

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.7
(402 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
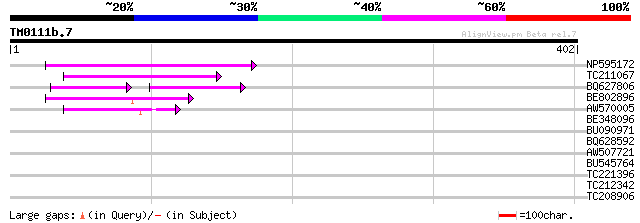
NP595172 polyprotein [Glycine max] 91 9e-19
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 76 2e-14
BQ627806 48 2e-10
BE802896 58 7e-09
AW570005 45 8e-05
BE348096 38 0.007
BU090971 37 0.016
BQ628592 33 0.23
AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine m... 32 0.67
BU545764 30 2.5
TC221396 similar to UP|O64989 (O64989) Steroid 22-alpha-hydroxyl... 29 4.3
TC212342 similar to PIR|T09217|T09217 protein sam2B - spinach {S... 28 7.4
TC208906 homologue to UP|Q9ZS21 (Q9ZS21) Glyoxalase I , partial... 28 9.6
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 90.9 bits (224), Expect = 9e-19
Identities = 52/150 (34%), Positives = 82/150 (54%)
Frame = +1
Query: 26 EESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMA 85
++ +F + +F ++KK +T APVL+LP +P+ + +AS G+G VL Q+ +A
Sbjct: 2566 QKDSFLWNNEAEAAFVKLKKAMTEAPVLSLPDFSQPFILETDASGIGVGAVLGQNGHPIA 2745
Query: 86 YASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMG 145
Y S +L + + EL + AL +RHYL G+ F + ++ + LK L D
Sbjct: 2746 YFSKKLAPRMQKQSAYTRELLAITEALSKFRHYLLGNKFIIRTDQRSLKSLMDQSLQTPE 2925
Query: 146 QRCWMNFIEDYDFTLLYHPGKANVVADALS 175
Q+ W++ YDF + Y PGK N ADALS
Sbjct: 2926 QQAWLHKFLGYDFKIEYKPGKDNQAADALS 3015
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 76.3 bits (186), Expect = 2e-14
Identities = 41/112 (36%), Positives = 63/112 (55%)
Frame = +1
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNY 98
+F +K++LT APVL LP + +E+ C+AS G+ VL+Q +AY S +L NY
Sbjct: 250 AFALLKEKLTKAPVLALPDFSKTFELECDASGVGVRAVLLQGGHPIAYFSEKLHSATLNY 429
Query: 99 LTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWM 150
T+D EL ++ A + W H+L F + S+H+ LKY+ LN W+
Sbjct: 430 PTYDKELYALIRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKWV 585
>BQ627806
Length = 435
Score = 47.8 bits (112), Expect(2) = 2e-10
Identities = 23/68 (33%), Positives = 37/68 (53%)
Frame = +1
Query: 100 THDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDFT 159
T+ ELA + A+K WR YL G F + ++H+ LK L Q+ ++ + +D+T
Sbjct: 232 TYVRELAAITVAVKKWRQYLLGHHFVILTDHRSLKELMSQAVQTPEQQIYLARLMGFDYT 411
Query: 160 LLYHPGKA 167
+ Y GKA
Sbjct: 412 IQYRAGKA 435
Score = 35.0 bits (79), Expect(2) = 2e-10
Identities = 19/57 (33%), Positives = 30/57 (52%)
Frame = +3
Query: 30 FHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAY 86
FH + +F ++K L APVL LP + V +AS G+G +L Q+ +A+
Sbjct: 21 FHWNEEADRAFSQLKLALCQAPVLGLPDFNSSFVVETDASGIGMGAILSQNHHPLAF 191
>BE802896
Length = 416
Score = 58.2 bits (139), Expect = 7e-09
Identities = 34/109 (31%), Positives = 56/109 (51%), Gaps = 4/109 (3%)
Frame = -2
Query: 26 EESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMA 85
+E F + + +F +K+ L P++ P P+E+ C+AS LG VL Q + +
Sbjct: 331 KEVEFDFNDKCK*AFDCLKRALITTPIIQAPDWTAPFELMCDASNYALGVVLAQKIDKLP 152
Query: 86 ----YASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNH 130
Y+S L + NY T + EL +VFAL+ + YL G+ V+ +H
Sbjct: 151 RVIYYSSRTLDAAQANYTTTEKELLAIVFALEKFHSYLLGTRIIVYIDH 5
>AW570005
Length = 413
Score = 44.7 bits (104), Expect = 8e-05
Identities = 30/85 (35%), Positives = 43/85 (50%), Gaps = 2/85 (2%)
Frame = -2
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQL--KLHER 96
+FQ +K +T VL LP +P+ V +AS +G VL Q +A+ S + KL
Sbjct: 262 AFQALKNVVTNTLVLALPDFTKPFTVETDASGSDMGAVLSQEGHPIAFFSKEFCPKLVRS 83
Query: 97 NYLTHDLELAVVVFALKIWRHYLYG 121
+ H ELA + +K WR YL G
Sbjct: 82 STYVH--ELAAITNVVKKWRQYLLG 14
>BE348096
Length = 445
Score = 38.1 bits (87), Expect = 0.007
Identities = 17/38 (44%), Positives = 21/38 (54%)
Frame = -2
Query: 82 EAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYL 119
+ Y +K H R Y THDLEL +V A WRHY+
Sbjct: 180 QVATYVLP*VKTH*RIYPTHDLELVAMVCAFTFWRHYI 67
>BU090971
Length = 447
Score = 37.0 bits (84), Expect = 0.016
Identities = 14/25 (56%), Positives = 20/25 (80%)
Frame = -3
Query: 153 IEDYDFTLLYHPGKANVVADALSHQ 177
++ Y+F +LY P K+N+V DALSHQ
Sbjct: 373 LQGYNFDILYKPDKSNIVVDALSHQ 299
>BQ628592
Length = 423
Score = 33.1 bits (74), Expect = 0.23
Identities = 14/45 (31%), Positives = 27/45 (59%)
Frame = -1
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREA 83
+F+++K+ L VL P+ P+ +Y + +GCVL+QH ++
Sbjct: 378 AFEKIKQSLANPSVLMPPVTGRPFLLYMTMLDESMGCVLVQHDDS 244
>AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine max}, partial
(3%)
Length = 464
Score = 31.6 bits (70), Expect = 0.67
Identities = 13/20 (65%), Positives = 15/20 (75%)
Frame = +2
Query: 156 YDFTLLYHPGKANVVADALS 175
YDF + Y PGK N+ ADALS
Sbjct: 20 YDFIIQYSPGKENIPADALS 79
>BU545764
Length = 639
Score = 29.6 bits (65), Expect = 2.5
Identities = 11/32 (34%), Positives = 19/32 (59%)
Frame = +1
Query: 285 LEVAFMVARNEEACCGVCSCVLNLTESKSGAL 316
L + ++ R EE+CC +CSC+ N + A+
Sbjct: 544 LHIIVVLLRTEESCCLLCSCLDNFHDCHCSAI 639
>TC221396 similar to UP|O64989 (O64989) Steroid 22-alpha-hydroxylase, partial
(12%)
Length = 690
Score = 28.9 bits (63), Expect = 4.3
Identities = 10/23 (43%), Positives = 16/23 (69%)
Frame = -2
Query: 285 LEVAFMVARNEEACCGVCSCVLN 307
L + ++ R EE+CC +CSC+ N
Sbjct: 134 LHIIVVLLRTEESCCLLCSCLNN 66
>TC212342 similar to PIR|T09217|T09217 protein sam2B - spinach {Spinacia
oleracea;} , partial (54%)
Length = 561
Score = 28.1 bits (61), Expect = 7.4
Identities = 16/55 (29%), Positives = 26/55 (47%), Gaps = 4/55 (7%)
Frame = +1
Query: 15 KDC----RTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVY 65
KDC R ++KTH S +R Q K +++ ++T+PL Y+ Y
Sbjct: 310 KDCIALYRQLEKTHPSVSIRRQAAELRYILQAPKLKISQEEMVTIPLIGSTYDSY 474
>TC208906 homologue to UP|Q9ZS21 (Q9ZS21) Glyoxalase I , partial (53%)
Length = 538
Score = 27.7 bits (60), Expect = 9.6
Identities = 12/24 (50%), Positives = 13/24 (54%)
Frame = +3
Query: 44 KKRLTAAPVLTLPLEKEPYEVYCN 67
+ RL PV TLPL K P CN
Sbjct: 180 RNRLPTTPVFTLPLTKPPRVTSCN 251
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.353 0.155 0.544
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,390,272
Number of Sequences: 63676
Number of extensions: 250864
Number of successful extensions: 2892
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 2166
Number of HSP's successfully gapped in prelim test: 83
Number of HSP's that attempted gapping in prelim test: 700
Number of HSP's gapped (non-prelim): 2280
length of query: 402
length of database: 12,639,632
effective HSP length: 99
effective length of query: 303
effective length of database: 6,335,708
effective search space: 1919719524
effective search space used: 1919719524
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0111b.7


