

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0113.7
(147 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
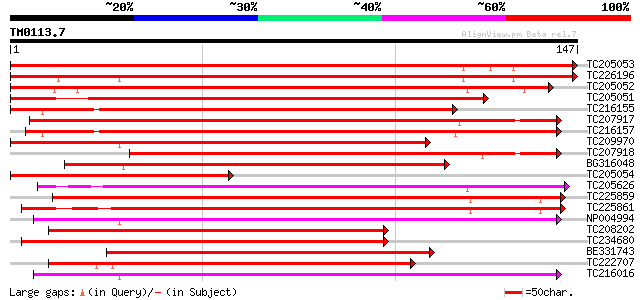
Score E
Sequences producing significant alignments: (bits) Value
TC205053 homologue to UP|Q8L5W2 (Q8L5W2) BZIP transcription fact... 212 6e-56
TC226196 UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2, comp... 199 5e-52
TC205052 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor... 182 5e-47
TC205051 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor... 150 2e-37
TC216155 weakly similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription... 140 3e-34
TC207917 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor... 134 2e-32
TC216157 weakly similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription... 132 6e-32
TC209970 weakly similar to UP|Q8GTZ1 (Q8GTZ1) BZIP transcription... 125 1e-29
TC207918 similar to UP|Q9FWS7 (Q9FWS7) F1B16.8 protein (At1g7539... 109 4e-25
BG316048 similar to GP|22597162|gb| bZIP transcription factor AT... 104 1e-23
TC205054 homologue to UP|Q8L5W2 (Q8L5W2) BZIP transcription fact... 103 4e-23
TC205626 similar to UP|Q9FWS7 (Q9FWS7) F1B16.8 protein (At1g7539... 97 3e-21
TC225859 similar to UP|O22677 (O22677) BZIP DNA-binding protein,... 95 1e-20
TC225861 similar to UP|Q941L6 (Q941L6) BZIP transcription factor... 93 5e-20
NP004994 G/HBF-1 85 1e-17
TC208202 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor... 83 4e-17
TC234680 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor... 81 2e-16
BE331743 similar to GP|24460973|gb| bZIP transcription factor {C... 81 2e-16
TC222707 homologue to UP|Q9AT29 (Q9AT29) BZip transcription fact... 79 6e-16
TC216016 similar to UP|P93839 (P93839) G/HBF-1, partial (61%) 77 4e-15
>TC205053 homologue to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (89%)
Length = 1432
Score = 212 bits (539), Expect = 6e-56
Identities = 115/160 (71%), Positives = 135/160 (83%), Gaps = 13/160 (8%)
Frame = +2
Query: 1 MASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQV 60
MASSSGTSSGSS++QNSGSEE+LQA+MDQRKRKRMISNRESARRSRMRKQKHLD+L +QV
Sbjct: 716 MASSSGTSSGSSLLQNSGSEEDLQAVMDQRKRKRMISNRESARRSRMRKQKHLDDLVSQV 895
Query: 61 AQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATN---- 116
AQLR ENQQ+LTS+N+TTQ+++ ++AENSVL AQ+GELSHRLESLNEI+D +NAT
Sbjct: 896 AQLRKENQQILTSVNITTQQYLSVEAENSVLRAQVGELSHRLESLNEIVDVLNATTVAGF 1075
Query: 117 GVFAANS-FVDPIP--------SAYYLNHPIMASADILQY 147
G A +S FV+PI + YLNHPIMASADILQY
Sbjct: 1076GAAATSSTFVEPINNNSFFNPLNMGYLNHPIMASADILQY 1195
>TC226196 UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2, complete
Length = 1518
Score = 199 bits (505), Expect = 5e-52
Identities = 111/166 (66%), Positives = 134/166 (79%), Gaps = 19/166 (11%)
Frame = +2
Query: 1 MASSSGTSSGS-SMIQNSGSEEELQALM-DQRKRKRMISNRESARRSRMRKQKHLDELAA 58
MA SSGTSSGS S++QNSGSEE+LQA+M DQRKRKRMISNRESARRSRMRKQKHLD+L +
Sbjct: 737 MACSSGTSSGSLSLLQNSGSEEDLQAMMEDQRKRKRMISNRESARRSRMRKQKHLDDLVS 916
Query: 59 QVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATN-- 116
QVAQLR ENQQ+LTS+N+TTQ+++ ++AENSVL AQ+GELSHRLESLNEI+D +NAT
Sbjct: 917 QVAQLRKENQQILTSVNITTQQYLSVEAENSVLRAQVGELSHRLESLNEIVDVLNATTTV 1096
Query: 117 ---GVFAANSFVDPIP------------SAYYLNHPIMASADILQY 147
G A+++FV+P+ + YLN PIMASADILQY
Sbjct: 1097AGFGAAASSTFVEPMNNNNNSFFNFNPLNMGYLNQPIMASADILQY 1234
>TC205052 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (58%)
Length = 953
Score = 182 bits (462), Expect = 5e-47
Identities = 104/149 (69%), Positives = 124/149 (82%), Gaps = 8/149 (5%)
Frame = +1
Query: 1 MASSSGTSSG--SSMIQN-SGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELA 57
MA SSGTSSG SSM+QN SGSEEELQALM+QRKRKRMISNRESARRSRMRKQKHLD+LA
Sbjct: 505 MACSSGTSSGATSSMLQNNSGSEEELQALMEQRKRKRMISNRESARRSRMRKQKHLDDLA 684
Query: 58 AQVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNG 117
+QV QLR+EN Q+LTS+NLTTQ+++ ++AENSVL AQ+ ELSH LESLNEII F+NAT+G
Sbjct: 685 SQVTQLRNENHQILTSVNLTTQKYLAVEAENSVLRAQVNELSHWLESLNEIIHFLNATDG 864
Query: 118 --VFAANSFVDPIPSAY---YLNHPIMAS 141
+SF +P + + YL+ PIMAS
Sbjct: 865 GPPPPPSSFFEPDATFFNKAYLSQPIMAS 951
>TC205051 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (63%)
Length = 773
Score = 150 bits (380), Expect = 2e-37
Identities = 79/124 (63%), Positives = 99/124 (79%)
Frame = +1
Query: 1 MASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQV 60
MA SSGTSSG+S ELQ +MDQRKRKRMISNRESARRSRMRKQKHLD+LA+Q+
Sbjct: 397 MACSSGTSSGTS--------SELQGMMDQRKRKRMISNRESARRSRMRKQKHLDDLASQL 552
Query: 61 AQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVFA 120
QLRS+NQQ+LTS+NLT+ +++ ++AENSVL AQ+ ELSHRL+SLN+II +N +
Sbjct: 553 TQLRSQNQQLLTSVNLTSHKYLAVEAENSVLRAQVNELSHRLDSLNQIIHLLNFFKPNTS 732
Query: 121 ANSF 124
N+F
Sbjct: 733 TNTF 744
>TC216155 weakly similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor
ATB2, partial (66%)
Length = 1264
Score = 140 bits (352), Expect = 3e-34
Identities = 72/119 (60%), Positives = 100/119 (83%), Gaps = 3/119 (2%)
Frame = +1
Query: 1 MASSSGT---SSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELA 57
MAS G+ SSGSS +QNSGSE + + +M+QRKRKRM+SNRESARRSR+RKQ+HL+ L+
Sbjct: 583 MASPGGSGTYSSGSSSLQNSGSEGD-RDIMEQRKRKRMLSNRESARRSRIRKQQHLEGLS 759
Query: 58 AQVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATN 116
AQ+ QL+ EN Q+ T++++TTQ ++ ++AEN++L AQMGELS+RL SLNE+I F+N+TN
Sbjct: 760 AQLDQLKKENAQINTNISITTQMYLNVEAENAILRAQMGELSNRLNSLNEMISFINSTN 936
>TC207917 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (75%)
Length = 869
Score = 134 bits (336), Expect = 2e-32
Identities = 76/151 (50%), Positives = 101/151 (66%), Gaps = 13/151 (8%)
Frame = +2
Query: 6 GTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRS 65
GTSS S ++Q+SGSEE+LQ LM+QRK+KR SNRESARRSRMRKQKHLD+L AQV L+
Sbjct: 5 GTSSTSILLQSSGSEEDLQLLMEQRKKKRKQSNRESARRSRMRKQKHLDDLIAQVDLLKK 184
Query: 66 ENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNAT---------- 115
+ +N+TTQ + ++AENS+L AQ EL+ L+SLN+II+ +N T
Sbjct: 185 QKSLSFKKVNITTQHCLKVEAENSILGAQKTELTQSLQSLNDIINLINTTTTSDHNNYYY 364
Query: 116 ---NGVFAANSFVDPIPSAYYLNHPIMASAD 143
N N ++P+ A YLN PI+A+AD
Sbjct: 365 NNNNNNNNNNFMMNPMHMA-YLNQPIVATAD 454
>TC216157 weakly similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor
ATB2, partial (68%)
Length = 875
Score = 132 bits (332), Expect = 6e-32
Identities = 79/156 (50%), Positives = 105/156 (66%), Gaps = 17/156 (10%)
Frame = +2
Query: 5 SGT-SSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQL 63
SGT SSGSS +QNSGSE + + +M+QRKRKRM+SNRESARRSRMRKQ+HL+ L+AQ+ QL
Sbjct: 107 SGTYSSGSSSLQNSGSEGD-RDIMEQRKRKRMLSNRESARRSRMRKQQHLEGLSAQLDQL 283
Query: 64 RSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNA--------- 114
+ EN QM T++ ++TQ ++ ++AEN++L AQM ELS RL SLNE+I +N+
Sbjct: 284 KKENTQMNTNIGISTQLYLNVEAENAILRAQMEELSKRLNSLNEMISLINSTTTTNNCLM 463
Query: 115 -------TNGVFAANSFVDPIPSAYYLNHPIMASAD 143
T +F F+D LN IMA AD
Sbjct: 464 FDEAQETTTQLFNDCGFMDAWNYGIPLNQQIMAYAD 571
>TC209970 weakly similar to UP|Q8GTZ1 (Q8GTZ1) BZIP transcription factor,
partial (62%)
Length = 1041
Score = 125 bits (313), Expect = 1e-29
Identities = 64/110 (58%), Positives = 88/110 (79%), Gaps = 1/110 (0%)
Frame = +2
Query: 1 MASSSGTSSGSSMIQNSGSEEELQALM-DQRKRKRMISNRESARRSRMRKQKHLDELAAQ 59
MASSSG SS S+ Q+SGSE +LQ ++ D+RK KR SNRESARRSRMRK+ HLD+L Q
Sbjct: 254 MASSSGNSSISTKSQSSGSEGDLQVVITDERKNKRKQSNRESARRSRMRKRNHLDQLTKQ 433
Query: 60 VAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEII 109
++QL N ++L ++++TTQ ++ ++AENS+L AQMGELS RL+SLN+I+
Sbjct: 434 LSQLAKNNGEILATIDITTQHYLNVEAENSILRAQMGELSQRLQSLNDIV 583
>TC207918 similar to UP|Q9FWS7 (Q9FWS7) F1B16.8 protein (At1g75390), partial
(40%)
Length = 882
Score = 109 bits (273), Expect = 4e-25
Identities = 61/130 (46%), Positives = 84/130 (63%), Gaps = 18/130 (13%)
Frame = +2
Query: 32 RKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAENSVL 91
+KR SNRESARRSRMRKQKHLD+L AQV L+ + L +++TT+ ++ + AENS+L
Sbjct: 2 KKRKQSNRESARRSRMRKQKHLDDLIAQVDHLKKQKSLTLMKVDITTKHYLEVKAENSIL 181
Query: 92 SAQMGELSHRLESLNEIIDFVNATNGVFAA------------------NSFVDPIPSAYY 133
AQ EL+ L+SLN+IID +N TNGV+ N+F++P+ A Y
Sbjct: 182 WAQKTELTQSLQSLNDIIDLINTTNGVYHTDCYDINNHNNHNNYYNNNNNFMNPMHMA-Y 358
Query: 134 LNHPIMASAD 143
LN PI+A+AD
Sbjct: 359 LNQPIVATAD 388
>BG316048 similar to GP|22597162|gb| bZIP transcription factor ATB2 {Glycine
max}, partial (55%)
Length = 304
Score = 104 bits (260), Expect = 1e-23
Identities = 59/101 (58%), Positives = 75/101 (73%), Gaps = 1/101 (0%)
Frame = +2
Query: 15 QNSGSEEELQALMD-QRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTS 73
QNSGSEE+ QA+M+ QRK RMISNR SARRS M+KQ H L VAQ R E + + +
Sbjct: 2 QNSGSEEDSQAMMEHQRKTTRMISNRYSARRSPMKKQSHFHYLVPPVAQPR*EIRHIPAT 181
Query: 74 LNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNA 114
++TTQ+ + ++AENSVL+A +GELSH LESLN I+D VNA
Sbjct: 182 GHITTQQCLSVEAENSVLTAHVGELSHMLESLN*ILDLVNA 304
>TC205054 homologue to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (35%)
Length = 462
Score = 103 bits (256), Expect = 4e-23
Identities = 51/58 (87%), Positives = 57/58 (97%)
Frame = +1
Query: 1 MASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAA 58
MASSSGTSSGSS++QNSGSEE+LQA+MDQRKRKRMISNRESARRSRMRKQKHLD+L +
Sbjct: 289 MASSSGTSSGSSLLQNSGSEEDLQAVMDQRKRKRMISNRESARRSRMRKQKHLDDLVS 462
>TC205626 similar to UP|Q9FWS7 (Q9FWS7) F1B16.8 protein (At1g75390), partial
(34%)
Length = 1184
Score = 97.1 bits (240), Expect = 3e-21
Identities = 59/141 (41%), Positives = 85/141 (59%), Gaps = 3/141 (2%)
Frame = +3
Query: 8 SSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSEN 67
SSGSS G + A MD+RKRKRM SNRESARRSRM+KQK L++L+ ++L+ EN
Sbjct: 435 SSGSS---EGGGDP---AAMDERKRKRMESNRESARRSRMKKQKLLEDLSDVASRLQGEN 596
Query: 68 QQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNG---VFAANSF 124
++ S+ + ++ I+A N +L AQ EL+ RL LN I++ + G F
Sbjct: 597 VRLAQSIKAKEEAYVEIEAANDILRAQTMELADRLRFLNSILEIADEVGGGGESFEIPQI 776
Query: 125 VDPIPSAYYLNHPIMASADIL 145
DP+ + + HP+MAS D+L
Sbjct: 777 PDPLFMPWQIPHPMMASPDML 839
>TC225859 similar to UP|O22677 (O22677) BZIP DNA-binding protein, partial
(59%)
Length = 1132
Score = 95.1 bits (235), Expect = 1e-20
Identities = 55/137 (40%), Positives = 85/137 (61%), Gaps = 4/137 (2%)
Frame = +1
Query: 12 SMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQML 71
S +SGSE ++D+RKRKRM+SNRESARRSRMRKQK L++L +V++L+S N+++
Sbjct: 448 STTTSSGSEGGDPHIIDERKRKRMLSNRESARRSRMRKQKQLEDLTDEVSRLQSANKKLA 627
Query: 72 TSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGV-FAANSFVDPIPS 130
++ + + +A NS+L AQ EL+ RL LN I++ G+ DP+
Sbjct: 628 ENIEAKEEACVETEAANSILRAQTMELADRLRFLNSILEIAEEVEGLSVEIPEIPDPLLK 807
Query: 131 AYYLNH---PIMASADI 144
+ + H PIMA+A++
Sbjct: 808 PWQIPHPIQPIMATANM 858
>TC225861 similar to UP|Q941L6 (Q941L6) BZIP transcription factor BZI-3,
partial (43%)
Length = 1239
Score = 92.8 bits (229), Expect = 5e-20
Identities = 56/145 (38%), Positives = 88/145 (60%), Gaps = 4/145 (2%)
Frame = +3
Query: 4 SSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQL 63
S+ TSSGS G + ++ +D+RKRKRM+SNRESARRSRMRKQK L++L +V++L
Sbjct: 465 STATSSGSE----GGGDPQM---IDERKRKRMLSNRESARRSRMRKQKQLEDLTDEVSRL 623
Query: 64 RSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGV-FAAN 122
+ N+++ ++ + + +A NS+L AQ EL+ RL LN I++ G+
Sbjct: 624 QGANKKLAENIKAKEEACVETEAANSILRAQTMELADRLRFLNSILEIAEEVEGLSVEIP 803
Query: 123 SFVDPIPSAYYLNH---PIMASADI 144
DP+ + + H PIMA+A++
Sbjct: 804 EIPDPLLKPWQIPHPIQPIMATANM 878
>NP004994 G/HBF-1
Length = 1137
Score = 84.7 bits (208), Expect = 1e-17
Identities = 55/144 (38%), Positives = 76/144 (52%), Gaps = 7/144 (4%)
Frame = +1
Query: 7 TSSGSSMIQNSGSEEELQALM-------DQRKRKRMISNRESARRSRMRKQKHLDELAAQ 59
T+SGSS Q+ E E +A D ++ +RM+SNRESARRSR RKQ HL EL Q
Sbjct: 433 TTSGSSREQSDDDEAEGEAETTQGMDPADAKRVRRMLSNRESARRSRRRKQAHLTELETQ 612
Query: 60 VAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVF 119
V+QLR EN +L L +Q++ +N VL A + L +++ E + V N +F
Sbjct: 613 VSQLRVENSSLLKRLTDISQKYNEAAVDNRVLKADVETLRTKVKMAEETVKRVTGLNPLF 792
Query: 120 AANSFVDPIPSAYYLNHPIMASAD 143
A S + + Y P SAD
Sbjct: 793 QAMSEISSMVMPSYSGSPSDTSAD 864
>TC208202 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor, partial
(51%)
Length = 758
Score = 83.2 bits (204), Expect = 4e-17
Identities = 44/88 (50%), Positives = 61/88 (69%)
Frame = +1
Query: 11 SSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQM 70
SS + ++E+ Q+L+++RK +RMISNRESARRSRMRKQKHLDEL +QV LR+EN Q+
Sbjct: 175 SSNSTSDEADEQQQSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVVWLRNENHQL 354
Query: 71 LTSLNLTTQRFMIIDAENSVLSAQMGEL 98
+ LN ++ + EN L + EL
Sbjct: 355 MDKLNHVSESHDKVAQENVQLREEASEL 438
>TC234680 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor, partial
(49%)
Length = 571
Score = 81.3 bits (199), Expect = 2e-16
Identities = 46/95 (48%), Positives = 64/95 (66%)
Frame = +2
Query: 4 SSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQL 63
S +S SS + ++E+ +L+++RK +RMISNRESARRSRMRKQKHLDEL +QV L
Sbjct: 269 SPQSSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVVWL 448
Query: 64 RSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGEL 98
R+EN Q++ LN ++ + EN+ L Q EL
Sbjct: 449 RNENHQLMDKLNHVSESHDQVMQENAQLKEQALEL 553
>BE331743 similar to GP|24460973|gb| bZIP transcription factor {Capsicum
chinense}, partial (35%)
Length = 292
Score = 81.3 bits (199), Expect = 2e-16
Identities = 40/85 (47%), Positives = 60/85 (70%)
Frame = +2
Query: 26 LMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIID 85
++D+RKRKRM+SNRESARRSRMRKQK L++L +V++L+ N+ + ++ + +
Sbjct: 26 MIDERKRKRMLSNRESARRSRMRKQKQLEDLTDEVSRLQGANKMLAENIKAKEDACVETE 205
Query: 86 AENSVLSAQMGELSHRLESLNEIID 110
A NS+L AQ EL+ RL LN I++
Sbjct: 206 AANSILRAQTMELADRLRCLNSILE 280
>TC222707 homologue to UP|Q9AT29 (Q9AT29) BZip transcription factor, complete
Length = 671
Score = 79.3 bits (194), Expect = 6e-16
Identities = 46/98 (46%), Positives = 64/98 (64%), Gaps = 3/98 (3%)
Frame = +3
Query: 11 SSMIQNSGSEE--ELQA-LMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSEN 67
S + NS S+E E+Q ++D+RK +RMISNRESARRSRMRKQKHLDEL +QV +LR+EN
Sbjct: 252 SCLSSNSTSDEADEIQFNIIDERKHRRMISNRESARRSRMRKQKHLDELWSQVVRLRTEN 431
Query: 68 QQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESL 105
++ LN ++ + EN+ L + L L +
Sbjct: 432 HNLIDKLNHVSESHDRVLQENARLKEEASALRQMLADM 545
>TC216016 similar to UP|P93839 (P93839) G/HBF-1, partial (61%)
Length = 1764
Score = 76.6 bits (187), Expect = 4e-15
Identities = 50/144 (34%), Positives = 74/144 (50%), Gaps = 7/144 (4%)
Frame = +3
Query: 7 TSSGSSMIQNSGSEEELQALM-------DQRKRKRMISNRESARRSRMRKQKHLDELAAQ 59
T SGSS Q+ E E + M D ++ +RM+SNRESARRSR RKQ HL EL Q
Sbjct: 645 TISGSSGEQSDDEEAEGEINMTGNMTPVDAKRVRRMLSNRESARRSRRRKQAHLTELETQ 824
Query: 60 VAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVF 119
V+QLRSEN +L +Q++ +N VL A + L +++ E + + + +
Sbjct: 825 VSQLRSENSSLLKRFTDVSQKYSNAAVDNRVLKADVETLRAKVKMAEETVKRITGLSPML 1004
Query: 120 AANSFVDPIPSAYYLNHPIMASAD 143
A + + + + P SAD
Sbjct: 1005HAMTEMSSLGMPLFDESPSETSAD 1076
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.124 0.319
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,792,556
Number of Sequences: 63676
Number of extensions: 30812
Number of successful extensions: 275
Number of sequences better than 10.0: 86
Number of HSP's better than 10.0 without gapping: 257
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 264
length of query: 147
length of database: 12,639,632
effective HSP length: 88
effective length of query: 59
effective length of database: 7,036,144
effective search space: 415132496
effective search space used: 415132496
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0113.7


