

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.9
(341 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
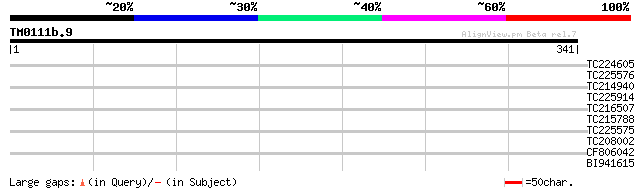
TC224605 homologue to UP|PRO1_SOYBN (O65809) Profilin 1 (GmPRO1)... 31 0.71
TC225576 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, ... 30 1.6
TC214940 homologue to UP|Q75NI2 (Q75NI2) Type 1 metallothionein,... 30 2.1
TC225914 homologue to UP|Q75NI2 (Q75NI2) Type 1 metallothionein,... 30 2.1
TC216507 similar to GB|AAL15354.1|16323240|AY057724 AT5g14920/F2... 30 2.1
TC215788 weakly similar to UP|O22627 (O22627) Auxin-induced prot... 29 2.7
TC225575 similar to UP|Q9M6E4 (Q9M6E4) Poly(A)-binding protein (... 29 2.7
TC208002 similar to PIR|D86282|D86282 protein F10B6.26 [imported... 29 2.7
CF806042 28 4.6
BI941615 28 4.6
>TC224605 homologue to UP|PRO1_SOYBN (O65809) Profilin 1 (GmPRO1) (Allergen
Gly m 3), complete
Length = 713
Score = 31.2 bits (69), Expect = 0.71
Identities = 16/38 (42%), Positives = 19/38 (49%), Gaps = 3/38 (7%)
Frame = +1
Query: 209 W*MERLAVVHSHP---YFRHSEAYGH*SCYCEEWRRSL 243
W + R +H HP + RHS G CYCEE R L
Sbjct: 235 WIVSRCHQIHGHPGRAWCRHSRQEGSWWCYCEEDRCGL 348
>TC225576 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, partial (93%)
Length = 2491
Score = 30.0 bits (66), Expect = 1.6
Identities = 15/35 (42%), Positives = 20/35 (56%)
Frame = -2
Query: 160 ASRRLLI*VGEWQHLLLV*GLEWAWGLGLVSSFCA 194
A+RR+ I W+HLLL W W LGL+ C+
Sbjct: 1824 AARRVTIDTSPWKHLLL---HHWHWLLGLLHRTCS 1729
>TC214940 homologue to UP|Q75NI2 (Q75NI2) Type 1 metallothionein, complete
Length = 426
Score = 29.6 bits (65), Expect = 2.1
Identities = 12/30 (40%), Positives = 20/30 (66%)
Frame = -2
Query: 289 C*YGYWSIM*FKLMTVDFDVILLPRRCALV 318
C*Y Y+S + FK++ +D + RRC+L+
Sbjct: 356 C*YIYYSYLCFKVINLDLTCSCMGRRCSLI 267
>TC225914 homologue to UP|Q75NI2 (Q75NI2) Type 1 metallothionein, complete
Length = 582
Score = 29.6 bits (65), Expect = 2.1
Identities = 12/30 (40%), Positives = 20/30 (66%)
Frame = -3
Query: 289 C*YGYWSIM*FKLMTVDFDVILLPRRCALV 318
C*Y Y+S + FK++ +D + RRC+L+
Sbjct: 361 C*YIYYSYLCFKVINLDLTCSCMGRRCSLI 272
>TC216507 similar to GB|AAL15354.1|16323240|AY057724 AT5g14920/F2G14_40
{Arabidopsis thaliana;} , partial (22%)
Length = 680
Score = 29.6 bits (65), Expect = 2.1
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = -2
Query: 70 FPSWISWLASSQVGHVGSFEAAWWSWYPRCDTREY 104
+ SW W +V H GS+ +WW +Y R R Y
Sbjct: 187 YRSWWRWSNRRRVHHWGSWCWSWWLYYWRLRGRGY 83
>TC215788 weakly similar to UP|O22627 (O22627) Auxin-induced protein, partial
(75%)
Length = 1350
Score = 29.3 bits (64), Expect = 2.7
Identities = 11/22 (50%), Positives = 16/22 (72%)
Frame = -2
Query: 66 LYLVFPSWISWLASSQVGHVGS 87
+YL+FPSW S L+S V +G+
Sbjct: 512 MYLIFPSWTSLLSSPMVSSMGT 447
>TC225575 similar to UP|Q9M6E4 (Q9M6E4) Poly(A)-binding protein (Fragment),
partial (38%)
Length = 791
Score = 29.3 bits (64), Expect = 2.7
Identities = 15/35 (42%), Positives = 20/35 (56%)
Frame = -2
Query: 160 ASRRLLI*VGEWQHLLLV*GLEWAWGLGLVSSFCA 194
A+RR+ I W+HLLL W W LGL+ C+
Sbjct: 160 AARRVTIDTPPWKHLLL---HHWHWLLGLLHRTCS 65
>TC208002 similar to PIR|D86282|D86282 protein F10B6.26 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(84%)
Length = 932
Score = 29.3 bits (64), Expect = 2.7
Identities = 14/31 (45%), Positives = 18/31 (57%)
Frame = +2
Query: 92 WWSWYPRCDTREYFLVEEIGVGLCAKI*EVV 122
WW W+P+ R +FLV VG C I +VV
Sbjct: 77 WWLWFPKKMLRHWFLVP---VGSCNVIEKVV 160
>CF806042
Length = 604
Score = 28.5 bits (62), Expect = 4.6
Identities = 9/16 (56%), Positives = 10/16 (62%)
Frame = +3
Query: 87 SFEAAWWSWYPRCDTR 102
SF WW W+PRC R
Sbjct: 195 SFLRCWWWWWPRCPRR 242
>BI941615
Length = 331
Score = 28.5 bits (62), Expect = 4.6
Identities = 7/13 (53%), Positives = 11/13 (83%)
Frame = -2
Query: 87 SFEAAWWSWYPRC 99
++E++WWSW P C
Sbjct: 39 AYESSWWSWMPSC 1
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.358 0.162 0.621
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,076,546
Number of Sequences: 63676
Number of extensions: 355576
Number of successful extensions: 4210
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 4143
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4201
length of query: 341
length of database: 12,639,632
effective HSP length: 98
effective length of query: 243
effective length of database: 6,399,384
effective search space: 1555050312
effective search space used: 1555050312
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0111b.9


