

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.4
(467 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
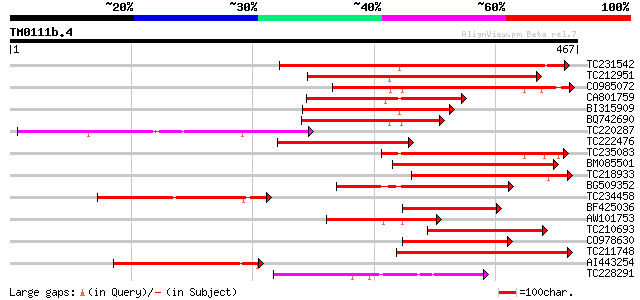
Score E
Sequences producing significant alignments: (bits) Value
TC231542 similar to UP|Q8LBK2 (Q8LBK2) BCS1 protein-like protein... 292 2e-79
TC212951 similar to UP|Q8LBK2 (Q8LBK2) BCS1 protein-like protein... 215 4e-56
CO985072 184 7e-47
CA801759 similar to GP|9294272|dbj mitochondrial protein-like {A... 173 2e-43
BI315909 similar to GP|18086343|gb AT3g50930/F18B3_210 {Arabidop... 167 1e-41
BQ742690 similar to GP|9294272|dbj mitochondrial protein-like {A... 155 4e-38
TC220287 similar to UP|Q9FRW0 (Q9FRW0) Mitochondrial protein-lik... 146 2e-35
TC222476 similar to UP|Q9FN75 (Q9FN75) AAA-type ATPase-like prot... 144 8e-35
TC235083 similar to UP|Q9LH82 (Q9LH82) Similarity to AAA-type AT... 143 1e-34
BM085501 similar to GP|9759053|dbj AAA-type ATPase-like protein ... 143 2e-34
TC218933 similar to UP|Q9FN78 (Q9FN78) AAA-type ATPase-like prot... 137 1e-32
BG509352 weakly similar to GP|18700084|gb At2g46620/F13A10.15 {A... 122 4e-28
TC234458 weakly similar to UP|Q8LBK2 (Q8LBK2) BCS1 protein-like ... 117 1e-26
BF425036 weakly similar to GP|9294274|dbj| mitochondrial protein... 114 1e-25
AW101753 113 2e-25
TC210693 weakly similar to UP|Q9FN78 (Q9FN78) AAA-type ATPase-li... 110 1e-24
CO978630 98 9e-21
TC211748 similar to GB|AAN28905.1|23506091|AY143966 At2g46620/F1... 98 9e-21
AI443254 weakly similar to GP|9759050|dbj| AAA-type ATPase-like ... 96 3e-20
TC228291 homologue to UP|Q9FXT8 (Q9FXT8) 26S proteasome regulato... 69 4e-12
>TC231542 similar to UP|Q8LBK2 (Q8LBK2) BCS1 protein-like protein, partial
(34%)
Length = 1025
Score = 292 bits (748), Expect = 2e-79
Identities = 141/241 (58%), Positives = 183/241 (75%), Gaps = 2/241 (0%)
Frame = +1
Query: 223 DLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNK 282
DL+RF++ KE+Y R GKAWKRGYLLYGPPGTGKSSLIAAMANYL +D+YDL+LT LN N
Sbjct: 1 DLERFVKRKEYYRRVGKAWKRGYLLYGPPGTGKSSLIAAMANYLKFDVYDLELTELNANS 180
Query: 283 DLKNLLLAMSNRSILVIEDIDCSIKLPNREEDEEAAK--NGDNKMTLSGLLNVIDGLWSC 340
+L+ LL+AM+NRSILV+EDIDC+++ +R + AA N D ++TLSGLLN IDGLWS
Sbjct: 181 ELRRLLIAMANRSILVVEDIDCTVEFHDRRAEARAASGHNNDRQVTLSGLLNFIDGLWSS 360
Query: 341 CGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQI 400
CG+ERII+FTTNH+++LDPALLRPGRMD+H+++SYCT F QLA YLGI +H LFE+I
Sbjct: 361 CGDERIIVFTTNHKDKLDPALLRPGRMDVHIHMSYCTPCGFRQLASNYLGIKEHSLFEKI 540
Query: 401 GGLLRHVQVTPAEVAGQLTKIADVEECLHGLVKFLQDKTQNDQNINQDNASQKEKKNQDH 460
++ QVTPAEVA QL K + +E L L+ F++ K + Q + Q+ K+ Q
Sbjct: 541 EEEMQKTQVTPAEVAEQLLKSSHIETSLEQLIDFMRKKKET-QKLEAKKKEQEAKEEQQR 717
Query: 461 E 461
+
Sbjct: 718 K 720
>TC212951 similar to UP|Q8LBK2 (Q8LBK2) BCS1 protein-like protein, partial
(26%)
Length = 808
Score = 215 bits (547), Expect = 4e-56
Identities = 114/204 (55%), Positives = 143/204 (69%), Gaps = 11/204 (5%)
Frame = +1
Query: 246 LLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDCS 305
LLYGP GTGKSSLIAA+ANYL D+YDL+L+++ N +L ++ + RSI+VIEDIDC+
Sbjct: 1 LLYGPXGTGKSSLIAAIANYLKXDVYDLELSSMXSNSELMRVMRETTXRSIIVIEDIDCN 180
Query: 306 IKLPNR-----------EEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHR 354
++ R + D + K + TLSGLLN +DGLWS GEERIIIFTTNHR
Sbjct: 181 KEVHARPTTKPFSDSDSDFDRKRVKVKPYRFTLSGLLNNMDGLWSSGGEERIIIFTTNHR 360
Query: 355 ERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTPAEV 414
ER+DPALLRPGRMDMH++LS+ AF LA YLGI H LFE+I GLL ++VTPA V
Sbjct: 361 ERIDPALLRPGRMDMHIHLSFLKGKAFRVLASNYLGIEDHSLFEEIDGLLEKLEVTPAVV 540
Query: 415 AGQLTKIADVEECLHGLVKFLQDK 438
A QL + D E L GLV+FL++K
Sbjct: 541 AEQLMRNEDPEVALEGLVEFLKEK 612
>CO985072
Length = 839
Score = 184 bits (467), Expect = 7e-47
Identities = 100/227 (44%), Positives = 145/227 (63%), Gaps = 28/227 (12%)
Frame = -3
Query: 267 SYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDCSIKLPNRE----------EDEE 316
+YD+YDL+LTA+ DN DL+ LL+ S++SI+VIEDIDCS+ L + E ++
Sbjct: 837 NYDVYDLELTAVKDNSDLRKLLINTSSKSIMVIEDIDCSLDLTGQRKKRKEKVEGREGKD 658
Query: 317 AAKNGD---------NKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRM 367
+ K GD +K+TLSGLLNVIDG+WS CG ERI++FTTN E+LDPAL+R GRM
Sbjct: 657 SRKRGDEDDDDDDRGSKVTLSGLLNVIDGIWSACGGERIMVFTTNFVEKLDPALIRRGRM 478
Query: 368 DMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTPAEVAGQLTKIA---DV 424
D H+ LSYC + AF+ LA YLG+ H+LF +I LL ++TPA+VA L + +V
Sbjct: 477 DKHIELSYCCYEAFKVLAQNYLGLESHQLFPKIEKLLEETKMTPADVAENLMPKSLDEEV 298
Query: 425 EECLHGLVKFLQ------DKTQNDQNINQDNASQKEKKNQDHEYGID 465
+ CLH L++ L+ +K + + Q N +K +++H G++
Sbjct: 297 DTCLHNLIQALERSKVDLEKKKAETERKQSNV---QKTSENHGEGME 166
>CA801759 similar to GP|9294272|dbj mitochondrial protein-like {Arabidopsis
thaliana}, partial (26%)
Length = 450
Score = 173 bits (438), Expect = 2e-43
Identities = 84/150 (56%), Positives = 112/150 (74%), Gaps = 18/150 (12%)
Frame = +3
Query: 245 YLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDC 304
YLLYGPPGTGKS++IAA+AN+++YD+YDL+LTA+ DN +L+ LL+ ++SI VIEDIDC
Sbjct: 3 YLLYGPPGTGKSTMIAAIANFMNYDVYDLELTAVKDNTELRKLLIETPSKSITVIEDIDC 182
Query: 305 SIKL------------------PNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERI 346
S+ L P R +EE++K+ +K+TLSGLLN IDG+WS CG ERI
Sbjct: 183 SLDLTGQRKKKKEENEDEEQKDPMRRNEEESSKS--SKVTLSGLLNFIDGIWSACGGERI 356
Query: 347 IIFTTNHRERLDPALLRPGRMDMHVNLSYC 376
I+FTTN+ E+LDPAL+R GRMD H+ +SYC
Sbjct: 357 IVFTTNYVEKLDPALIRRGRMDKHIEMSYC 446
>BI315909 similar to GP|18086343|gb AT3g50930/F18B3_210 {Arabidopsis
thaliana}, partial (19%)
Length = 417
Score = 167 bits (422), Expect = 1e-41
Identities = 83/138 (60%), Positives = 103/138 (74%), Gaps = 13/138 (9%)
Frame = +2
Query: 242 KRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIED 301
KRGYLL GPPGTGKSSLIAAMANYL++D+YDL+LT + N DL+ LL+ NRSILV+ED
Sbjct: 2 KRGYLLSGPPGTGKSSLIAAMANYLNFDVYDLELTDVRRNTDLRKLLIGTGNRSILVVED 181
Query: 302 IDCSIKLPNREEDEEAAK-------------NGDNKMTLSGLLNVIDGLWSCCGEERIII 348
IDCS+ L +R ++++ N ++TLSG LN IDGLWS CG+ERII+
Sbjct: 182 IDCSLTLQDRLAKPKSSQPVAITPWPFHPHDNPKPQVTLSGFLNFIDGLWSSCGDERIIV 361
Query: 349 FTTNHRERLDPALLRPGR 366
FTTNH+ +LDPALLRPGR
Sbjct: 362 FTTNHKNKLDPALLRPGR 415
>BQ742690 similar to GP|9294272|dbj mitochondrial protein-like {Arabidopsis
thaliana}, partial (24%)
Length = 405
Score = 155 bits (392), Expect = 4e-38
Identities = 76/134 (56%), Positives = 100/134 (73%), Gaps = 16/134 (11%)
Frame = +2
Query: 241 WKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIE 300
WKRGYLLYGPPGTGKS++IAAMAN+L YD+YDL+LTA+ DN +L+ LL+ S++SI+VIE
Sbjct: 2 WKRGYLLYGPPGTGKSTMIAAMANFLGYDLYDLELTAVKDNTELRKLLIETSSKSIIVIE 181
Query: 301 DIDCSIKLPNR---------EEDEEAAKNG-------DNKMTLSGLLNVIDGLWSCCGEE 344
DIDCS+ L + E+D+ + G +++TLSGLLN IDGLWS CG E
Sbjct: 182 DIDCSLDLTGQRRKKKEEVEEKDQRQKQQGMQEREVKSSQVTLSGLLNFIDGLWSACGGE 361
Query: 345 RIIIFTTNHRERLD 358
R+I+FTTN+ E+LD
Sbjct: 362 RLIVFTTNYVEKLD 403
>TC220287 similar to UP|Q9FRW0 (Q9FRW0) Mitochondrial protein-like protein
(Fragment), partial (44%)
Length = 980
Score = 146 bits (368), Expect = 2e-35
Identities = 88/254 (34%), Positives = 143/254 (55%), Gaps = 10/254 (3%)
Frame = +3
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQG--M 64
K L + SL AT + + +I F P L + + L+ F+ + EF G +
Sbjct: 210 KELWAQMGSLMATIVFMYTIFERFFPPHLREKLQAYTQKLTNHFNPYIQISFPEFSGERL 389
Query: 65 TKNEVYEAAEAYLSTKATLSAQRVKAGKSEDDKT-LSFGADRDEEVTDDFEGVRVTWKLT 123
K+E Y A + YLS ++ A+R+KA D +T L D +EE+TD+F G+++ W+ +
Sbjct: 390 KKSEAYTAIQTYLSANSSQRAKRLKAEVVNDSQTPLVLSMDDNEEITDEFHGIKL-WR-S 563
Query: 124 CIRVDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMA 183
+V ++ R + S E R Y+L+FHK+H++ + SY+ +VL+ K I+ N
Sbjct: 564 ANKVSNNPQRYNPFSYYGS-SDEKRFYKLTFHKRHRDIVTMSYIKHVLDEGKDIEMRNRQ 740
Query: 184 VKIYTVE-------YEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNR 236
+K+YT Y+ + F+HP TF+TLA+D K++++ DL +F +GK++Y +
Sbjct: 741 LKLYTNNPSSGWYGYKQSKWSHIVFEHPATFETLAMDRRKKEDILKDLVKFKKGKDYYAK 920
Query: 237 TGKAWKRGYLLYGP 250
GKAWKRGYLLYGP
Sbjct: 921 IGKAWKRGYLLYGP 962
>TC222476 similar to UP|Q9FN75 (Q9FN75) AAA-type ATPase-like protein, partial
(18%)
Length = 480
Score = 144 bits (363), Expect = 8e-35
Identities = 68/112 (60%), Positives = 87/112 (76%)
Frame = +1
Query: 221 VSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALND 280
+ DLDRF++ KEFY R G+AWKRGYLLYGPPGTGKSSLIAAMANYL +D++DL+L ++
Sbjct: 16 IEDLDRFVKRKEFYKRVGRAWKRGYLLYGPPGTGKSSLIAAMANYLKFDVFDLELGSIVR 195
Query: 281 NKDLKNLLLAMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLN 332
+ DL+ LLLA +NRSILVIEDIDCS+ LP R + K D +++ S L+
Sbjct: 196 DSDLRKLLLATANRSILVIEDIDCSVDLPERRHGDHGRKQADVQVSNSESLS 351
>TC235083 similar to UP|Q9LH82 (Q9LH82) Similarity to AAA-type ATPase,
partial (24%)
Length = 724
Score = 143 bits (361), Expect = 1e-34
Identities = 78/165 (47%), Positives = 104/165 (62%), Gaps = 11/165 (6%)
Frame = +3
Query: 307 KLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGR 366
K P+++++EE KN +K+TLSGLLN IDG+WS CG ERIIIFTTN ++LDPAL+R GR
Sbjct: 72 KDPSKKDEEEGNKN--SKVTLSGLLNFIDGIWSACGGERIIIFTTNFVDKLDPALIRTGR 245
Query: 367 MDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTPAEVAGQLTKIA---D 423
MD H+ LSYC F AF+ LA YL + H LF +I LL VTPA++A L D
Sbjct: 246 MDKHIELSYCRFEAFKVLAKNYLDVDSHYLFARIANLLEVTNVTPADIAENLMPKCLNED 425
Query: 424 VEECLHGLVKFLQDKT----QNDQNINQDNA----SQKEKKNQDH 460
VE CL L++ L K + + +N++ +Q+E KN H
Sbjct: 426 VESCLLNLIQSLDKKVAEEEEEEAGLNEEKVKGEPTQQENKNNGH 560
>BM085501 similar to GP|9759053|dbj AAA-type ATPase-like protein {Arabidopsis
thaliana}, partial (21%)
Length = 421
Score = 143 bits (360), Expect = 2e-34
Identities = 69/138 (50%), Positives = 94/138 (68%), Gaps = 1/138 (0%)
Frame = +3
Query: 316 EAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSY 375
+ K D +++L GLLN IDGLWS CG+ERIII TTNH+ERLDPALLRPGRMDMH+++SY
Sbjct: 3 DGRKQPDVQLSLCGLLNFIDGLWSSCGDERIIILTTNHKERLDPALLRPGRMDMHIHMSY 182
Query: 376 CTFSAFEQLAFTYLGIS-QHKLFEQIGGLLRHVQVTPAEVAGQLTKIADVEECLHGLVKF 434
C++ F+ LA YL I+ H+L +I GL+ +Q+TPA+VA +L K D + L G +K
Sbjct: 183 CSYHGFKVLASNYLDIAPDHRLVGEIEGLIEDMQITPAQVAEELMKSEDADTALEGFLKL 362
Query: 435 LQDKTQNDQNINQDNASQ 452
L+ K D + +
Sbjct: 363 LKRKKMEGDVCENDGSDK 416
>TC218933 similar to UP|Q9FN78 (Q9FN78) AAA-type ATPase-like protein, partial
(21%)
Length = 803
Score = 137 bits (345), Expect = 1e-32
Identities = 69/136 (50%), Positives = 91/136 (66%), Gaps = 4/136 (2%)
Frame = +3
Query: 332 NVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGI 391
N IDGLWS CG+ERIIIFTTNH+ERLDPALLRPGRMDMH+++SYC++ F+ LA YL
Sbjct: 3 NFIDGLWSSCGDERIIIFTTNHKERLDPALLRPGRMDMHIHMSYCSYQGFKILASNYLET 182
Query: 392 -SQHKLFEQIGGLLRHVQVTPAEVAGQLTKIADVEECLHGLVKFLQDKTQND---QNINQ 447
S H LF ++ GL+ +Q+TPA+VA +L K D E L G VK L+ K +N
Sbjct: 183 PSDHPLFGEVEGLIEDIQITPAQVAEELMKNEDPEATLEGFVKLLKRKKMEGDVCENSTP 362
Query: 448 DNASQKEKKNQDHEYG 463
D A ++++ + G
Sbjct: 363 DKAEPTHQQSKRRKVG 410
>BG509352 weakly similar to GP|18700084|gb At2g46620/F13A10.15 {Arabidopsis
thaliana}, partial (30%)
Length = 436
Score = 122 bits (305), Expect = 4e-28
Identities = 62/148 (41%), Positives = 99/148 (66%), Gaps = 2/148 (1%)
Frame = +1
Query: 270 IYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
+YD+DL+ + + DL LL + +S++V+ED+D + E + E A +T SG
Sbjct: 1 VYDVDLSKIRGDSDLMFLLTETTAKSVIVVEDLDRFL-----EPEPETA----TAVTASG 153
Query: 330 LLNVIDGLWS-CCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTY 388
+ + +DG+ S CCGEER+++FT N +E +DP LLRPGR+D+H++ C FSAF+ LA +Y
Sbjct: 154 IQSFMDGIVSACCGEERVMVFTMNSKECVDPDLLRPGRVDVHIHFPVCDFSAFKTLASSY 333
Query: 389 LGISQHKLFEQIGGLLRH-VQVTPAEVA 415
LG+ +HKLF Q+ + RH ++PAE++
Sbjct: 334 LGVRKHKLFAQVEEIFRHGATLSPAEIS 417
>TC234458 weakly similar to UP|Q8LBK2 (Q8LBK2) BCS1 protein-like protein,
partial (19%)
Length = 450
Score = 117 bits (292), Expect = 1e-26
Identities = 60/151 (39%), Positives = 93/151 (60%), Gaps = 8/151 (5%)
Frame = +3
Query: 73 AEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRVDSSSS 132
AE YL K + + +R+K K E D T + +R+E +TD F ++ W L C +V+S
Sbjct: 3 AETYLGAKISPNTRRLKVSKPETDTTFALTMERNESLTDVFRSMKFNWVLVCRQVESRGF 182
Query: 133 RTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKIYTVEY- 191
++++++ SEVRS EL+F+KKHK+ + +YLPY+L AK++KQ A+KI+TV+Y
Sbjct: 183 HNP-RDLNATMKSEVRSLELTFNKKHKDMVLQTYLPYILNEAKSMKQATKALKIFTVDYQ 359
Query: 192 -------EAWCRNGTRFDHPMTFKTLAIDAG 215
+AW G + DHP TF TLA++ G
Sbjct: 360 NMYGNISDAWV--GMKLDHPATFDTLAMERG 446
>BF425036 weakly similar to GP|9294274|dbj| mitochondrial protein-like
{Arabidopsis thaliana}, partial (15%)
Length = 416
Score = 114 bits (284), Expect = 1e-25
Identities = 50/82 (60%), Positives = 65/82 (78%)
Frame = -2
Query: 324 KMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQ 383
++TLSGLLN DGLWSCCGEERI++FTTNHR+ +DPAL+R GRMD+HV+L+ C AF +
Sbjct: 415 RVTLSGLLNFTDGLWSCCGEERIVVFTTNHRDSVDPALVRCGRMDVHVSLATCGAHAFRE 236
Query: 384 LAFTYLGISQHKLFEQIGGLLR 405
LA YLG+ H LF+ + G +R
Sbjct: 235 LARNYLGLESHVLFQAVEGCIR 170
>AW101753
Length = 341
Score = 113 bits (283), Expect = 2e-25
Identities = 56/112 (50%), Positives = 77/112 (68%), Gaps = 18/112 (16%)
Frame = +2
Query: 262 MANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDCSI--------------- 306
MAN+L YD+YDL+LT ++DN +L++LL+ +NRSI+VIEDIDCS+
Sbjct: 2 MANFLCYDVYDLELTKVSDNSELRSLLIQTTNRSIIVIEDIDCSVDITADRTVKVKKSQG 181
Query: 307 -KLPNREEDEEAAKNGD--NKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRE 355
KL R +++ + ++TLSGLLN DGLWSCCGEERI++FTTNHR+
Sbjct: 182 AKLSLRSSNKKGQTGCEESGRVTLSGLLNFTDGLWSCCGEERIVVFTTNHRD 337
>TC210693 weakly similar to UP|Q9FN78 (Q9FN78) AAA-type ATPase-like protein,
partial (18%)
Length = 520
Score = 110 bits (276), Expect = 1e-24
Identities = 52/99 (52%), Positives = 71/99 (71%)
Frame = +1
Query: 345 RIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLL 404
RII+FTTNH+ +LDPALLRPGRMD+H++++YCT F+ LAF YLGI++H LF ++ LL
Sbjct: 1 RIIVFTTNHKNKLDPALLRPGRMDVHIDMTYCTPCGFKMLAFNYLGITEHPLFVEVETLL 180
Query: 405 RHVQVTPAEVAGQLTKIADVEECLHGLVKFLQDKTQNDQ 443
+ VTPAEV Q K D E L L++ L +K +N +
Sbjct: 181 KTTNVTPAEVGEQFLKNEDPEIALESLMELLIEKGRNHE 297
>CO978630
Length = 660
Score = 97.8 bits (242), Expect = 9e-21
Identities = 44/93 (47%), Positives = 69/93 (73%), Gaps = 2/93 (2%)
Frame = -1
Query: 324 KMTLSGLLNVIDGLW-SCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFE 382
+++ SG+LN +D L SCC EER+++FT N +E +DP LLRPGR+D+H++ C FSAF+
Sbjct: 657 EISASGILNFMDALLTSCCAEERVMVFTMNTKEHVDPNLLRPGRVDVHIHFPLCDFSAFK 478
Query: 383 QLAFTYLGISQHKLFEQIGGLLRH-VQVTPAEV 414
LA +YLG+ +HKLF Q+ + ++ ++PAE+
Sbjct: 477 TLASSYLGVKEHKLFPQVQEIFQNGASLSPAEI 379
>TC211748 similar to GB|AAN28905.1|23506091|AY143966 At2g46620/F13A10.15
{Arabidopsis thaliana;} , partial (22%)
Length = 786
Score = 97.8 bits (242), Expect = 9e-21
Identities = 52/148 (35%), Positives = 90/148 (60%), Gaps = 3/148 (2%)
Frame = +2
Query: 319 KNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNH-RERLDPALLRPGRMDMHVNLSYCT 377
K+ N +LS +LN +DG+ SCCGEER+++FT N +E +D A+LRPGR+D+H++ C
Sbjct: 20 KSKSNTTSLSSVLNFMDGIVSCCGEERVMVFTMNETKEEVDQAVLRPGRIDVHIHFPLCD 199
Query: 378 FSAFEQLAFTYLGISQHKLFEQIGGLLR-HVQVTPAEVAG-QLTKIADVEECLHGLVKFL 435
FS F+ LA +YLG+ +HKLF Q+ + + +++PAE+ ++ L ++ L
Sbjct: 200 FSTFKILASSYLGLKEHKLFPQVEEVFQTGARLSPAELGEIMISNRNSPTRALKTVISAL 379
Query: 436 QDKTQNDQNINQDNASQKEKKNQDHEYG 463
Q ++ + + + S + + D+E G
Sbjct: 380 QVQSNGPREGQRLSHSGSGRNSDDNEPG 463
>AI443254 weakly similar to GP|9759050|dbj| AAA-type ATPase-like protein
{Arabidopsis thaliana}, partial (4%)
Length = 392
Score = 96.3 bits (238), Expect = 3e-20
Identities = 48/129 (37%), Positives = 79/129 (61%), Gaps = 5/129 (3%)
Frame = +2
Query: 86 QRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRVDSSSSRTRYTNMSSSLMS 145
+R+K KS +K L+ ++ E+V D F+G W+ C + ++ N S S+ S
Sbjct: 8 ERLKISKSAKEKKLTVRLEKGEKVVDCFDGACFKWRFICAESEKNNPNDHSNNNSISVRS 187
Query: 146 EVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKIYTVEYEAWCRNGTRFD--- 202
E RS+ELSF KK+KE + +SYLP++LE+AK +K E +K++T+ ++C +G ++D
Sbjct: 188 EKRSFELSFPKKYKEMVLDSYLPFILEKAKEMKDEERVLKMHTLN-TSYCYSGVKWDSIN 364
Query: 203 --HPMTFKT 209
HP TF+T
Sbjct: 365 LEHPSTFET 391
>TC228291 homologue to UP|Q9FXT8 (Q9FXT8) 26S proteasome regulatory particle
triple-A ATPase subunit4, partial (62%)
Length = 1079
Score = 68.9 bits (167), Expect = 4e-12
Identities = 57/186 (30%), Positives = 94/186 (49%), Gaps = 9/186 (4%)
Frame = +3
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTA 277
+EL ++ L E + R G +G LLYGPPGTGK+ L A+A+ + + + +A
Sbjct: 6 RELRESIELPLMNPELFIRVGIKPPKGVLLYGPPGTGKTLLARAIASNIEANFLKVVSSA 185
Query: 278 LNDN--KDLKNLLLAMSNRS------ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
+ D + L+ M + I+ +++ID + R E + + + + TL
Sbjct: 186 IIDKYIGESARLIREMFGYARDHQPCIIFMDEIDA---IGGRRFSEGTSADREIQRTLME 356
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCT-FSAFEQLAFTY 388
LLN +DG + G+ ++I+ TN + LDPALLRPGR+D + + S E L
Sbjct: 357 LLNQLDG-FDQLGKVKMIM-ATNRPDVLDPALLRPGRLDRKIEIPLPNEQSRMEILKIHA 530
Query: 389 LGISQH 394
GI++H
Sbjct: 531 AGIAKH 548
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,666,807
Number of Sequences: 63676
Number of extensions: 215577
Number of successful extensions: 1282
Number of sequences better than 10.0: 123
Number of HSP's better than 10.0 without gapping: 1244
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1250
length of query: 467
length of database: 12,639,632
effective HSP length: 101
effective length of query: 366
effective length of database: 6,208,356
effective search space: 2272258296
effective search space used: 2272258296
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0111b.4


