

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111a.1
(863 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
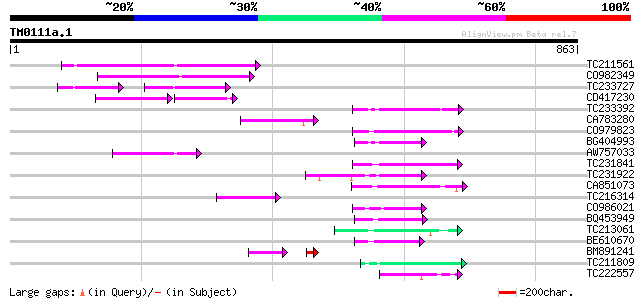
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 146 3e-35
CO982349 108 1e-23
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 60 2e-20
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 63 8e-18
TC233392 77 4e-14
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 75 9e-14
CO979823 74 4e-13
BG404993 71 2e-12
AW757033 69 1e-11
TC231841 67 3e-11
TC231922 65 1e-10
CA851073 62 8e-10
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 62 1e-09
CO986021 61 2e-09
BQ453949 61 2e-09
TC213061 60 3e-09
BE610670 60 5e-09
BM891241 52 1e-08
TC211809 59 1e-08
TC222557 57 5e-08
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 146 bits (369), Expect = 3e-35
Identities = 95/305 (31%), Positives = 148/305 (48%), Gaps = 2/305 (0%)
Frame = +1
Query: 79 LIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTSFGFSEKWVDRIMQCV 138
LIA+E K + K F K++ KAYD + W+FL + FS KW+ I +CV
Sbjct: 13 LIANEVIDEAKRSNKSCLVF---KVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIEECV 183
Query: 139 TGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLKAQEEGRIQGLRMA 198
AS VNG + +PQRG+RQG PL+P+LF + AE ++ L+ +A EE + +
Sbjct: 184 KSASISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRRAVEENLYKPYLVG 363
Query: 199 PECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQMISIEKS--GIIFSRNTL 256
PI+ L +ADD++FF +A +E + IL + L S I+ KS G+
Sbjct: 364 ANGVPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKSCFGVF---GVT 534
Query: 257 SSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVLDKIDGWKEKLLNQAG 316
K + ++ L S P YLG+P R + + I + K+ WK + ++ G
Sbjct: 535 DQWKQEAANYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKCERKLSKWK*RHISFGG 714
Query: 317 KEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKERGIHWKDWHFISKAK 376
+ LIK+VL ++ Y S R PQ+ + F W ++ I W W + K
Sbjct: 715 RVTLIKSVLTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGGGPDQKKISWIRWEKLCLPK 894
Query: 377 CKGGL 381
+GG+
Sbjct: 895 ERGGI 909
>CO982349
Length = 795
Score = 108 bits (269), Expect = 1e-23
Identities = 71/241 (29%), Positives = 111/241 (45%), Gaps = 2/241 (0%)
Frame = +3
Query: 134 IMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLKAQEEGRIQ 193
I C+ AS VN PQRG+RQG PL+P LF + AE ++ L+ +A + R
Sbjct: 72 IRGCLHSASVSILVNESPINEFSPQRGLRQGDPLAPLLFNIVAEGLTGLMREAVDRKRFN 251
Query: 194 GLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQMISIEKS--GIIF 251
+ P++ L +ADD++FF +A E + +IL + LASG I+ +S G I+
Sbjct: 252 SFLVGKNKEPVSILQYADDTIFFEEATMENMKVIKTILRCFELASGLKINFAESRFGAIW 431
Query: 252 SRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVLDKIDGWKEKL 311
+ + + L S P YLG+P E + I + K+ WK++
Sbjct: 432 KPDHWCK---EAAEFLNCSMLSMPFSYLGIPIEANPRCREI*DPIIRKCETKLARWKQRH 602
Query: 312 LNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKERGIHWKDWHF 371
++ G+ LI A+L A+ Y S R P Q + +F W +R I W W
Sbjct: 603 ISLGGRVTLINAILTALPIYFFSFFRAPSKVVQKLVNIQRKFLWGGGSXQRRIAWVKWET 782
Query: 372 I 372
+
Sbjct: 783 V 785
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 60.5 bits (145), Expect(2) = 2e-20
Identities = 38/131 (29%), Positives = 62/131 (47%)
Frame = -2
Query: 205 THLLFADDSLFFAKANREEAYQLISILNDYTLASGQMISIEKSGIIFSRNTLSSTKHQIS 264
+H++ ADD + F K + LI+ Y + GQ+IS KS I F + IS
Sbjct: 451 SHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKI-FIGSIPGQRLRVIS 275
Query: 265 SILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVLDKIDGWKEKLLNQAGKEVLIKAV 324
+L+ D P Y G+P GK L + ++ K+ WK +L+ G+ L+ +V
Sbjct: 274 DLLQYFHSDIPFNYFGVPIFKGKPTRRHLYCVADKIKCKLASWKGSILSIMGRIQLVNSV 95
Query: 325 LQAMSAYTMSI 335
+ M Y+ S+
Sbjct: 94 IHGMLLYSFSV 62
Score = 58.2 bits (139), Expect(2) = 2e-20
Identities = 35/99 (35%), Positives = 52/99 (52%)
Frame = -3
Query: 74 IQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTSFGFSEKWVDR 133
I+D + E + L G +A+K+++ KA+D L+ DFL + L FG+S + +
Sbjct: 834 IKDCTCVTSEVINMLDKKVFGGN--LALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNW 661
Query: 134 IMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLF 172
I + A VNGE QRG+RQG PLSP L+
Sbjct: 660 IRVILLSAKLSISVNGEPVGFFSCQRGVRQGDPLSPSLY 544
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 63.2 bits (152), Expect(2) = 8e-18
Identities = 34/117 (29%), Positives = 62/117 (52%)
Frame = -3
Query: 131 VDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLKAQEEG 190
V + C++ + NGEV P G+RQ P++PYLFVL E +SHL+ +
Sbjct: 686 VQLVWYCMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFVLCLERLSHLINLVMNKK 507
Query: 191 RIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQMISIEKS 247
+ +++ I+HL F DD + F +AN ++ + L ++ +SG ++++K+
Sbjct: 506 L*RSIQVNRGGLLISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKT 336
Score = 46.2 bits (108), Expect(2) = 8e-18
Identities = 30/96 (31%), Positives = 49/96 (50%)
Frame = -2
Query: 251 FSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVLDKIDGWKEK 310
FS+N + ISS L + + D GKYLG+P ++T ++I +V + WK
Sbjct: 324 FSKNVNWWVEEDISSGLGLQSTDDLGKYLGVPIFHKNPTSHTFDFIIDKVNKRFSRWKSN 145
Query: 311 LLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAI 346
LL+ G+ L K V+ A+ ++ I++F C I
Sbjct: 144 LLSMVGRLTLTK*VV*ALPSH---IMQFDNLLCMGI 46
>TC233392
Length = 1145
Score = 76.6 bits (187), Expect = 4e-14
Identities = 48/170 (28%), Positives = 75/170 (43%), Gaps = 1/170 (0%)
Frame = -1
Query: 522 QVPDKLCWPHTKDGEYTVKTGYQVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMF 581
QVPD W H + Y+ ++ Y++ L+ G D +W +++P K +F
Sbjct: 665 QVPDTWVWKHEPNELYSTRSAYKL---LQGHLEGED----QDGALQDLWKLKIPAKASIF 507
Query: 582 LWKACNEALPVKASLFRRHIA-PDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQH 640
W+ + LP K++L RR + D +C C EE H+ L C R +W SQ V+
Sbjct: 506 AWRLIRDRLPTKSNLHRRQVVLEDSLCPFCRIREEDASHIFLECNKIRPLWWESQTWVRS 327
Query: 641 LHQGSLTVKDWLTQLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQ 690
L + + Q V S+ +R T W +WK RN+ FQ
Sbjct: 326 LGVSPINKRQHFLQHV-HGTPGSKRYNRWKTWWIALTWSLWKHRNQVIFQ 180
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 75.5 bits (184), Expect = 9e-14
Identities = 35/123 (28%), Positives = 57/123 (45%), Gaps = 5/123 (4%)
Frame = +2
Query: 352 RFWWRSPGKERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVR 411
RF W R I W +W + K KGGLG KD T N LL K W ++ W +
Sbjct: 59 RFLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTTLLGKWRWDLFYIQQEPWAK 238
Query: 412 TLKGLYFPNSSFLQSGRTARNSWGWQSILQGKEF-----LKEAGLWIIGDGSEVKVWKDK 466
L+ Y + + +++S W+ +++ ++ LK +W +G G +K W+D
Sbjct: 239 VLQSKYGGWRALEEGSSGSKDSAWWKDLIKTQQLQRNIPLKRETIWKVGGGDRIKFWEDL 418
Query: 467 WVS 469
W +
Sbjct: 419 WTN 427
>CO979823
Length = 853
Score = 73.6 bits (179), Expect = 4e-13
Identities = 48/170 (28%), Positives = 72/170 (42%), Gaps = 1/170 (0%)
Frame = -2
Query: 522 QVPDKLCWPHTKDGEYTVKTGYQVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMF 581
Q DK W G Y+ K+GY V + G + D + +IW +++P K +F
Sbjct: 711 QREDKWLWKPEPGGHYSTKSGYHVLW-------GELTEEIQDADFAEIWKLKIPTKAAVF 553
Query: 582 LWKACNEALPVKASLFRRHI-APDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQH 640
W+ + LP K++L RR + D VC +CN EE HL C T +W S V
Sbjct: 552 AWRLVRDRLPTKSNLRRRQVMVQDMVCPLCNNIEEGAAHLFFNCTKTLPLWWESMSWVNL 373
Query: 641 LHQGSLTVKDWLTQLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQ 690
T + Q + + + +A W IW+ RN+ FQ
Sbjct: 372 KTAMPQTPRQHFLQYGTDIADGLKSKRWKCWWIA-LTWTIWQHRNKVVFQ 226
>BG404993
Length = 412
Score = 71.2 bits (173), Expect = 2e-12
Identities = 38/111 (34%), Positives = 54/111 (48%), Gaps = 1/111 (0%)
Frame = -3
Query: 525 DKLCWPHTKDGEYTVKTGYQVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWK 584
D W G Y+ KT Y++ L+E RG D I+ Q+W +++P K +F W+
Sbjct: 332 DSWTWKADPSGNYSTKTAYEL---LQEAIRGDN----EDGIFVQLWKLKIPPKAPVFTWR 174
Query: 585 ACNEALPVKASLFRRHI-APDPVCFICNQDEESVYHLLLGCPWTRAVWLSS 634
N+ LP K +L RR + D +C +CN EE HL C T W S
Sbjct: 173 LINDRLPTKVNLRRRQVEISDSLCPLCNNSEEDAAHLFFNCNKTLPFWWES 21
>AW757033
Length = 441
Score = 68.6 bits (166), Expect = 1e-11
Identities = 44/138 (31%), Positives = 69/138 (49%), Gaps = 2/138 (1%)
Frame = -2
Query: 157 PQRGIRQGYPLSPYLFVLAAETMSHLLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFF 216
P RG+RQG PL+P LF +AAE ++ L+ +A + + P++ L +ADD++FF
Sbjct: 425 PHRGLRQGDPLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKKPVSILQYADDTIFF 246
Query: 217 AKANREEAYQLISILNDYTLASGQMISIEKS--GIIFSRNTLSSTKHQISSILRMSAWDS 274
+A E + +IL + LASG I+ KS G I + + + + + S
Sbjct: 245 GEATMENVRVIKTILRGFELASGLKINFAKSRFGAI---SVPD*WRKEAAEFMNCSLLSL 75
Query: 275 PGKYLGLPAEWGKSRNNT 292
P YLG+P R T
Sbjct: 74 PFSYLGIPIGANPRRRET 21
>TC231841
Length = 791
Score = 67.0 bits (162), Expect = 3e-11
Identities = 48/171 (28%), Positives = 71/171 (41%), Gaps = 3/171 (1%)
Frame = +3
Query: 522 QVPDKLCWPHTKDGEYTVKTGYQVAYS--LKEQARGPQPHILNDPIWNQIWGIQVP*KIK 579
Q D+ W G+YT K+ Y V + +EQ D ++ ++W +++P KI
Sbjct: 54 QQGDQWVWKADPSGQYTAKSAYGVLWGEMFEEQ---------QDGVFEELWKLKLPSKIT 206
Query: 580 MFLWKACNEALPVKASLFRRHI-APDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHV 638
+F W+ + LP +++L R+ I DP C C EES HL C VW S V
Sbjct: 207 IFAWRLIRDRLPTRSNLRRKQIEVDDPRCPFCRSAEESAAHLFFHCSRIAPVWWESLSWV 386
Query: 639 QHLHQGSLTVKDWLTQLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHF 689
L + Q + + SR W IWK+RN F
Sbjct: 387 NLLGVFPNHPRQHFLQHIYGVTAGMR-ASRWKWWWLALTWTIWKQRNNMIF 536
>TC231922
Length = 803
Score = 65.5 bits (158), Expect = 1e-10
Identities = 47/199 (23%), Positives = 82/199 (40%), Gaps = 15/199 (7%)
Frame = -3
Query: 451 LWIIGDGSEVKVWKDKWVSD---------QVLQAPEAGDAELTVNSLILPDVKQWNLPKL 501
+W +G G ++K W+D W+S+ Q+ + ++ + + W
Sbjct: 708 VWRVGCGDKIKFWQDSWLSEGCNLQQKYNQLYTISRQ*NLTISKMGKFSQNARSWEFKWR 529
Query: 502 LSTFPRQIAFKV-FQIPLS----LTQVPDKLCWPHTKDGEYTVKTGYQVAYSLKEQARGP 556
F + A V F +S Q D + W G Y+ K+ AY L + P
Sbjct: 528 RRLFDYEYAMAVDFMNEISGISIQNQAHDSMFWKADSSGVYSTKS----AYRLLMPSISP 361
Query: 557 QPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFRRHI-APDPVCFICNQDEE 615
P + I+ +W +++P + +F W+ + LP + +L RR I D +C +C E
Sbjct: 360 AP---SRRIFQILWHLKIPPRAAVFSWRLFLDRLPTRGNLSRRSIPIQDIMCPLCGCQHE 190
Query: 616 SVYHLLLGCPWTRAVWLSS 634
HL C T+ +W S
Sbjct: 189 EAGHLFFHCKMTKGLWWES 133
>CA851073
Length = 633
Score = 62.4 bits (150), Expect = 8e-10
Identities = 41/183 (22%), Positives = 74/183 (40%), Gaps = 7/183 (3%)
Frame = +1
Query: 521 TQVPDKLCWPHTKDGEYTVKTGYQVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKM 580
+ + D + W G Y+ K+ Y++ + H+ I+ IW +++P + +
Sbjct: 16 SHLQDTMVWKADPCGVYSTKSAYRILMTCNR-------HVSEANIFKTIWKLKIPPRAAV 174
Query: 581 FLWKACNEALPVKASLFRRHIA-PDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQ 639
F W+ + LP + +L RR+++ + C +C ++E HL C TR +W S Q
Sbjct: 175 FSWRLIKDRLPTRHNLLRRNVSIQENECPLCGYEQEEADHLFFNCKMTRGLWWESMRWNQ 354
Query: 640 HLHQGSLTVKDWLTQLVIRLQENSEDLSRGLTIVAYYLW------GIWKERNECHFQQRP 693
+ S++ Q + G + W IWK RN FQ
Sbjct: 355 IVGPLSVSPASHFVQF-------CDGFGAGRNHTRWCGWWIALTSSIWKHRNLLIFQGNQ 513
Query: 694 PDP 696
+P
Sbjct: 514 FEP 522
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 61.6 bits (148), Expect = 1e-09
Identities = 31/96 (32%), Positives = 42/96 (43%)
Frame = +2
Query: 316 GKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKERGIHWKDWHFISKA 375
G+ LIK+VL A+ +S + PQ + +F W I W W I
Sbjct: 8 GRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADICNP 187
Query: 376 KCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVR 411
K GGLG KD S N AL + W + + + LW R
Sbjct: 188 KIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWAR 295
>CO986021
Length = 900
Score = 60.8 bits (146), Expect = 2e-09
Identities = 34/114 (29%), Positives = 55/114 (47%), Gaps = 1/114 (0%)
Frame = -1
Query: 522 QVPDKLCWPHTKDGEYTVKTGYQVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMF 581
Q+ D + W G Y+ K+ AY L P H N I +W +++P + ++F
Sbjct: 792 QLQDSMLWKADPTGIYSTKS----AYRLLLPNNRPGQHSSNFKI---LWKLKIPPRAELF 634
Query: 582 LWKACNEALPVKASLFRRHIA-PDPVCFICNQDEESVYHLLLGCPWTRAVWLSS 634
W+ + L ++A+L RRH+A D +C +C +E HL C T +W S
Sbjct: 633 SWRLFRDRLSIRANLLRRHVALQDIMCPLCGNHQEEAGHLFFHCRMTIGLWWES 472
>BQ453949
Length = 450
Score = 60.8 bits (146), Expect = 2e-09
Identities = 32/113 (28%), Positives = 55/113 (48%), Gaps = 1/113 (0%)
Frame = +3
Query: 525 DKLCWPHTKDGEYTVKTGYQVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWK 584
D L W DG Y+ K+ Y + Q + D + IW +++P + F +
Sbjct: 30 DLLLWGSNSDGSYSTKSAYNFLKNEDSQT-------IEDSAFKNIWNLKLPPRACAFS*R 188
Query: 585 ACNEALPVKASLFRRHI-APDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQL 636
+P K +L RRH+ P C +C+++EE+V H++ C TR +W ++L
Sbjct: 189 IF*NRIPTKVNLRRRHVKLPSSNCPLCDEEEETVGHVMYSCIKTRHLWWETEL 347
>TC213061
Length = 823
Score = 60.5 bits (145), Expect = 3e-09
Identities = 51/208 (24%), Positives = 79/208 (37%), Gaps = 13/208 (6%)
Frame = +2
Query: 495 QWNLPKLLSTFPRQIAF-KVFQIPLSLTQVPDKLCWPHTKDGEYTVKTGYQVAYSLKEQA 553
QW P S +AF + + + + D+ W G Y ++ Y+V
Sbjct: 35 QWRRPLFDSEIEMLVAFIQEVEGHKIRSDIEDQWVWAAESSGSYLARSAYRVI------- 193
Query: 554 RGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFRRHI-APDPVCFICNQ 612
R P D + ++W ++VP K+ MF W+ + LP + +L R+ + + VC +C
Sbjct: 194 REGIPEEEQDREFKELWKLKVPMKVTMFAWRLLKDILPTRDNLRRKRVELHEYVCPLCRS 373
Query: 613 DEESVYHLLLGCPWTRAVWLSSQLHVQ-----------HLHQGSLTVKDWLTQLVIRLQE 661
+ES HL C +W S V+ H HQ S V L
Sbjct: 374 MDESASHLFFHCSKILPIWWESLSWVKLVGAFPHHPRHHFHQHSHEVYQGLQG------- 532
Query: 662 NSEDLSRGLTIVAYYLWGIWKERNECHF 689
+R W IWK RN+ F
Sbjct: 533 -----NRWKWWWLALTWTIWKHRNDIIF 601
>BE610670
Length = 368
Score = 59.7 bits (143), Expect = 5e-09
Identities = 32/108 (29%), Positives = 51/108 (46%), Gaps = 1/108 (0%)
Frame = -2
Query: 525 DKLCWPHTKDGEYTVKTGYQVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWK 584
D L W G Y+ K+ Y + +G D IW +++P + F W+
Sbjct: 313 DSLIWRADPSGAYSTKSAYNLL-------KGEGSFDNEDSASKIIWNLKIPPRTIAFSWR 155
Query: 585 ACNEALPVKASLFRRHI-APDPVCFICNQDEESVYHLLLGCPWTRAVW 631
LP KA+L RR + P C +C+ +EE+V H++ C TR++W
Sbjct: 154 IFKNRLPTKANLRRRQVELPSYRCPLCDLEEENVGHIMFSCTRTRSLW 11
>BM891241
Length = 407
Score = 51.6 bits (122), Expect(2) = 1e-08
Identities = 24/60 (40%), Positives = 29/60 (48%)
Frame = -2
Query: 364 IHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNSSF 423
I W W + K KGGLG KD S N+ALL K W + + LW R + Y S F
Sbjct: 382 IPWVKWEVVCLPKTKGGLGIKDISKFNIALLGKWIWALASDQQQLWARIINSKYGGWSDF 203
Score = 26.9 bits (58), Expect(2) = 1e-08
Identities = 7/19 (36%), Positives = 12/19 (62%)
Frame = -3
Query: 452 WIIGDGSEVKVWKDKWVSD 470
W +G G ++ W DKW+ +
Sbjct: 102 WKVGCGDKINFWTDKWLGE 46
>TC211809
Length = 895
Score = 58.5 bits (140), Expect = 1e-08
Identities = 46/162 (28%), Positives = 63/162 (38%), Gaps = 1/162 (0%)
Frame = +2
Query: 535 GEYTVKTGYQVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKA 594
G Y+ K+ AY L +A G P D + +W +++P K +F W+ + LP
Sbjct: 200 GNYSTKS----AYRLLMEATGEFPV---DRTYVDLWNLKIPSKAIVFAWRLIKDRLPTWT 358
Query: 595 SLFRRHI-APDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWLT 653
+L R + D C +CN EE HL C T +W +Q V L KD
Sbjct: 359 NLRVRQVELNDSRCPLCNSSEEDAAHLFFHCTKTEXLWWETQSWVNSLGAFPHNPKDHFL 538
Query: 654 QLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPD 695
Q R R + W IWK RN F D
Sbjct: 539 QHDNR-SAGGVXARRWKCLWVALTWVIWKHRNGVVFHNHSFD 661
>TC222557
Length = 1002
Score = 56.6 bits (135), Expect = 5e-08
Identities = 39/139 (28%), Positives = 58/139 (41%), Gaps = 12/139 (8%)
Frame = +1
Query: 563 DPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFRRHI-APDPVCFICNQDEESVYHLL 621
D +W +++P K F W+ E +P K +L+RR + + +C CN+ EE HL
Sbjct: 421 DEALEDLWQLKIPLKATTFAWRLIKERIPTKGNLWRRRVQLNNLMCPFCNRQEEEASHLF 600
Query: 622 LGCP-----------WTRAVWLSSQLHVQHLHQGSLTVKDWLTQLVIRLQENSEDLSRGL 670
CP WT+ V + S + Q+ Q +L K LQ + L
Sbjct: 601 FNCPRILPLWWESLAWTKTVGVFSPIPRQNYMQHTLISKG------KHLQARWKCWWVAL 762
Query: 671 TIVAYYLWGIWKERNECHF 689
T W IW+ RN F
Sbjct: 763 T------WSIWRHRNRIVF 801
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,494,523
Number of Sequences: 63676
Number of extensions: 757293
Number of successful extensions: 3526
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 3468
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3501
length of query: 863
length of database: 12,639,632
effective HSP length: 105
effective length of query: 758
effective length of database: 5,953,652
effective search space: 4512868216
effective search space used: 4512868216
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0111a.1


