

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.15
(184 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
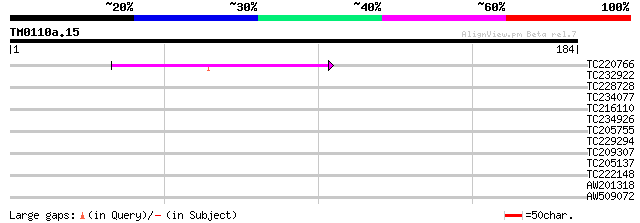
Score E
Sequences producing significant alignments: (bits) Value
TC220766 homologue to UP|Q9Q5L3 (Q9Q5L3) EBNA-2, partial (4%) 55 2e-08
TC232922 weakly similar to UP|Q84VZ3 (Q84VZ3) At5g52990, partial... 37 0.004
TC228728 weakly similar to UP|Q84TD4 (Q84TD4) At2g17540, partial... 30 0.64
TC234077 homologue to UP|Q6WV72 (Q6WV72) Histone H4, complete 28 2.4
TC216110 weakly similar to UP|P93333 (P93333) PR10-1 protein, pa... 27 4.2
TC234926 similar to UP|Q9SYN8 (Q9SYN8) F9H16.2 protein, partial ... 27 4.2
TC205755 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-ric... 27 5.5
TC229294 27 5.5
TC209307 similar to GB|AAO64839.1|29028920|BT005904 At3g53440 {A... 27 5.5
TC205137 similar to PIR|T52133|T52133 potassium channel beta sub... 27 7.1
TC222148 weakly similar to UP|Q75ZI4 (Q75ZI4) Prolyl 4-hydroxyla... 27 7.1
AW201318 27 7.1
AW509072 26 9.3
>TC220766 homologue to UP|Q9Q5L3 (Q9Q5L3) EBNA-2, partial (4%)
Length = 885
Score = 54.7 bits (130), Expect = 2e-08
Identities = 32/73 (43%), Positives = 43/73 (58%), Gaps = 1/73 (1%)
Frame = +2
Query: 34 IYFAIFDHRHTKSQTLTFLNRIRSSLKETL-DSVNDSTAPPPLSLQTPFDLILQEILHLD 92
++FAI H KSQTL FL+ IRSS ++TL S N++ PLS Q+ F I Q LH
Sbjct: 2 VFFAISHHSLLKSQTLPFLHCIRSSFRQTLPPSSNNNHHFAPLSFQSQFTSIFQNALHHH 181
Query: 93 DGNRSPPAVVGIS 105
+ SPP+ +S
Sbjct: 182 HHHHSPPSATSLS 220
>TC232922 weakly similar to UP|Q84VZ3 (Q84VZ3) At5g52990, partial (15%)
Length = 584
Score = 37.4 bits (85), Expect = 0.004
Identities = 35/112 (31%), Positives = 50/112 (44%), Gaps = 2/112 (1%)
Frame = +2
Query: 45 KSQTLTFLNRIRSSLKETLDSVNDSTAPPPLSLQTPFDLILQEILHLDDGNRSPPAVVGI 104
KSQTL FL+RIRSS ++TL N+ PLS Q+ F I + +
Sbjct: 5 KSQTLPFLHRIRSSFRQTLPPTNNHHF-APLSFQSQFHTIFHIFASRQQQIAATTTTLSX 181
Query: 105 SH--DEGLKKKKMVVDSATVIANRTSAKGLAAAAATELVELEKDAVEEALVG 154
S GL++K+ A A R + A A A + + + +EE VG
Sbjct: 182 SQPSSSGLEEKEKAQ*RAR-HARRLRRRRFAQAPAPQPQQQGQARLEETRVG 334
>TC228728 weakly similar to UP|Q84TD4 (Q84TD4) At2g17540, partial (25%)
Length = 1093
Score = 30.0 bits (66), Expect = 0.64
Identities = 22/90 (24%), Positives = 38/90 (41%), Gaps = 2/90 (2%)
Frame = +3
Query: 65 SVNDSTAPPPLSLQTPFDLILQEIL--HLDDGNRSPPAVVGISHDEGLKKKKMVVDSATV 122
+V++ +P S + F+ L+ +L H D+ N +P VV S + K +
Sbjct: 279 TVSEVASPSLTSYSSEFEARLKNLLGSHSDNPNLTPEEVVEASKAAAVAASKAAKAARAA 458
Query: 123 IANRTSAKGLAAAAATELVELEKDAVEEAL 152
+ A AAA ++L EEA+
Sbjct: 459 AKEKAETAAKAVAAAKRALDLVSSFSEEAV 548
>TC234077 homologue to UP|Q6WV72 (Q6WV72) Histone H4, complete
Length = 605
Score = 28.1 bits (61), Expect = 2.4
Identities = 12/24 (50%), Positives = 17/24 (70%)
Frame = -3
Query: 83 LILQEILHLDDGNRSPPAVVGISH 106
L L+ I ++ NR PP+V+GISH
Sbjct: 345 LSLESINNIHGSNRLPPSVLGISH 274
>TC216110 weakly similar to UP|P93333 (P93333) PR10-1 protein, partial (92%)
Length = 772
Score = 27.3 bits (59), Expect = 4.2
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = -3
Query: 11 STHSLFSHTVNNRTYTFLIDPPF 33
S+ S+ S V N T TFL+DPPF
Sbjct: 527 SSLSVASPLVKNWTLTFLMDPPF 459
>TC234926 similar to UP|Q9SYN8 (Q9SYN8) F9H16.2 protein, partial (8%)
Length = 467
Score = 27.3 bits (59), Expect = 4.2
Identities = 9/23 (39%), Positives = 16/23 (69%)
Frame = +3
Query: 13 HSLFSHTVNNRTYTFLIDPPFIY 35
HSL+SHT+ ++Y + PP+ +
Sbjct: 372 HSLYSHTL*TQSYDISLPPPYYF 440
>TC205755 weakly similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich
glycoprotein DZ-HRGP precursor, partial (6%)
Length = 1036
Score = 26.9 bits (58), Expect = 5.5
Identities = 21/73 (28%), Positives = 32/73 (43%), Gaps = 10/73 (13%)
Frame = +2
Query: 106 HDEGLKKKKMVVDSATVIANRTSAKGLAAA---AATELVELEK-------DAVEEALVGS 155
+ ++ K+ V A ++A +K +AA A L E EK DA L
Sbjct: 62 NSRAVQAKRHFVQGAQLLAQARRSKSASAATSLAKQALAEAEKALALDPKDAASHLLKSM 241
Query: 156 AIQIEGFRRVWLD 168
A+ ++GFR LD
Sbjct: 242 ALDLQGFRTNALD 280
>TC229294
Length = 741
Score = 26.9 bits (58), Expect = 5.5
Identities = 26/74 (35%), Positives = 33/74 (44%), Gaps = 4/74 (5%)
Frame = -3
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHT----KSQTLTFLNRIRSSLKETLDSVND 68
HS HT T +F+ P H H KS TL +S LK L S+
Sbjct: 472 HSCVHHT---ETCSFVASPS-------SHTHLSF**KSSTLDQ----QSLLKPPLPSL-- 341
Query: 69 STAPPPLSLQTPFD 82
T+PPP++L PFD
Sbjct: 340 -TSPPPMTLPHPFD 302
>TC209307 similar to GB|AAO64839.1|29028920|BT005904 At3g53440 {Arabidopsis
thaliana;} , partial (6%)
Length = 1111
Score = 26.9 bits (58), Expect = 5.5
Identities = 12/26 (46%), Positives = 14/26 (53%)
Frame = +1
Query: 14 SLFSHTVNNRTYTFLIDPPFIYFAIF 39
SL VNN Y FL PP ++F F
Sbjct: 988 SLLPSLVNNYFYFFLNQPPVVFFCSF 1065
>TC205137 similar to PIR|T52133|T52133 potassium channel beta subunit homolog
[imported] - Arabidopsis thaliana {Arabidopsis thaliana;}
, complete
Length = 1525
Score = 26.6 bits (57), Expect = 7.1
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = -1
Query: 162 FRRVWLDLRSRLSLLHESRSE 182
FR +WL L + +LLH +RS+
Sbjct: 1126 FRALWLALNNSFNLLHHNRSQ 1064
>TC222148 weakly similar to UP|Q75ZI4 (Q75ZI4) Prolyl 4-hydroxylase, partial
(40%)
Length = 619
Score = 26.6 bits (57), Expect = 7.1
Identities = 15/49 (30%), Positives = 25/49 (50%), Gaps = 3/49 (6%)
Frame = -1
Query: 70 TAPPPLSLQT---PFDLILQEILHLDDGNRSPPAVVGISHDEGLKKKKM 115
+A P L +Q PF L + + HL R PP V + + GL ++++
Sbjct: 451 SAKPVLMVQVWALPFQLFVGILPHL*SLERKPPVVSNLKSEMGLNQQRV 305
>AW201318
Length = 475
Score = 26.6 bits (57), Expect = 7.1
Identities = 15/50 (30%), Positives = 24/50 (48%)
Frame = +1
Query: 25 YTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDSTAPPP 74
Y+ L F YF + HT + + R++ L+ D+ +DSTA P
Sbjct: 46 YSRLYSNIFNYFCRLPNPHTPFPRVPHIVRLQEPLRGLYDARSDSTATRP 195
>AW509072
Length = 428
Score = 26.2 bits (56), Expect = 9.3
Identities = 13/35 (37%), Positives = 17/35 (48%)
Frame = -1
Query: 70 TAPPPLSLQTPFDLILQEILHLDDGNRSPPAVVGI 104
T PPPLS TPF + + +DD + P I
Sbjct: 425 TPPPPLS*YTPFSIWRRNTSSIDDSPLNFPIAFSI 321
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,797,868
Number of Sequences: 63676
Number of extensions: 104830
Number of successful extensions: 583
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 581
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 582
length of query: 184
length of database: 12,639,632
effective HSP length: 91
effective length of query: 93
effective length of database: 6,845,116
effective search space: 636595788
effective search space used: 636595788
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0110a.15


