, partial (36%) 74 3e-13
TC220018 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana geno... 69 7e-12
TC208379 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana geno... 68 1e-11
CO983685 65 7e-11
TC229098 similar to PIR|T46032|T46032 WD-40 repeat regulatory pr... 65 1e-10
TC216518 weakly similar to UP|Q9LW87 (Q9LW87) Coatomer protein c... 59 5e-09
TC228105 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {A... 58 1e-08
>TC226136 similar to UP|O48847 (O48847) Expressed protein
(At2g32700/F24L7.16), partial (42%)
Length = 1463
Score = 362 bits (928), Expect = e-100
Identities = 176/336 (52%), Positives = 241/336 (71%)
Frame = +3
Query: 326 DAKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKV 385
D ++NVESFLS +D D R + F LKR + + + +KGFS E+G + KS +KV
Sbjct: 72 DVGSLEDNVESFLSQDD--GDGR-DLFGTLKRNPSEHATDASKGFSFSEVGSIRKSNSKV 242
Query: 386 LTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFD 445
+ CHFSSDGK++ASAGH+KKV +WNME Q ++ + HS +ITDVRF+P ST ATSSFD
Sbjct: 243 VCCHFSSDGKLLASAGHDKKVVLWNMETLQTESTPEEHSLIITDVRFRPNSTQLATSSFD 422
Query: 446 RSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHT 505
+VRLWDA++P L +GH V+SLDFHP++ +L CS D+N+ IR W+++Q
Sbjct: 423 TTVRLWDAADPTFPLHTYSGHTSHVVSLDFHPKKTELFCSCDNNNEIRFWSISQYSSTRV 602
Query: 506 TRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLA 565
+GGS QVRFQP+ G LLA A GN +++ DVETD +H L+GH +V +CWD +G++LA
Sbjct: 603 FKGGSTQVRFQPRLGHLLAAASGNVVSLFDVETDRQMHTLQGHSAEVHCVCWDTNGDYLA 782
Query: 566 SVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
SVS++S ++WS AS G+CI EL+S GN F SC+FHP Y +LL+IGGYQSLE W+ E +K
Sbjct: 783 SVSQESVKVWSLAS-GECIHELNSSGNMFHSCVFHPSYSTLLVIGGYQSLELWNMAE-NK 956
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+I AH+ +I+ LA SP+ ++ASASHD SVK+WK
Sbjct: 957 CMTIPAHECVISALAQSPLTGMVASASHDKSVKIWK 1064
>TC220207 similar to UP|Q6H975 (Q6H975) STY-L protein, partial (43%)
Length = 1244
Score = 282 bits (722), Expect = 3e-76
Identities = 147/362 (40%), Positives = 208/362 (56%), Gaps = 7/362 (1%)
Frame = +3
Query: 257 TKRKNKHPKIPKTGSSKNDSQKMEQIISTGEHQCNQDQQIQMQSQNVEKSTKRMITESTR 316
+ RK K TG+ + ST H Q+V+KS TE+T
Sbjct: 30 SNRKRKSGAANSTGTGNTAAPSPTSSPST--HTPGDGLNTASSMQHVQKSMMMYGTEATG 203
Query: 317 PVESAQDC-------ADAKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKG 369
+ S+ + D D+NVESFLS + N + +K+ A + +KG
Sbjct: 204 GLASSSNLLDDMDRFGDVDALDDNVESFLSNDGGDGGNL---YGTVKQSPAEQQKESSKG 374
Query: 370 FSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITD 429
F+ E+GC +KV CHFSSDGK +ASAG + KV IWNM+ + ++ H +ITD
Sbjct: 375 FTFAEVGCRRTRNSKVTCCHFSSDGKWLASAGDDMKVDIWNMDTLETESTPAEHKSVITD 554
Query: 430 VRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSN 489
VRF+P S+ AT+S D+SVRLWD +NP R L +GH +MSLDFHP++ +L C D
Sbjct: 555 VRFRPNSSQLATASTDKSVRLWDTTNPSRCLQEYSGHSSAIMSLDFHPKKTELFCFCDGE 734
Query: 490 DVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHD 549
+ IR WN+N + C T+G S QVRFQP+ G+ LA A ++I DVE+D+ ++ L+GH
Sbjct: 735 NEIRYWNINSATCTRVTKGASAQVRFQPRLGRFLAAASDKGVSIFDVESDTQIYTLQGHP 914
Query: 550 KDVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLII 609
+ V ICWD +G+ LASVS + ++WS S G+CI E S G++ SC+FHP Y +LL+I
Sbjct: 915 EPVSYICWDGNGDALASVSPNLVKVWSLTSGGECIHEFSSTGSQLHSCVFHPSYSTLLVI 1094
Query: 610 GG 611
GG
Sbjct: 1095GG 1100
>TC226134 similar to UP|Q6H975 (Q6H975) STY-L protein, partial (20%)
Length = 826
Score = 223 bits (569), Expect = 2e-58
Identities = 104/209 (49%), Positives = 144/209 (68%)
Frame = +2
Query: 453 ASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQ 512
A++P + H V+SLDF P++ +L CS D+N+ IR W+++Q +GGS Q
Sbjct: 8 AADPXXXXHTYSXHTSHVVSLDFXPKKTELFCSCDNNNEIRFWSISQYSSTRIFKGGSTQ 187
Query: 513 VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSA 572
VRFQP+ G LLA A G+ +++ DVETD +H L+GH +V +CWD +G++LASVS++S
Sbjct: 188 VRFQPRLGHLLAAASGSVVSLFDVETDRQIHTLQGHSAEVRCVCWDTNGDYLASVSQESV 367
Query: 573 RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAH 632
++WS AS G+CI EL S GN F SC+FHP Y +LL+IGGYQSLE W+ E AAH
Sbjct: 368 KVWSLAS-GECIHELTSSGNMFHSCVFHPSYSTLLVIGGYQSLELWNMAENKCMTVSAAH 544
Query: 633 QGLIAGLADSPVGELIASASHDNSVKVWK 661
+ +I+ LA SPV ++ASASHD SVK+WK
Sbjct: 545 ECVISALAHSPVTRMVASASHDKSVKIWK 631
>TC207542 similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial (18%)
Length = 670
Score = 188 bits (478), Expect = 6e-48
Identities = 90/162 (55%), Positives = 120/162 (73%), Gaps = 2/162 (1%)
Frame = +1
Query: 502 CMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG 561
C ++GG+ Q+RFQP+ G+ LA A N ++I+DVET + + LKGH K + S+CWD SG
Sbjct: 7 CARVSKGGAVQMRFQPRLGRYLAAAAENVVSILDVETQASRYSLKGHTKSIHSVCWDPSG 186
Query: 562 NFLASVSEDSARIWS--AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
FLASVSEDS R+W+ + S+G+C+ EL GNKF SC+FHP Y SLL++G YQSLE W+
Sbjct: 187 EFLASVSEDSVRVWTLGSGSEGECVHELSCNGNKFHSCVFHPTYSSLLVVGCYQSLELWN 366
Query: 620 PTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
TE +KT +++AH+GLIA LA S V L+ASASHD VK+WK
Sbjct: 367 MTE-NKTMTLSAHEGLIAALAVSTVNGLVASASHDKFVKLWK 489
>TC207541 similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial (16%)
Length = 682
Score = 154 bits (389), Expect = 1e-37
Identities = 79/151 (52%), Positives = 103/151 (67%), Gaps = 2/151 (1%)
Frame = +1
Query: 513 VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSA 572
+RFQ LA A N ++I DVET + + LKGH K V +CWD SG LASVSEDS
Sbjct: 7 MRFQXXXXXYLAAAAENIVSIFDVETQACRYSLKGHTKPVDCVCWDPSGELLASVSEDSV 186
Query: 573 RIWS--AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIA 630
R+W+ + S+G+C+ EL GNKF + +FHP Y SLL+IG YQSLE W+ +E +KT +++
Sbjct: 187 RVWTLGSGSEGECVHELSCNGNKFHASVFHPTYPSLLVIGCYQSLELWNMSE-NKTMTLS 363
Query: 631 AHQGLIAGLADSPVGELIASASHDNSVKVWK 661
AH GLI LA S V L+ASASHD +K+WK
Sbjct: 364 AHDGLITSLAVSTVNGLVASASHDKFLKLWK 456
>TC231604 weakly similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial (10%)
Length = 424
Score = 142 bits (357), Expect = 6e-34
Identities = 65/141 (46%), Positives = 94/141 (66%)
Frame = +1
Query: 497 VNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSIC 556
+N S C T G S QVR QP+ G+ LA A ++I DVE+D+ ++ L+GH + V IC
Sbjct: 1 INSSTCTRVTNGVSAQVRSQPRLGRFLAAASDKGVSIFDVESDTQIYTLQGHPEPVSYIC 180
Query: 557 WDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLE 616
WD +G+ LASVS + ++WS S G+CI E S GN+F SC+FHP Y +LL++GG SLE
Sbjct: 181 WDGNGDALASVSSNLVKVWSLTSGGECIHEFSSPGNQFHSCVFHPSYSTLLVVGGISSLE 360
Query: 617 AWSPTEGSKTWSIAAHQGLIA 637
W+ TE +K+ +I H+ +I+
Sbjct: 361 LWNMTE-NKSMTITTHENVIS 420
>TC229333 similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial (32%)
Length = 982
Score = 128 bits (322), Expect(2) = 2e-31
Identities = 58/118 (49%), Positives = 79/118 (66%)
Frame = +3
Query: 370 FSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITD 429
F+ ++ + S +KV CHFSSDGK++AS GH+K+V +W ++ + A+ + HS LITD
Sbjct: 624 FTFSDVNSVRASTSKVACCHFSSDGKLLASGGHDKRVVLWYTDSLKQKATLEEHSSLITD 803
Query: 430 VRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSD 487
VRF P ATSSFD++V +WD NP SL TGH VMSL+FHP + DL+CS D
Sbjct: 804 VRFSPSMPRLATSSFDKTVXVWDVDNPGYSLHTFTGHSTSVMSLNFHPNKDDLICSCD 977
Score = 26.2 bits (56), Expect(2) = 2e-31
Identities = 25/110 (22%), Positives = 46/110 (41%), Gaps = 5/110 (4%)
Frame = +1
Query: 242 APACTPNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQ----IISTGEHQCNQDQQIQ 297
+P+ P+ ++ PG P +P +GSS +++ +Q D+ ++
Sbjct: 334 SPSSAPSTPSTHTPGDVISM--PALPHSGSSSKPLMMFSTDGTGTLTSPSNQLWDDKDLE 507
Query: 298 MQSQNVEKSTKRMITESTRPVESAQDCADAKPADENVESFLSFED-EPAD 346
+Q+ R + + + DENVESFLS +D +P D
Sbjct: 508 LQAD-----VDRFVEDGS--------------LDENVESFLSHDDTDPRD 600
>TC226137 similar to UP|O48847 (O48847) Expressed protein
(At2g32700/F24L7.16), partial (15%)
Length = 417
Score = 114 bits (285), Expect = 1e-25
Identities = 68/190 (35%), Positives = 96/190 (49%)
Frame = +3
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
++NVESFLS +D D R + F LKR + + + +KGFS E+G + KS +KV+ CHF
Sbjct: 6 EDNVESFLSQDD--GDGR-DLFGTLKRNPSEHATDASKGFSFDEVGSIRKSNSKVVCCHF 176
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
SSDGK+++SAGH+KK
Sbjct: 177 SSDGKLLSSAGHDKK--------------------------------------------- 221
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
P L +GH V+SLDFHP++ +L CS D+N+ IR W+++Q +GGS
Sbjct: 222 -----PTFPLHTYSGHTSHVVSLDFHPKKTELFCSCDNNNEIRFWSISQYSSTRIFKGGS 386
Query: 511 KQVRFQPQYG 520
QVRFQP+ G
Sbjct: 387 TQVRFQPRLG 416
>TC219061 similar to UP|Q6H975 (Q6H975) STY-L protein, partial (10%)
Length = 671
Score = 112 bits (279), Expect = 7e-25
Identities = 51/105 (48%), Positives = 75/105 (70%)
Frame = +3
Query: 557 WDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLE 616
WD +G+ LASVS + ++WS S G+CI E S G++ SC+FHP Y +LL+IGG SLE
Sbjct: 3 WDGNGDALASVSPNLVKVWSLTSGGECIHEFSSTGSQLHSCVFHPSYSTLLVIGGSSSLE 182
Query: 617 AWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
W+ T+ +K+ ++ AH+ +I+ LA S V ++ASAS+DN VK+WK
Sbjct: 183 LWNMTD-NKSLTVPAHENVISALAQSSVTGMVASASYDNYVKLWK 314
Score = 32.3 bits (72), Expect = 0.69
Identities = 24/96 (25%), Positives = 48/96 (50%), Gaps = 4/96 (4%)
Frame = +3
Query: 360 ATCSRNENKGFSLQEIG-CLHK---SINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQ 415
A+ S N K +SL G C+H+ + +++ +C F + G + +WNM + +
Sbjct: 27 ASVSPNLVKVWSLTSGGECIHEFSSTGSQLHSCVFHPSYSTLLVIGGSSSLELWNMTDNK 206
Query: 416 YFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLW 451
+ H ++I+ + + M A++S+D V+LW
Sbjct: 207 SL-TVPAHENVISALAQSSVTGMVASASYDNYVKLW 311
>TC226138 similar to UP|O48847 (O48847) Expressed protein
(At2g32700/F24L7.16), partial (10%)
Length = 940
Score = 101 bits (252), Expect = 9e-22
Identities = 47/81 (58%), Positives = 58/81 (71%)
Frame = -3
Query: 581 GKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLA 640
G+CI EL S GN F SC+FHP Y +LL+IGGYQSLE W+ E AAH+ +I+ LA
Sbjct: 386 GECIHELTSSGNMFHSCVFHPSYSTLLVIGGYQSLELWNMAENKCMTVSAAHECVISALA 207
Query: 641 DSPVGELIASASHDNSVKVWK 661
SPV ++ASASHD SVK+WK
Sbjct: 206 HSPVTRMVASASHDKSVKIWK 144
>TC220739 weakly similar to UP|PRP4_HUMAN (O43172) U4/U6 small nuclear
ribonucleoprotein Prp4 (U4/U6 snRNP 60 kDa protein) (WD
splicing factor Prp4) (hPrp4), partial (34%)
Length = 1206
Score = 90.1 bits (222), Expect = 3e-18
Identities = 66/278 (23%), Positives = 121/278 (42%), Gaps = 5/278 (1%)
Frame = +1
Query: 388 CHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRS 447
C FS DGK +A+ +W+M + + H+ TDV + P AT+S DR+
Sbjct: 91 CSFSRDGKWLATCSLTGASKLWSMPKIKKHSIFKGHTERATDVAYSPVHDHLATASADRT 270
Query: 448 VRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTR 507
+ W+ + +++ GH +++ + FHP L ++ + RLW++ + +
Sbjct: 271 AKYWNQGSLLKT---FEGHLDRLARIAFHPSG-KYLGTASFDKTWRLWDIETGDELLLQE 438
Query: 508 GGSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFL 564
G S+ V F + + + + D+ T + L+GH K VL I + +G L
Sbjct: 439 GHSRSVYGLAFHNDGSLAASCGLDSLARVWDLRTGRSILALEGHVKPVLGISFSPNGYHL 618
Query: 565 ASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGY-QSLEAWSPTE 622
A+ ED + RIW K + + N F P L+ Y + + WS +
Sbjct: 619 ATGGEDNTCRIWDLRKK-KSFYTIPAHSNLISQVKFEPHEGYFLVTASYDMTAKVWSGRD 795
Query: 623 GSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+++ H+ + + G I + SHD ++K+W
Sbjct: 796 FKPVKTLSGHEAKVTSVDVLGDGGSIVTVSHDRTIKLW 909
Score = 58.2 bits (139), Expect = 1e-08
Identities = 35/130 (26%), Positives = 67/130 (50%), Gaps = 1/130 (0%)
Frame = +1
Query: 373 QEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRF 432
+ I L + VL FS +G +A+ G + IW++ + F + HS+LI+ V+F
Sbjct: 544 RSILALEGHVKPVLGISFSPNGYHLATGGEDNTCRIWDLRKKKSFYTIPAHSNLISQVKF 723
Query: 433 QPGSTMF-ATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDV 491
+P F T+S+D + ++W + + + L+GH+ +V S+D ++ S
Sbjct: 724 EPHEGYFLVTASYDMTAKVW-SGRDFKPVKTLSGHEAKVTSVDVLGDGGSIVTVSHDR-T 897
Query: 492 IRLWNVNQSE 501
I+LW+ N ++
Sbjct: 898 IKLWSSNPTD 927
>TC207354 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat protein 5,
partial (57%)
Length = 1124
Score = 84.7 bits (208), Expect = 1e-16
Identities = 55/202 (27%), Positives = 100/202 (49%), Gaps = 5/202 (2%)
Frame = +1
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V FS+DG ++ASA +K + IW+ HS I+D+ + S ++
Sbjct: 139 NAVSCVKFSNDGTLLASASLDKTLIIWSSATLTLCHRLVGHSEGISDLAWSSDSHYICSA 318
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S DR++R+WDA+ + IL GHD+ V ++F+P + + S ++ I++W+V +C
Sbjct: 319 SDDRTLRIWDATVGGGCIKILRGHDDAVFCVNFNP-QSSYIVSGSFDETIKVWDVKTGKC 495
Query: 503 MHTTRGGS---KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSIC-WD 558
+HT +G + V + +++ + S I D ET + L L +S +
Sbjct: 496 VHTIKGHTMPVTSVHYNRDGNLIISASHDGSCKIWDTETGNLLKTLIEDKAPAVSFAKFS 675
Query: 559 RSGN-FLASVSEDSARIWSAAS 579
+G LA+ D+ ++W+ S
Sbjct: 676 PNGKLILAATLNDTLKLWNYGS 741
Score = 64.7 bits (156), Expect = 1e-10
Identities = 42/144 (29%), Positives = 69/144 (47%), Gaps = 3/144 (2%)
Frame = +1
Query: 520 GQLLATA-IGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSA 577
G LLA+A + ++ I T + H L GH + + + W +++ S S+D + RIW A
Sbjct: 172 GTLLASASLDKTLIIWSSATLTLCHRLVGHSEGISDLAWSSDSHYICSASDDRTLRIWDA 351
Query: 578 ASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGY-QSLEAWSPTEGSKTWSIAAHQGLI 636
G CI L + F+P S ++ G + ++++ W G +I H +
Sbjct: 352 TVGGGCIKILRGHDDAVFCVNFNP-QSSYIVSGSFDETIKVWDVKTGKCVHTIKGHTMPV 528
Query: 637 AGLADSPVGELIASASHDNSVKVW 660
+ + G LI SASHD S K+W
Sbjct: 529 TSVHYNRDGNLIISASHDGSCKIW 600
>AW761058
Length = 355
Score = 75.9 bits (185), Expect = 5e-14
Identities = 38/114 (33%), Positives = 66/114 (57%), Gaps = 13/114 (11%)
Frame = +3
Query: 334 VESFLSFEDEPADNRIEPFSNLKR-------------ISATCSRNENKGFSLQEIGCLHK 380
VESFLS +D + + ++ + ++++ ++ +K F+ ++ +
Sbjct: 6 VESFLSHDDTDPRDTVGRCMDVSKGAVNTILH*S*NILASSLQKSFSKCFTYSDVNSVRA 185
Query: 381 SINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQP 434
S +KV CHFSSDGK++AS GH+K+V +W ++ + A+ + HS LITDVRF P
Sbjct: 186 STSKVACCHFSSDGKLLASGGHDKRVVLWYTDSLKQKATLEEHSSLITDVRFSP 347
>TC208265 similar to UP|Q6S7B0 (Q6S7B0) TAF5, partial (36%)
Length = 1048
Score = 73.6 bits (179), Expect = 3e-13
Identities = 50/197 (25%), Positives = 92/197 (46%), Gaps = 4/197 (2%)
Frame = +2
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS G + S+ + + +W+ + H++ + DV+F P FA+SS DR+ R
Sbjct: 11 FSPVGDFILSSSADSTIRLWSTKLNANLVCYKGHNYPVWDVQFSPVGHYFASSSHDRTAR 190
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHT---T 506
+W + + I+ L I+ GH V + +H + + + S+ +RLW+V EC+
Sbjct: 191 IW-SMDRIQPLRIMAGHLSDVDCVQWH-ANCNYIATGSSDKTVRLWDVQSGECVRVFVGH 364
Query: 507 RGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
RG + P + + +I + D+ + L L GH V S+ + G+ +AS
Sbjct: 365 RGMILSLAMSPDGRYMASGDEDGTIMMWDLSSGRCLTPLIGHTSCVWSLAFSSEGSVIAS 544
Query: 567 VSED-SARIWSAASDGK 582
S D + ++W + K
Sbjct: 545 GSADCTVKLWDVNTSTK 595
Score = 57.8 bits (138), Expect = 2e-08
Identities = 45/184 (24%), Positives = 79/184 (42%), Gaps = 4/184 (2%)
Frame = +2
Query: 481 DLLCSSDSNDVIRLWNVNQSECMHTTRGGSK---QVRFQPQYGQLLATAIGNSITIIDVE 537
D + SS ++ IRLW+ + + +G + V+F P +++ + I ++
Sbjct: 26 DFILSSSADSTIRLWSTKLNANLVCYKGHNYPVWDVQFSPVGHYFASSSHDRTARIWSMD 205
Query: 538 TDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQS 596
L + GH DV + W + N++A+ S D + R+W S G+C+ S
Sbjct: 206 RIQPLRIMAGHLSDVDCVQWHANCNYIATGSSDKTVRLWDVQS-GECVRVFVGHRGMILS 382
Query: 597 CIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNS 656
P R + ++ W + G + H + LA S G +IAS S D +
Sbjct: 383 LAMSPDGRYMASGDEDGTIMMWDLSSGRCLTPLIGHTSCVWSLAFSSEGSVIASGSADCT 562
Query: 657 VKVW 660
VK+W
Sbjct: 563 VKLW 574
Score = 43.1 bits (100), Expect = 4e-04
Identities = 27/111 (24%), Positives = 49/111 (43%), Gaps = 15/111 (13%)
Frame = +2
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
+L+ S DG+ MAS + + +W++ + + H+ + + F ++ A+ S
Sbjct: 374 ILSLAMSPDGRYMASGDEDGTIMMWDLSSGRCLTPLIGHTSCVWSLAFSSEGSVIASGSA 553
Query: 445 DRSVRLWD---------------ASNPIRSLGILTGHDEQVMSLDFHPRRM 480
D +V+LWD ++N +RSL L V SL F R +
Sbjct: 554 DCTVKLWDVNTSTKVSRAEEKGGSANRLRSLKTLPTKSTPVYSLRFSRRNL 706
Score = 31.2 bits (69), Expect = 1.5
Identities = 16/62 (25%), Positives = 26/62 (41%)
Frame = +2
Query: 599 FHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVK 658
F P +L ++ WS + H + + SPVG AS+SHD + +
Sbjct: 11 FSPVGDFILSSSADSTIRLWSTKLNANLVCYKGHNYPVWDVQFSPVGHYFASSSHDRTAR 190
Query: 659 VW 660
+W
Sbjct: 191 IW 196
>TC220018 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K8K14 (AT5g67320/K8K14_4),
partial (42%)
Length = 1050
Score = 68.9 bits (167), Expect = 7e-12
Identities = 57/235 (24%), Positives = 108/235 (45%), Gaps = 15/235 (6%)
Frame = +1
Query: 378 LHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGST 437
L K + + ++ G + + ++ +W+++ ++ + HS DV ++ +
Sbjct: 19 LSKHKGPIFSLKWNKKGDYLLTGSCDQTAIVWDVKAEEWKQQFEFHSGPTLDVDWR-NNV 195
Query: 438 MFATSSFDRSVRLWDA--SNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLW 495
FATSS D + + ++PI++ TGH +V + + P LL S + ++W
Sbjct: 196 SFATSSTDNMIHVCKIGETHPIKTF---TGHQGEVNCVKWDPTG-SLLASCSDDITAKIW 363
Query: 496 NVNQSECMHTTRGGSKQVRF-----------QPQYGQLLATA-IGNSITIIDVETDSHLH 543
++ Q +H R SK++ P + +LA+A +++ + DVE ++
Sbjct: 364 SMKQDTYLHDLREHSKEIYTIRWSPTGPGTNNPNHKLVLASASFDSTVKLWDVELGKLIY 543
Query: 544 DLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSC 597
L GH V S+ + +G++L S S D S IWS DGK + G F+ C
Sbjct: 544 SLDGHRHPVYSVAFSPNGDYLVSGSLDRSMHIWS-LRDGKIVKTYTGNGGIFEVC 705
Score = 41.6 bits (96), Expect = 0.001
Identities = 50/211 (23%), Positives = 85/211 (39%), Gaps = 13/211 (6%)
Frame = +1
Query: 463 LTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQL 522
L+ H + SL ++ ++ D L + + +W+V E +F+ G
Sbjct: 19 LSKHKGPIFSLKWN-KKGDYLLTGSCDQTAIVWDVKAEEWKQ---------QFEFHSGPT 168
Query: 523 LATAIGNSITIIDVETDSHLHDLK-----------GHDKDVLSICWDRSGNFLASVSED- 570
L N+++ TD+ +H K GH +V + WD +G+ LAS S+D
Sbjct: 169 LDVDWRNNVSFATSSTDNMIHVCKIGETHPIKTFTGHQGEVNCVKWDPTGSLLASCSDDI 348
Query: 571 SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIA 630
+A+IWS D + +L + + WSPT G T
Sbjct: 349 TAKIWSMKQD-TYLHDLREHSKEIYTI-------------------RWSPT-GPGT---- 453
Query: 631 AHQGLIAGLADSPVGELI-ASASHDNSVKVW 660
++P +L+ ASAS D++VK+W
Sbjct: 454 ----------NNPNHKLVLASASFDSTVKLW 516
>TC208379 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K8K14 (AT5g67320/K8K14_4),
partial (42%)
Length = 1062
Score = 68.2 bits (165), Expect = 1e-11
Identities = 53/234 (22%), Positives = 104/234 (43%), Gaps = 14/234 (5%)
Frame = +2
Query: 378 LHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGST 437
L K + + ++ G + + ++ +W+++ ++ + HS DV ++ +
Sbjct: 23 LSKHKGPIFSLKWNKKGDYILTGSCDQTAIVWDVKAEEWKQQFEFHSGWTLDVDWR-NNV 199
Query: 438 MFATSSFDRSVRLWDASN--PIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLW 495
FATSS D + + PIR+ GH +V + + P LL S + ++W
Sbjct: 200 SFATSSTDTKIHVCKIGEKLPIRTF---VGHQSEVNCIKWDPTG-SLLASCSDDMTAKIW 367
Query: 496 NVNQSECMHTTRGGSKQVRF-----------QPQYGQLLATA-IGNSITIIDVETDSHLH 543
++ Q + +H R SK++ P +LA+A +++ + DVE L+
Sbjct: 368 SMKQDKYLHEFREHSKEIYTIRWSPTGPGTNNPNKNLVLASASFDSTVKLWDVELGKLLY 547
Query: 544 DLKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSC 597
L GH V S+ + +G ++AS S D + + + +GK + G F+ C
Sbjct: 548 SLNGHRDRVYSVAFSPNGEYIASGSPDRSMLIWSLKEGKIVKTYTGDGGIFEVC 709
Score = 62.4 bits (150), Expect = 6e-10
Identities = 54/219 (24%), Positives = 94/219 (42%), Gaps = 21/219 (9%)
Frame = +2
Query: 463 LTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQL 522
L+ H + SL ++ ++ D + + + +W+V E +F+ G
Sbjct: 23 LSKHKGPIFSLKWN-KKGDYILTGSCDQTAIVWDVKAEEWKQ---------QFEFHSGWT 172
Query: 523 LATAIGNSITIIDVETDSHLHDLK-----------GHDKDVLSICWDRSGNFLASVSED- 570
L N+++ TD+ +H K GH +V I WD +G+ LAS S+D
Sbjct: 173 LDVDWRNNVSFATSSTDTKIHVCKIGEKLPIRTFVGHQSEVNCIKWDPTGSLLASCSDDM 352
Query: 571 SARIWSAASDGKCIGELHSVGNKFQSCIF--------HPGYRSLLIIGGYQS-LEAWSPT 621
+A+IWS D K + E + + + +P +L + S ++ W
Sbjct: 353 TAKIWSMKQD-KYLHEFREHSKEIYTIRWSPTGPGTNNPNKNLVLASASFDSTVKLWDVE 529
Query: 622 EGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G +S+ H+ + +A SP GE IAS S D S+ +W
Sbjct: 530 LGKLLYSLNGHRDRVYSVAFSPNGEYIASGSPDRSMLIW 646
>CO983685
Length = 720
Score = 65.5 bits (158), Expect = 7e-11
Identities = 39/81 (48%), Positives = 53/81 (65%)
Frame = -3
Query: 326 DAKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKV 385
D ++NVESFLS +D D R + F LK + + + +KGFS E+G + KS NKV
Sbjct: 325 DVGSLEDNVESFLSQDD--GDGR-DLFGTLKP-NPSEHADASKGFSFSEVGSICKSNNKV 158
Query: 386 LTCHFSSDGKVMASAGHEKKV 406
+ CHFSSDGK++AS G +KKV
Sbjct: 157 VCCHFSSDGKLLASVGLDKKV 95
>TC229098 similar to PIR|T46032|T46032 WD-40 repeat regulatory protein tup1
homolog - Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (59%)
Length = 580
Score = 65.1 bits (157), Expect = 1e-10
Identities = 41/171 (23%), Positives = 78/171 (44%), Gaps = 7/171 (4%)
Frame = +3
Query: 408 IWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHD 467
+W++ + H++ + V F P S + + SFD +VR+WD + + L +L H
Sbjct: 6 LWDVPTGSLIKTLHGHTNYVFCVNFNPQSNIIVSGSFDETVRVWDVKSG-KCLKVLPAHS 182
Query: 468 EQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQ----VRFQPQYGQLL 523
+ V ++DF+ R L+ SS + + R+W+ + CM T V+F P +L
Sbjct: 183 DPVTAVDFN-RDGSLIVSSSYDGLCRIWDASTGHCMKTLIDDDNPPVSFVKFSPNAKFIL 359
Query: 524 ATAIGNSITIIDVETDSHLHDLKGHDKD---VLSICWDRSGNFLASVSEDS 571
+ N++ + + T L GH + S +G ++ SE++
Sbjct: 360 VGTLDNTLRLWNYSTGKFLKTYTGHVNSKYCISSTFSTTNGKYIVGGSEEN 512
Score = 57.0 bits (136), Expect = 3e-08
Identities = 43/180 (23%), Positives = 81/180 (44%), Gaps = 6/180 (3%)
Frame = +3
Query: 375 IGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQP 434
I LH N V +F+ ++ S ++ V +W++++ + HS +T V F
Sbjct: 33 IKTLHGHTNYVFCVNFNPQSNIIVSGSFDETVRVWDVKSGKCLKVLPAHSDPVTAVDFNR 212
Query: 435 GSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRL 494
++ +SS+D R+WDAS ++ + V + F P +L + N +RL
Sbjct: 213 DGSLIVSSSYDGLCRIWDASTGHCMKTLIDDDNPPVSFVKFSPNAKFILVGTLDN-TLRL 389
Query: 495 WNVNQSECMHTTRG--GSK---QVRFQPQYGQ-LLATAIGNSITIIDVETDSHLHDLKGH 548
WN + + + T G SK F G+ ++ + N I + D+++ + L+GH
Sbjct: 390 WNYSTGKFLKTYTGHVNSKYCISSTFSTTNGKYIVGGSEENYIYLWDLQSRKIVQKLEGH 569
Score = 37.0 bits (84), Expect = 0.028
Identities = 34/130 (26%), Positives = 60/130 (46%), Gaps = 4/130 (3%)
Frame = +3
Query: 535 DVETDSHLHDLKGHDKDVLSICWDRSGNFLASVS-EDSARIWSAASDGKCIGELHSVGNK 593
DV T S + L GH V + ++ N + S S +++ R+W S GKC+ L + +
Sbjct: 12 DVPTGSLIKTLHGHTNYVFCVNFNPQSNIIVSGSFDETVRVWDVKS-GKCLKVLPAHSDP 188
Query: 594 FQSCIFHPGYRSLLIIGGYQSL-EAWSPTEG--SKTWSIAAHQGLIAGLADSPVGELIAS 650
+ F+ SL++ Y L W + G KT I ++ + SP + I
Sbjct: 189 VTAVDFNRD-GSLIVSSSYDGLCRIWDASTGHCMKT-LIDDDNPPVSFVKFSPNAKFILV 362
Query: 651 ASHDNSVKVW 660
+ DN++++W
Sbjct: 363 GTLDNTLRLW 392
>TC216518 weakly similar to UP|Q9LW87 (Q9LW87) Coatomer protein complex, beta
prime; beta'-COP protein, partial (29%)
Length = 1189
Score = 59.3 bits (142), Expect = 5e-09
Identities = 50/242 (20%), Positives = 98/242 (39%), Gaps = 12/242 (4%)
Frame = +1
Query: 350 EPFSNLKRISATCS------RNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHE 403
EP+ L S T S + E K + E + V + F + + +A +
Sbjct: 130 EPWILLGLYSGTISIWNYQTKTEEKSLKISE--------SPVRSAKFIARENWIVAATDD 285
Query: 404 KKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGIL 463
K + ++N + + H I + P ++S D+ ++LW+
Sbjct: 286 KNIHVYNYDKMEKIVEFAEHKDYIRSLAVHPVLPYVISASDDQVLKLWNWRKGWSCYENF 465
Query: 464 TGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQY---- 519
GH VM + F+P+ S+ + +++W+++ S T G K V +
Sbjct: 466 EGHSHYVMQVAFNPKDPSTFASASLDGTLKIWSLDSSAPNFTLEGHQKGVNCVDYFITND 645
Query: 520 -GQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSA 577
LL+ + + + D + + + L+GH+ +V +IC + + SEDS +IW A
Sbjct: 646 KQYLLSGSDDYTAKVWDYHSRNCVQTLEGHENNVTAICAHPELPIIITASEDSTVKIWDA 825
Query: 578 AS 579
+
Sbjct: 826 VT 831
>TC228105 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;} , complete
Length = 1113
Score = 58.2 bits (139), Expect = 1e-08
Identities = 56/281 (19%), Positives = 114/281 (39%), Gaps = 23/281 (8%)
Frame = +3
Query: 403 EKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGI 462
++ V +W ++ + H + V P ++ A+SS D VR++D + ++
Sbjct: 111 DETVRLWRSDDLVLDRTNTGHCLGVASVAAHPLGSVAASSSLDSFVRVFDVDSNA-TIAT 287
Query: 463 LTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTT------------RGGS 510
L +V + F P+ L + + ++LW+ + E + T + GS
Sbjct: 288 LEAPPSEVWQMRFDPKGAILAVAGGGSASVKLWDTSSWELVATLSIPRPEGQKPTDKSGS 467
Query: 511 KQ----VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
K+ V + P +L ++ +I++ DV LH L+GH V S+ + L
Sbjct: 468 KKFVLSVAWSPDGKRLACGSMDGTISVFDVPRAKFLHHLEGHFMPVRSLVYSPYDPRLLF 647
Query: 567 VSEDSARIWSAASDGKCIGELHSVGNKFQSCI-FHPGYRSLLIIGGYQSLEAWSPTEGSK 625
+ D + ++GK + S + C+ P ++ +S+ W +
Sbjct: 648 TASDDGNVHMYDAEGKALIGTMSGHASWVLCVDVSPDGAAIATGSSDRSVRLWDLNMRAS 827
Query: 626 TWSIAAHQGLIAGLADSPV------GELIASASHDNSVKVW 660
+++ H + G+A P G +AS S D S+ ++
Sbjct: 828 VQTMSNHSDQVWGVAFRPPGGSDVRGGRLASVSDDKSISLY 950
Score = 44.7 bits (104), Expect = 1e-04
Identities = 25/85 (29%), Positives = 41/85 (47%)
Frame = +3
Query: 393 DGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWD 452
D +++ +A + V +++ E + H+ + V P AT S DRSVRLWD
Sbjct: 630 DPRLLFTASDDGNVHMYDAEGKALIGTMSGHASWVLCVDVSPDGAAIATGSSDRSVRLWD 809
Query: 453 ASNPIRSLGILTGHDEQVMSLDFHP 477
N S+ ++ H +QV + F P
Sbjct: 810 L-NMRASVQTMSNHSDQVWGVAFRP 881
Score = 34.7 bits (78), Expect = 0.14
Identities = 24/86 (27%), Positives = 41/86 (46%), Gaps = 6/86 (6%)
Frame = +3
Query: 375 IGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQP 434
IG + + VL S DG +A+ ++ V +W++ + HS + V F+P
Sbjct: 702 IGTMSGHASWVLCVDVSPDGAAIATGSSDRSVRLWDLNMRASVQTMSNHSDQVWGVAFRP 881
Query: 435 --GSTM----FATSSFDRSVRLWDAS 454
GS + A+ S D+S+ L+D S
Sbjct: 882 PGGSDVRGGRLASVSDDKSISLYDYS 959
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.130 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,981,577
Number of Sequences: 63676
Number of extensions: 467806
Number of successful extensions: 2647
Number of sequences better than 10.0: 153
Number of HSP's better than 10.0 without gapping: 2383
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2559
length of query: 661
length of database: 12,639,632
effective HSP length: 103
effective length of query: 558
effective length of database: 6,081,004
effective search space: 3393200232
effective search space used: 3393200232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0114.1


